HAFNI report on finalized components on task: SOCIAL |
Totally 3 copes finalized |
Subject: 1 |
GLM result cope: 01, threshold: 4
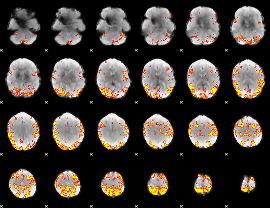 |
GLM result cope: 02, threshold: 4
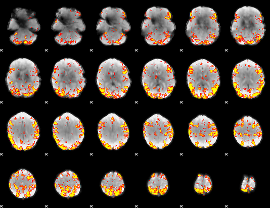 |
GLM result cope: 06, threshold: 2.5
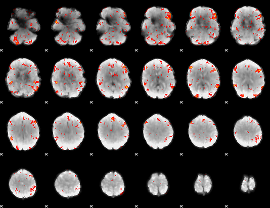 |
HAFNI result component: 266
Overlap rate: 0.33792
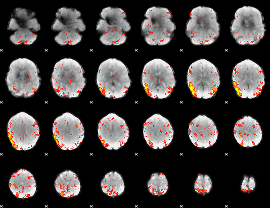 |
HAFNI result component: 266
Overlap rate: 0.30058
 |
HAFNI result component: 361
Overlap rate: 0.55988
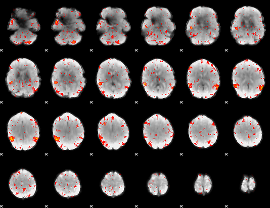 |
Correlation: 0.21187
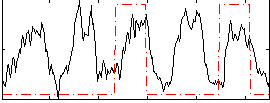  |
Correlation: 0.41513
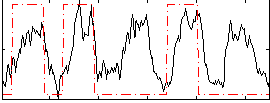  |
Correlation: 0.40107
  |
Subject: 2 |
GLM result cope: 01, threshold: 3.5
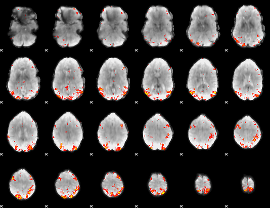 |
GLM result cope: 02, threshold: 4
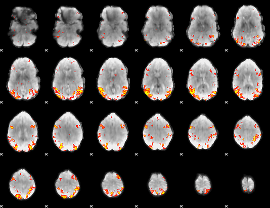 |
GLM result cope: 06, threshold: 2
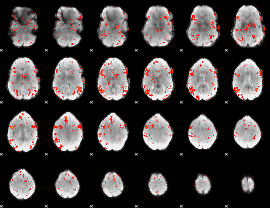 |
HAFNI result component: 321
Overlap rate: 0.55019
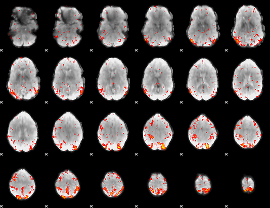 |
HAFNI result component: 332
Overlap rate: 0.42013
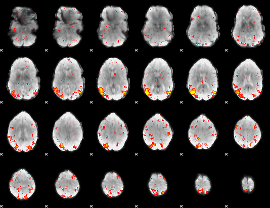 |
HAFNI result component: 144
Overlap rate: 0.64489
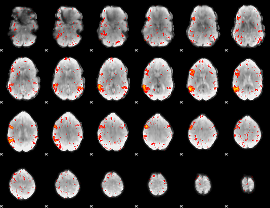 |
Correlation: 0.21734
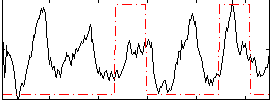  |
Correlation: 0.48842
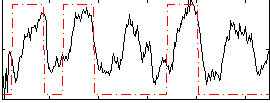  |
Correlation: 0.19135
  |
Subject: 3 |
GLM result cope: 01, threshold: 4
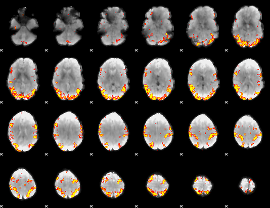 |
GLM result cope: 02, threshold: 4
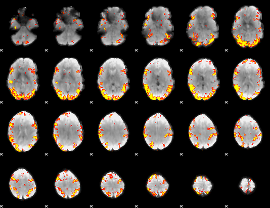 |
GLM result cope: 06, threshold: 3
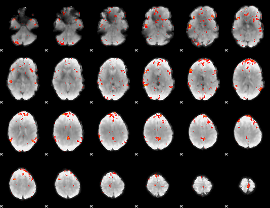 |
HAFNI result component: 253
Overlap rate: 0.39687
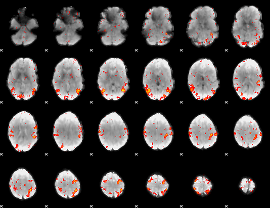 |
HAFNI result component: 011
Overlap rate: 0.2934
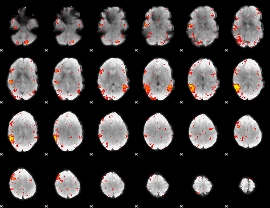 |
HAFNI result component: 031
Overlap rate: 0.33618
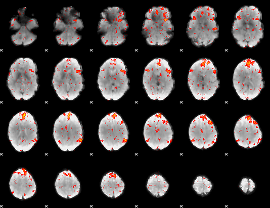 |
Correlation: 0.36141
  |
Correlation: 0.32164
 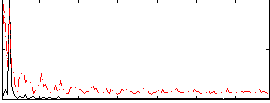 |
Correlation: 0.14099
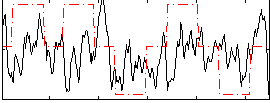 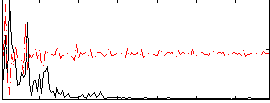 |
Subject: 4 |
GLM result cope: 01, threshold: 4
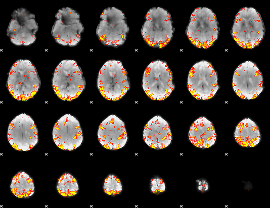 |
GLM result cope: 02, threshold: 4
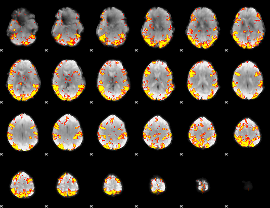 |
GLM result cope: 06, threshold: 3
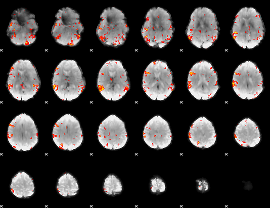 |
HAFNI result component: 037
Overlap rate: 0.34045
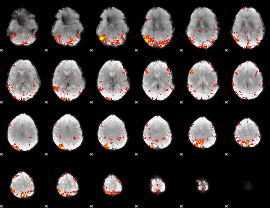 |
HAFNI result component: 037
Overlap rate: 0.29415
 |
HAFNI result component: 010
Overlap rate: 0.50925
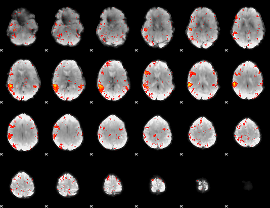 |
Correlation: -0.030963
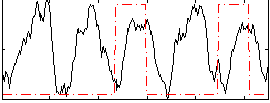 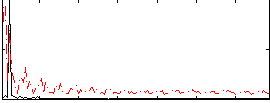 |
Correlation: 0.1964
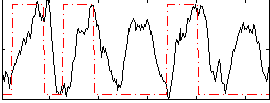 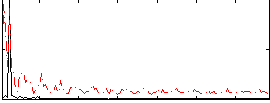 |
Correlation: 0.37817
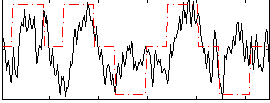 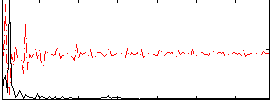 |
Subject: 5 |
GLM result cope: 01, threshold: 3.5
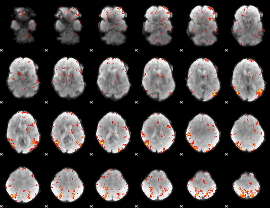 |
GLM result cope: 02, threshold: 3.5
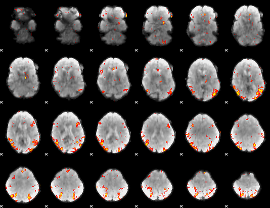 |
GLM result cope: 06, threshold: 2
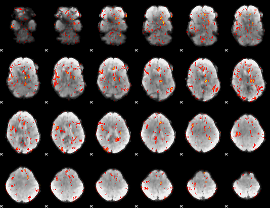 |
HAFNI result component: 199
Overlap rate: 0.43364
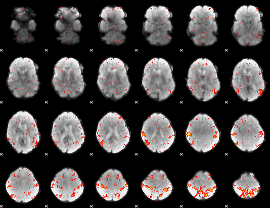 |
HAFNI result component: 380
Overlap rate: 0.45447
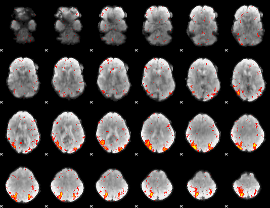 |
HAFNI result component: 136
Overlap rate: 0.33333
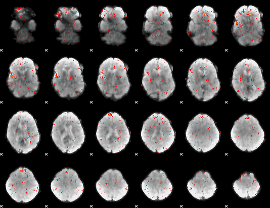 |
Correlation: 0.37023
  |
Correlation: 0.18888
  |
Correlation: 0.62964
  |
Subject: 6 |
GLM result cope: 01, threshold: 4
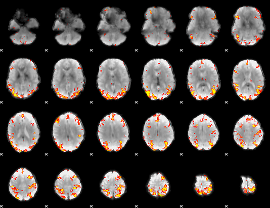 |
GLM result cope: 02, threshold: 4
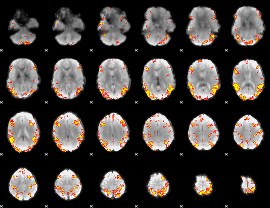 |
GLM result cope: 06, threshold: 3
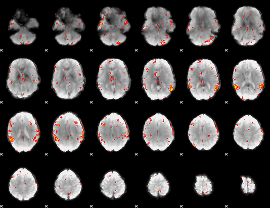 |
HAFNI result component: 101
Overlap rate: 0.36069
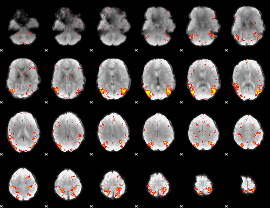 |
HAFNI result component: 101
Overlap rate: 0.34881
 |
HAFNI result component: 336
Overlap rate: 0.65848
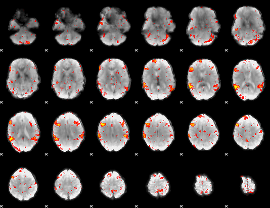 |
Correlation: 0.19758
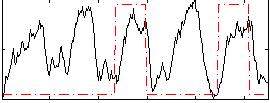  |
Correlation: 0.4396
  |
Correlation: 0.36287
  |
Subject: 7 |
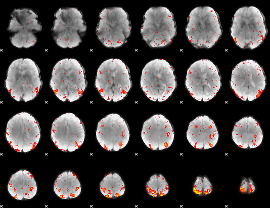
GLM result cope: 01, threshold: 4
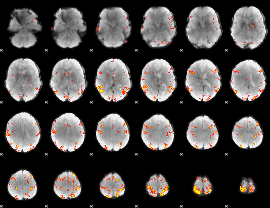 |
GLM result cope: 02, threshold: 4
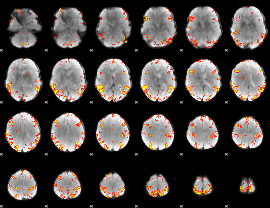 |
GLM result cope: 06, threshold: 2.5
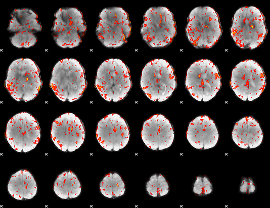 |
HAFNI result component: 275
Overlap rate: 0.40985
 |
HAFNI result component: 389
Overlap rate: 0.31482
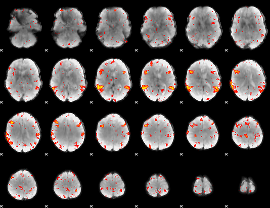 |
HAFNI result component: 112
Overlap rate: 0.2557
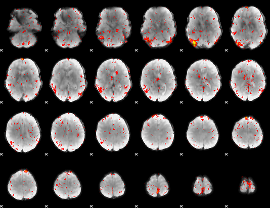 |
Correlation: 0.28862
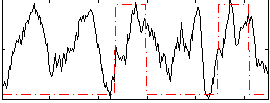  |
Correlation: 0.45549
  |
Correlation: 0.34602
  |
Subject: 8 |
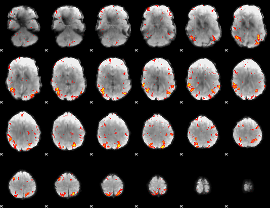
GLM result cope: 01, threshold: 2.5
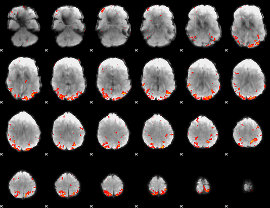 |
GLM result cope: 02, threshold: 4
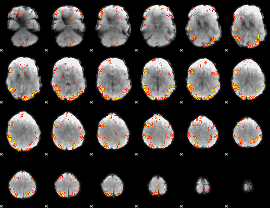 |
GLM result cope: 06, threshold: 2
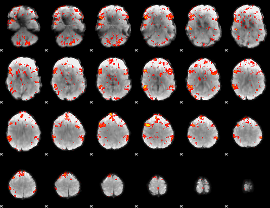 |
HAFNI result component: 244
Overlap rate: 0.63476
 |
HAFNI result component: 244
Overlap rate: 0.39577
 |
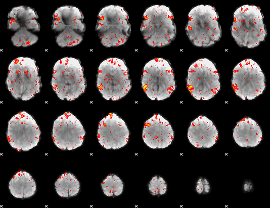
HAFNI result component: 005
Overlap rate: 0.711
 |
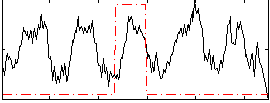
Correlation: 0.12033
  |
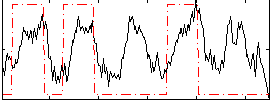
Correlation: 0.39885
  |
Correlation: 0.30737
  |
Subject: 9 |
GLM result cope: 01, threshold: 4
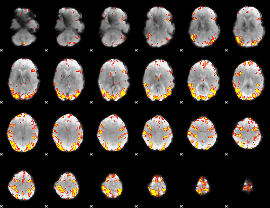 |
GLM result cope: 02, threshold: 4
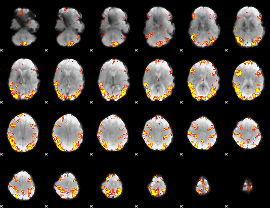 |
GLM result cope: 06, threshold: 1.5
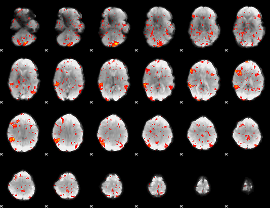 |
HAFNI result component: 384
Overlap rate: 0.32384
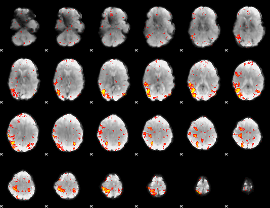 |
HAFNI result component: 400
Overlap rate: 0.30078
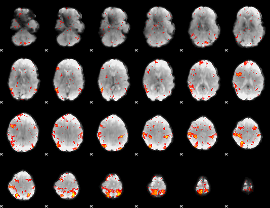 |
HAFNI result component: 295
Overlap rate: 0.85623
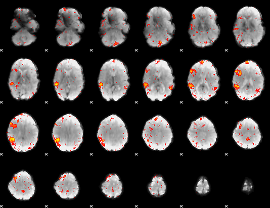 |
Correlation: 0.37239
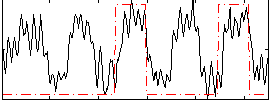 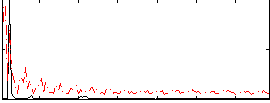 |
Correlation: 0.34543
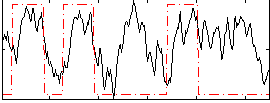  |
Correlation: 0.42245
  |
Subject: 10 |
GLM result cope: 01, threshold: 3
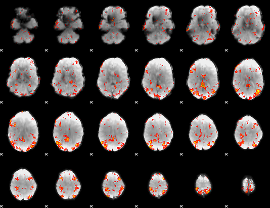 |
GLM result cope: 02, threshold: 4
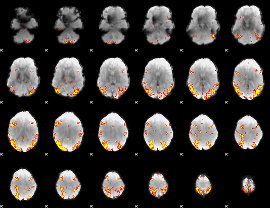 |
GLM result cope: 06, threshold: 2.5
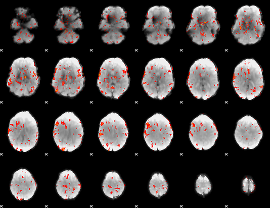 |
HAFNI result component: 380
Overlap rate: 0.40548
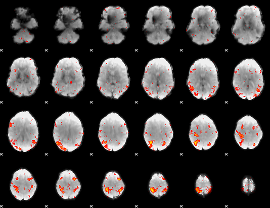 |
HAFNI result component: 322
Overlap rate: 0.39979
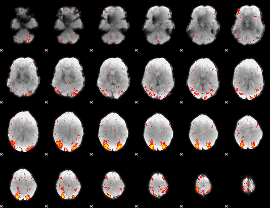 |
HAFNI result component: 163
Overlap rate: 0.35254
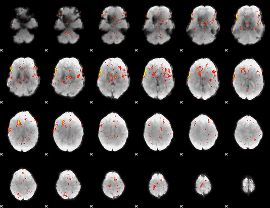 |
Correlation: -0.077176
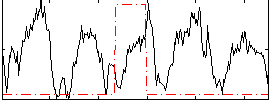  |
Correlation: 0.25788
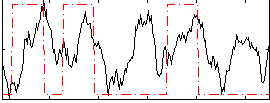 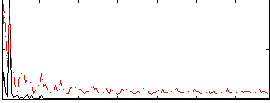 |
Correlation: 0.49753
  |
Subject: 11 |
GLM result cope: 01, threshold: 4
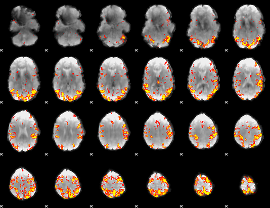 |
GLM result cope: 02, threshold: 3.5
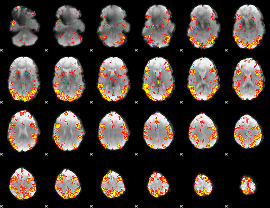 |
GLM result cope: 06, threshold: 1.5
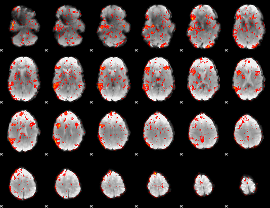 |
HAFNI result component: 334
Overlap rate: 0.39909
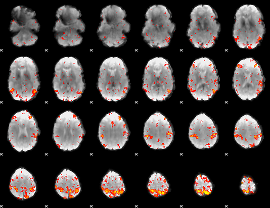 |
HAFNI result component: 334
Overlap rate: 0.31954
 |
HAFNI result component: 384
Overlap rate: 0.57363
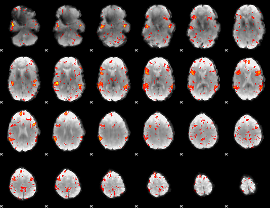 |
Correlation: 0.21035
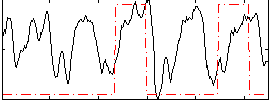  |
Correlation: 0.079741
  |
Correlation: 0.35914
  |
Subject: 12 |
GLM result cope: 01, threshold: 4
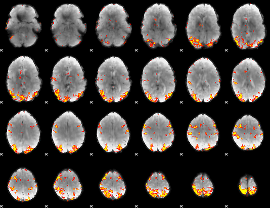 |
GLM result cope: 02, threshold: 4
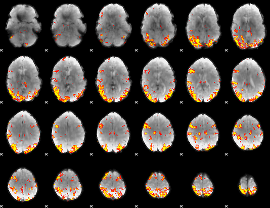 |
GLM result cope: 06, threshold: 3
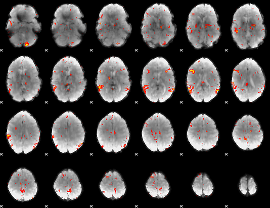 |
HAFNI result component: 382
Overlap rate: 0.44142
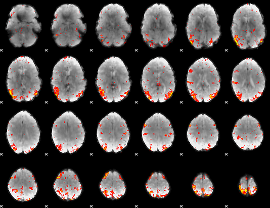 |
HAFNI result component: 382
Overlap rate: 0.40285
 |
HAFNI result component: 271
Overlap rate: 0.59112
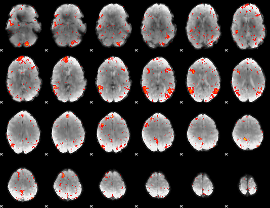 |
Correlation: 0.19011
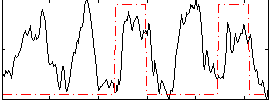 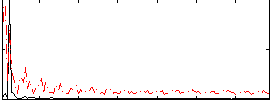 |
Correlation: 0.31998
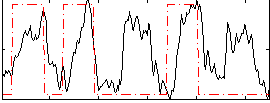 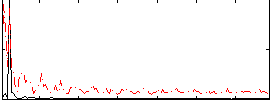 |
Correlation: 0.32905
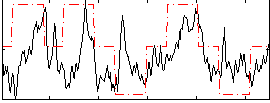 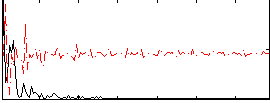 |
Subject: 13 |
GLM result cope: 01, threshold: 4
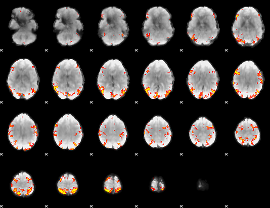 |
GLM result cope: 02, threshold: 4
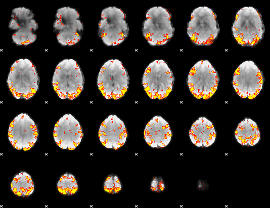 |
GLM result cope: 06, threshold: 2.5
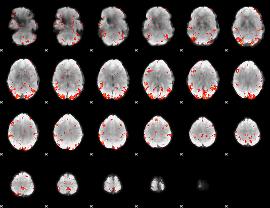 |
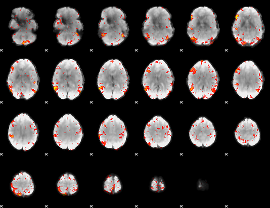
HAFNI result component: 237
Overlap rate: 0.48462
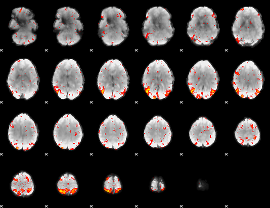 |
HAFNI result component: 280
Overlap rate: 0.31657
 |
HAFNI result component: 280
Overlap rate: 0.41859
 |
Correlation: 0.32953
  |
Correlation: 0.31004
 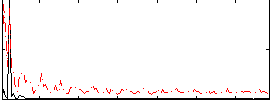 |
Correlation: 0.22417
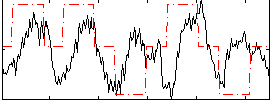 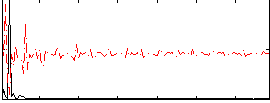 |
Subject: 14 |
GLM result cope: 01, threshold: 4
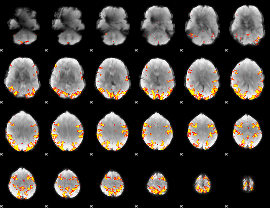 |
GLM result cope: 02, threshold: 4
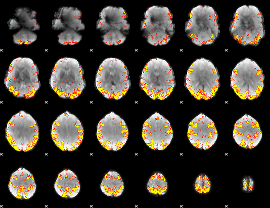 |
GLM result cope: 06, threshold: 3
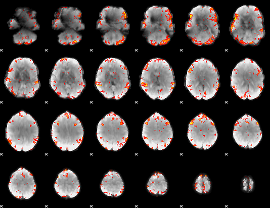 |
HAFNI result component: 325
Overlap rate: 0.45657
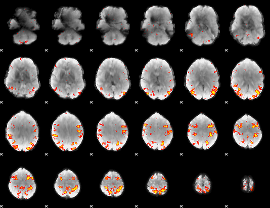 |
HAFNI result component: 375
Overlap rate: 0.33427
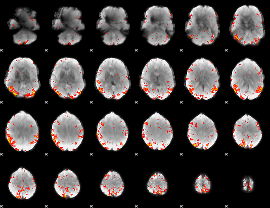 |
HAFNI result component: 215
Overlap rate: 0.37684
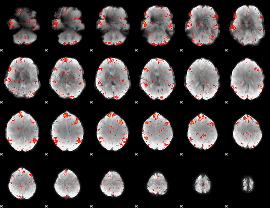 |
Correlation: 0.30257
  |
Correlation: 0.27637
  |
Correlation: 0.44685
  |
Subject: 15 |
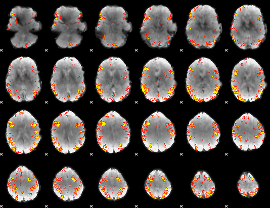
GLM result cope: 01, threshold: 3.5
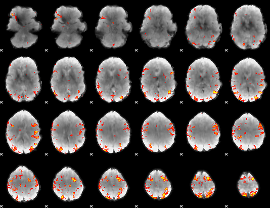 |
GLM result cope: 02, threshold: 4
 |
GLM result cope: 06, threshold: 2.5
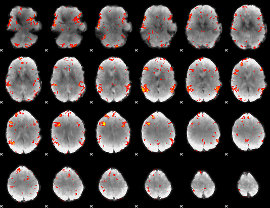 |
HAFNI result component: 061
Overlap rate: 0.48487
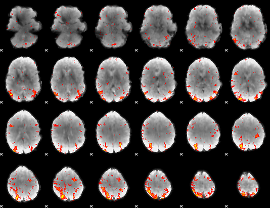 |
HAFNI result component: 061
Overlap rate: 0.32224
 |
HAFNI result component: 337
Overlap rate: 0.55621
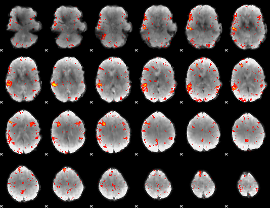 |
Correlation: 0.019049
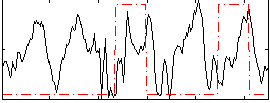  |
Correlation: 0.32668
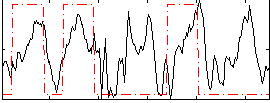  |
Correlation: 0.35429
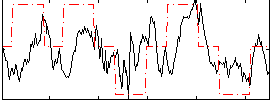 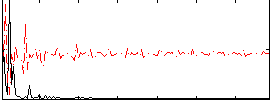 |
Subject: 16 |
GLM result cope: 01, threshold: 3.5
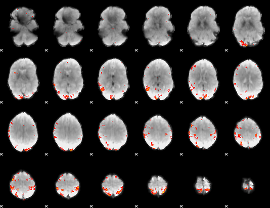 |
GLM result cope: 02, threshold: 4
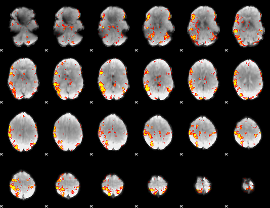 |
GLM result cope: 06, threshold: 2.5
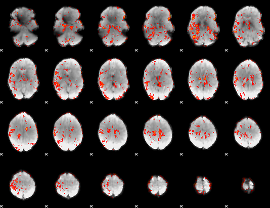 |
HAFNI result component: 386
Overlap rate: 0.48314
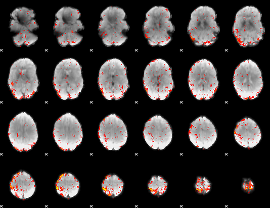 |
HAFNI result component: 220
Overlap rate: 0.31997
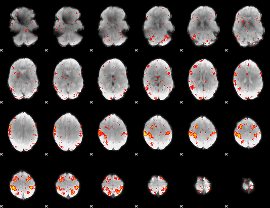 |
HAFNI result component: 185
Overlap rate: 0.22912
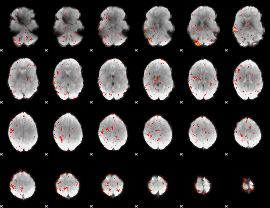 |
Correlation: 0.025948
  |
Correlation: 0.28875
  |
Correlation: 0.26834
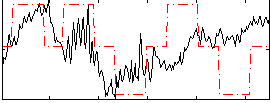  |
Subject: 17 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 06, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 18 |
GLM result cope: 01, threshold: 4
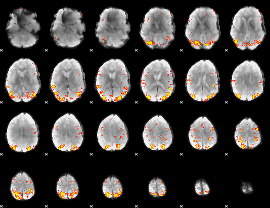 |
GLM result cope: 02, threshold: 3.5
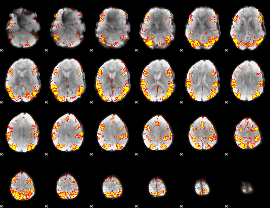 |
GLM result cope: 06, threshold: 3
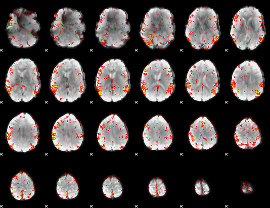 |
HAFNI result component: 340
Overlap rate: 0.43741
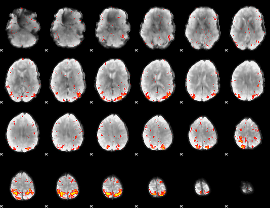 |
HAFNI result component: 253
Overlap rate: 0.25794
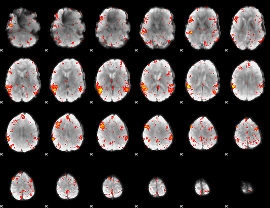 |
HAFNI result component: 253
Overlap rate: 0.42638
 |
Correlation: 0.38742
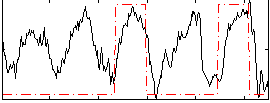  |
Correlation: 0.36175
  |
Correlation: 0.40233
  |
Subject: 19 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 06, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 20 |
GLM result cope: 01, threshold: 4
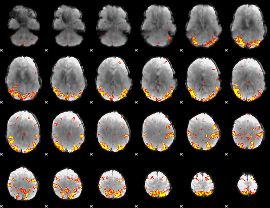 |
GLM result cope: 02, threshold: 4
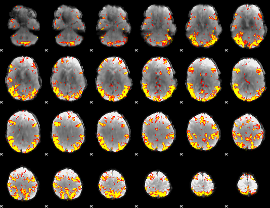 |
GLM result cope: 06, threshold: 3
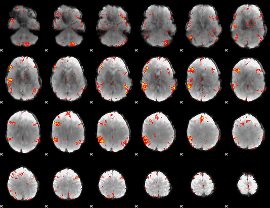 |
HAFNI result component: 163
Overlap rate: 0.45057
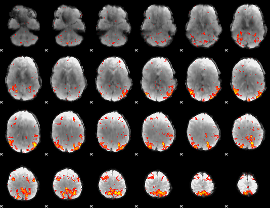 |
HAFNI result component: 163
Overlap rate: 0.26987
 |
HAFNI result component: 389
Overlap rate: 0.6004
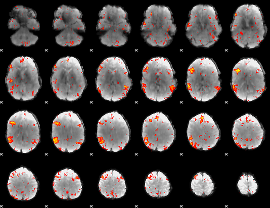 |
Correlation: 0.30218
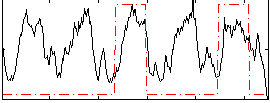 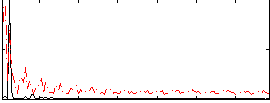 |
Correlation: 0.44065
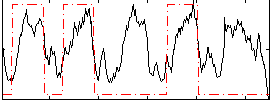 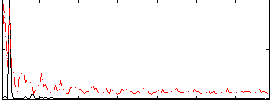 |
Correlation: 0.42598
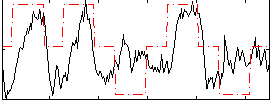 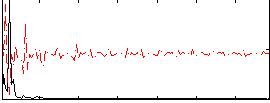 |
Subject: 21 |
GLM result cope: 01, threshold: 4
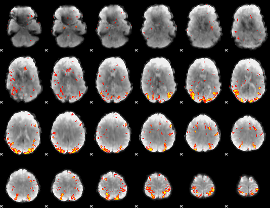 |
GLM result cope: 02, threshold: 4
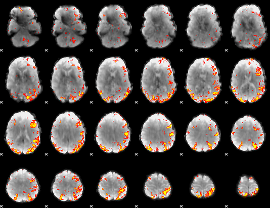 |
GLM result cope: 06, threshold: 2
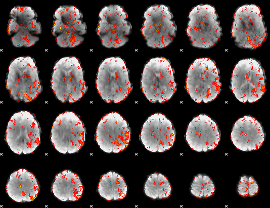 |
HAFNI result component: 269
Overlap rate: 0.37899
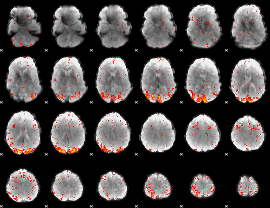 |
HAFNI result component: 125
Overlap rate: 0.36485
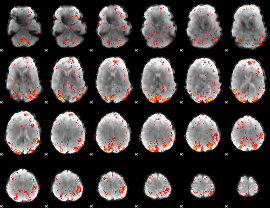 |
HAFNI result component: 157
Overlap rate: 0.2597
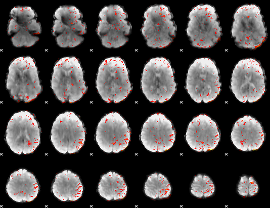 |
Correlation: 0.33099
  |
Correlation: 0.4197
  |
Correlation: 0.51369
  |
Subject: 22 |
GLM result cope: 01, threshold: 4
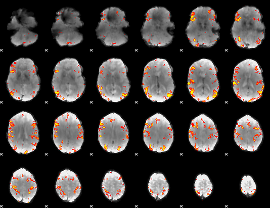 |
GLM result cope: 02, threshold: 4
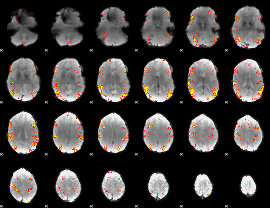 |
GLM result cope: 06, threshold: 1.5
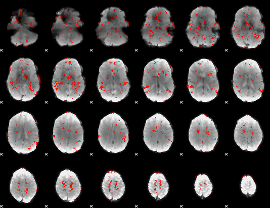 |
HAFNI result component: 293
Overlap rate: 0.39253
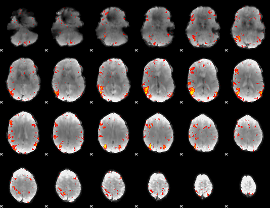 |
HAFNI result component: 293
Overlap rate: 0.4893
 |
HAFNI result component: 330
Overlap rate: 0.2459
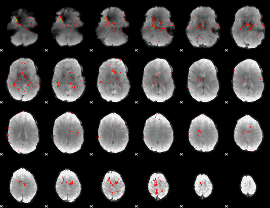 |
Correlation: 0.056429
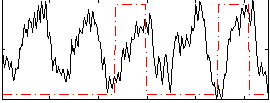  |
Correlation: 0.18681
  |
Correlation: 0.12155
  |
Subject: 23 |
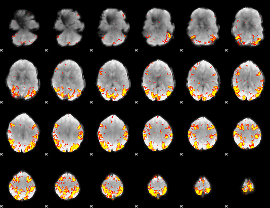
GLM result cope: 01, threshold: 4
 |
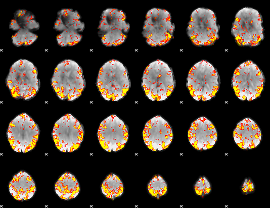
GLM result cope: 02, threshold: 4
 |
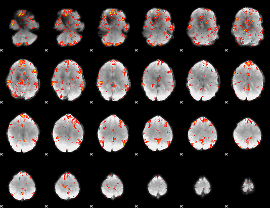
GLM result cope: 06, threshold: 3
 |
HAFNI result component: 398
Overlap rate: 0.36586
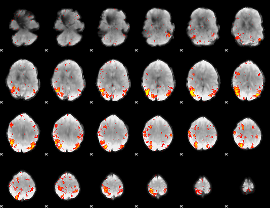 |
HAFNI result component: 025
Overlap rate: 0.28374
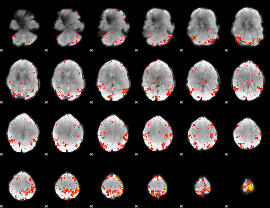 |
HAFNI result component: 213
Overlap rate: 0.38376
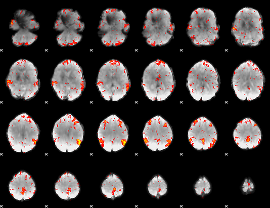 |
Correlation: 0.20084
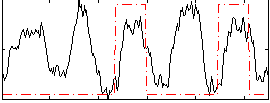  |
Correlation: 0.24123
  |
Correlation: 0.33996
  |
Subject: 24 |
GLM result cope: 01, threshold: 4
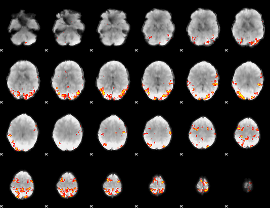 |
GLM result cope: 02, threshold: 4
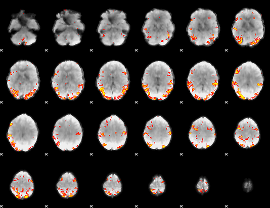 |
GLM result cope: 06, threshold: 2
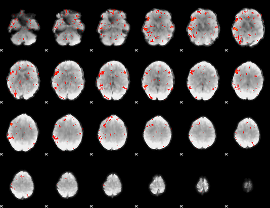 |
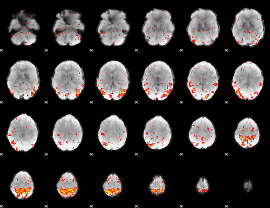
HAFNI result component: 255
Overlap rate: 0.5104
 |
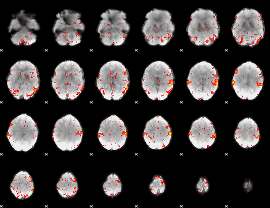
HAFNI result component: 201
Overlap rate: 0.36794
 |
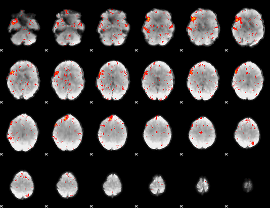
HAFNI result component: 282
Overlap rate: 0.76733
 |
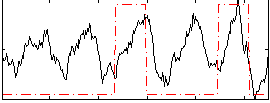
Correlation: 0.31957
  |
Correlation: 0.20888
  |
Correlation: 0.32631
  |
Subject: 25 |
GLM result cope: 01, threshold: 4
 |
GLM result cope: 02, threshold: 4
 |
GLM result cope: 06, threshold: 2
 |
HAFNI result component: 343
Overlap rate: 0.31806
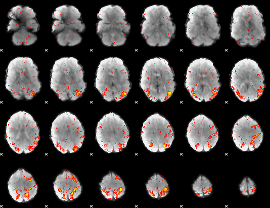 |
HAFNI result component: 239
Overlap rate: 0.34042
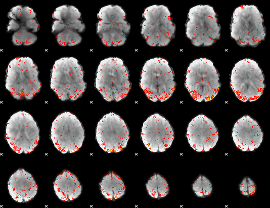 |
HAFNI result component: 384
Overlap rate: 0.24177
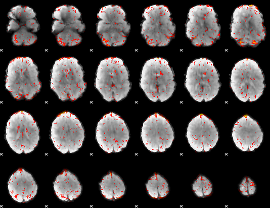 |
Correlation: 0.32453
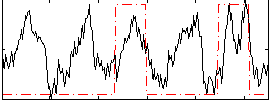  |
Correlation: 0.27165
  |
Correlation: 0.17493
  |
Subject: 26 |
GLM result cope: 01, threshold: 4
 |
GLM result cope: 02, threshold: 4
 |
GLM result cope: 06, threshold: 2.5
 |
HAFNI result component: 349
Overlap rate: 0.46483
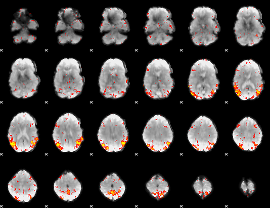 |
HAFNI result component: 261
Overlap rate: 0.27921
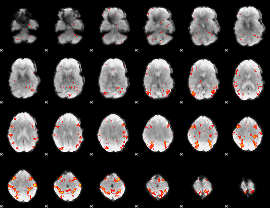 |
HAFNI result component: 202
Overlap rate: 0.43199
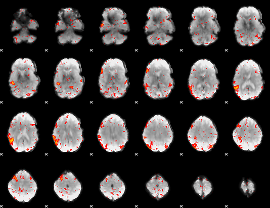 |
Correlation: 0.10544
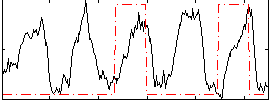  |
Correlation: 0.25395
  |
Correlation: 0.33483
  |
Subject: 27 |
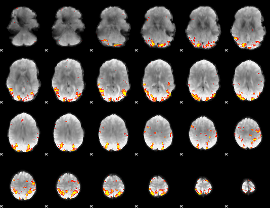
GLM result cope: 01, threshold: 4
 |
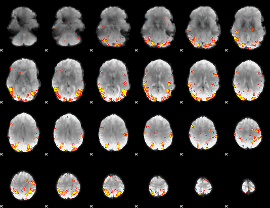
GLM result cope: 02, threshold: 4
 |
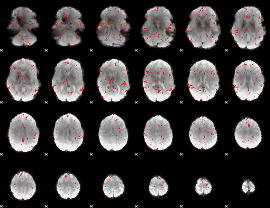
GLM result cope: 06, threshold: 2
 |
HAFNI result component: 312
Overlap rate: 0.51707
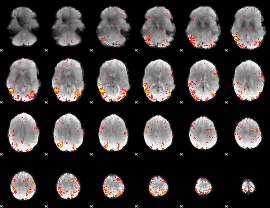 |
HAFNI result component: 312
Overlap rate: 0.44516
 |
HAFNI result component: 127
Overlap rate: 0.37931
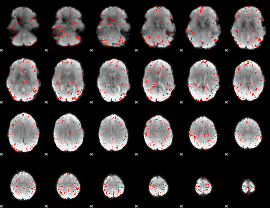 |
Correlation: 0.36171
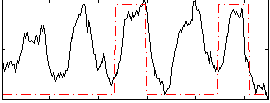 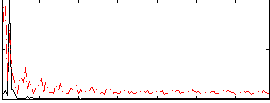 |
Correlation: 0.25988
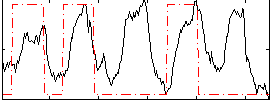 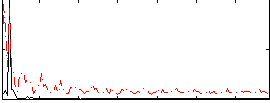 |
Correlation: 0.30022
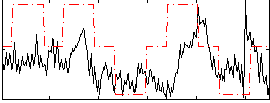 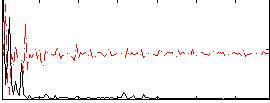 |
Subject: 28 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 06, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 29 |
GLM result cope: 01, threshold: 4
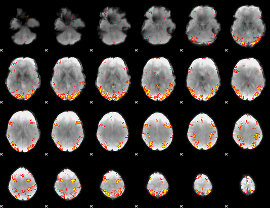 |
GLM result cope: 02, threshold: 4
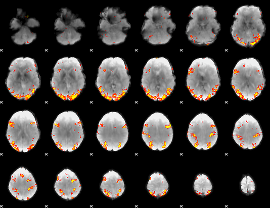 |
GLM result cope: 06, threshold: 1.5
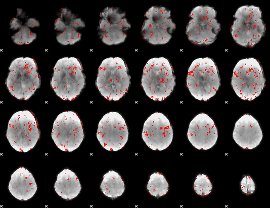 |
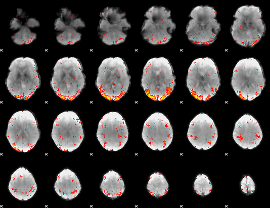
HAFNI result component: 400
Overlap rate: 0.31754
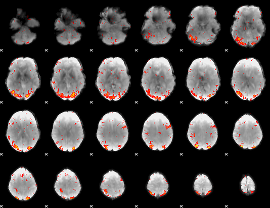 |
HAFNI result component: 203
Overlap rate: 0.36671
 |
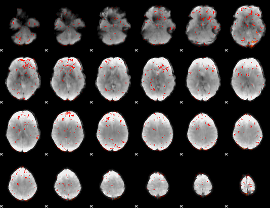
HAFNI result component: 358
Overlap rate: 0.38202
 |
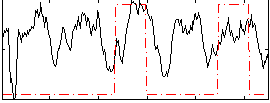
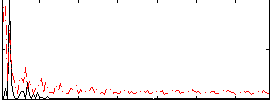
Correlation: 0.22826
  |
Correlation: 0.0068332
  |
Correlation: 0.22327
  |
Subject: 30 |
GLM result cope: 01, threshold: 2
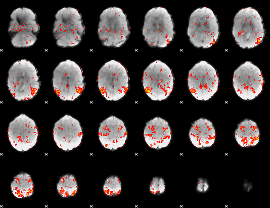 |
GLM result cope: 02, threshold: 3
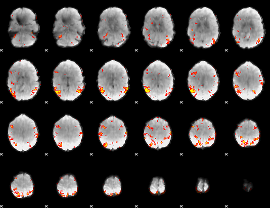 |
GLM result cope: 06, threshold: 2
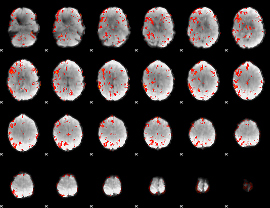 |
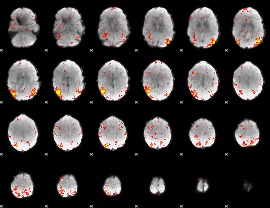
HAFNI result component: 280
Overlap rate: 0.58799
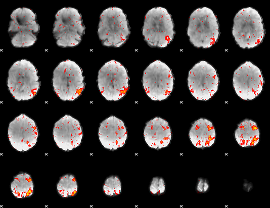 |
HAFNI result component: 247
Overlap rate: 0.54757
 |
HAFNI result component: 385
Overlap rate: 0.41486
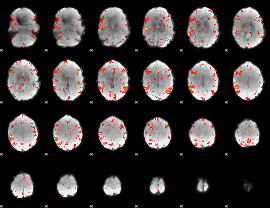 |
Correlation: 0.23073
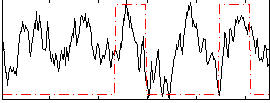 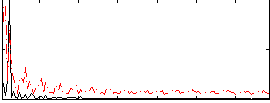 |
Correlation: 0.44634
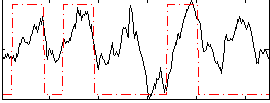 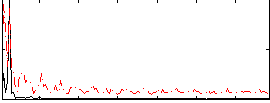 |
Correlation: 0.22099
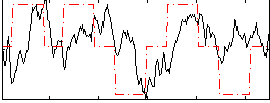 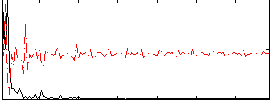 |
Subject: 31 |
GLM result cope: 01, threshold: 4
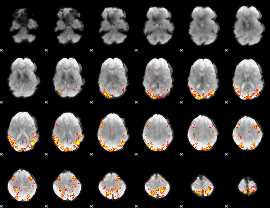 |
GLM result cope: 02, threshold: 4
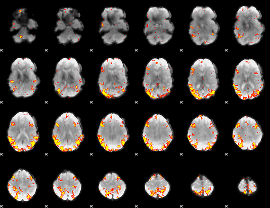 |
GLM result cope: 06, threshold: 2.5
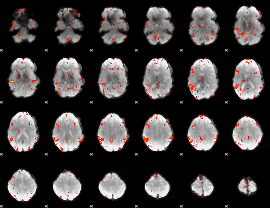 |
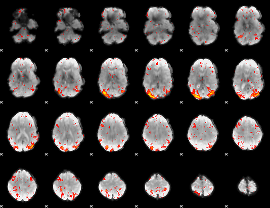
HAFNI result component: 238
Overlap rate: 0.55915
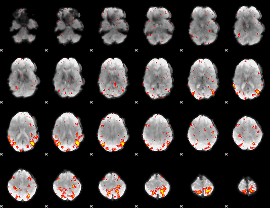 |
HAFNI result component: 304
Overlap rate: 0.35262
 |
HAFNI result component: 090
Overlap rate: 0.42607
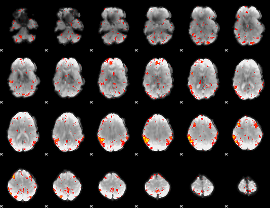 |
Correlation: 0.27512
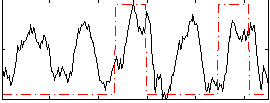 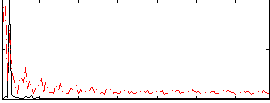 |
Correlation: 0.34784
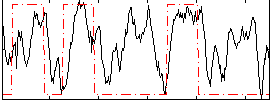 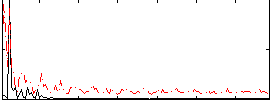 |
Correlation: 0.34085
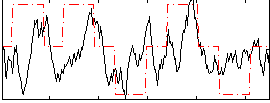 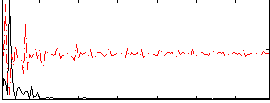 |
Subject: 32 |
GLM result cope: 01, threshold: 4
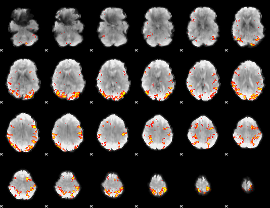 |
GLM result cope: 02, threshold: 4
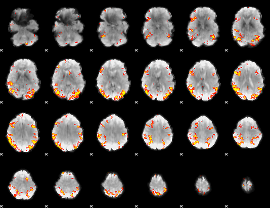 |
GLM result cope: 06, threshold: 2
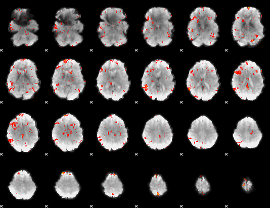 |
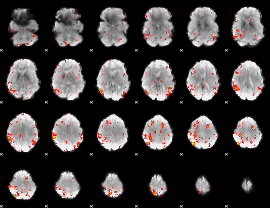
HAFNI result component: 356
Overlap rate: 0.4488
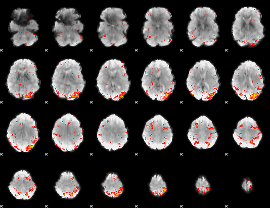 |
HAFNI result component: 073
Overlap rate: 0.34535
 |
HAFNI result component: 274
Overlap rate: 0.50661
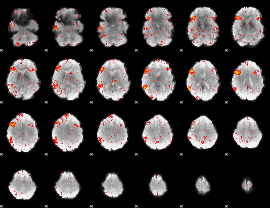 |
Correlation: 0.31481
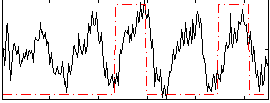 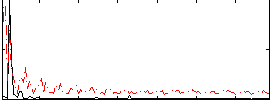 |
Correlation: 0.37045
  |
Correlation: 0.24419
  |
Subject: 33 |
GLM result cope: 01, threshold: 4
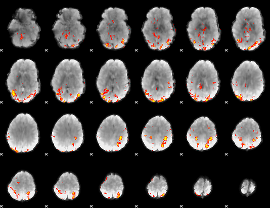 |
GLM result cope: 02, threshold: 4
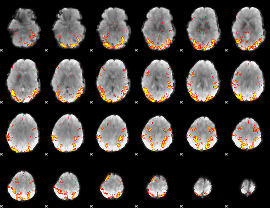 |
GLM result cope: 06, threshold: 3
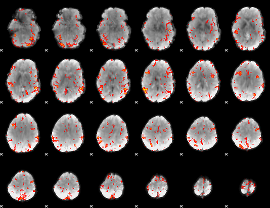 |
HAFNI result component: 378
Overlap rate: 0.50202
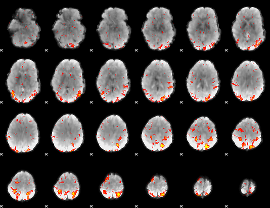 |
HAFNI result component: 378
Overlap rate: 0.36557
 |
HAFNI result component: 004
Overlap rate: 0.46994
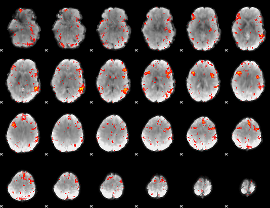 |
Correlation: 0.20018
  |
Correlation: 0.27775
 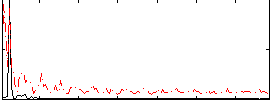 |
Correlation: 0.4495
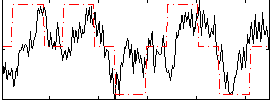 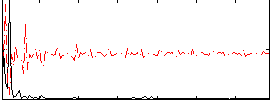 |
Subject: 34 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 06, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 35 |
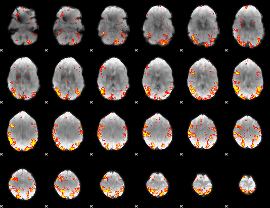
GLM result cope: 01, threshold: 4
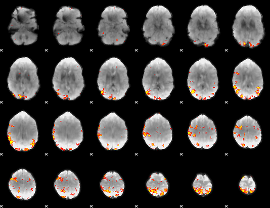 |
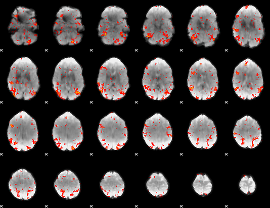
GLM result cope: 02, threshold: 3
 |
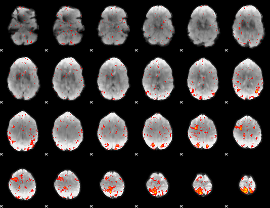
GLM result cope: 06, threshold: 2
 |
HAFNI result component: 071
Overlap rate: 0.42104
 |
HAFNI result component: 259
Overlap rate: 0.49151
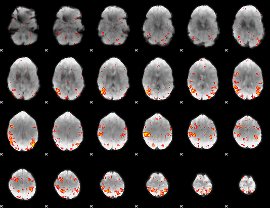 |
HAFNI result component: 162
Overlap rate: 0.64328
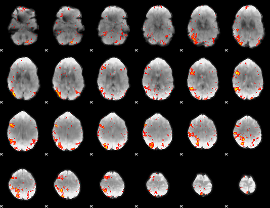 |
Correlation: 0.26482
  |
Correlation: 0.4733
 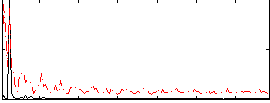 |
Correlation: 0.34171
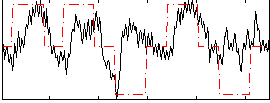 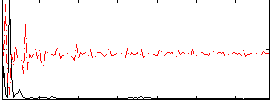 |
Subject: 36 |
GLM result cope: 01, threshold: 4
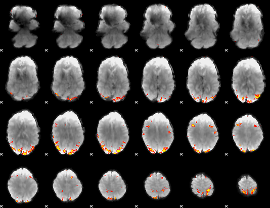 |
GLM result cope: 02, threshold: 3.5
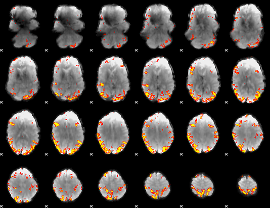 |
GLM result cope: 06, threshold: 3
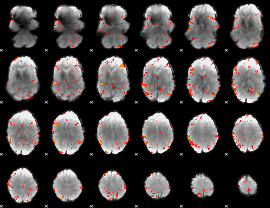 |
HAFNI result component: 312
Overlap rate: 0.62733
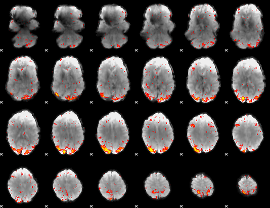 |
HAFNI result component: 069
Overlap rate: 0.41689
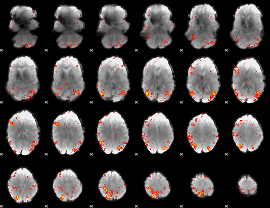 |
HAFNI result component: 369
Overlap rate: 0.54465
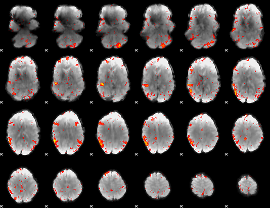 |
Correlation: 0.086234
  |
Correlation: 0.57821
 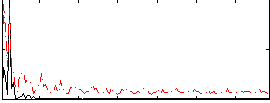 |
Correlation: 0.46495
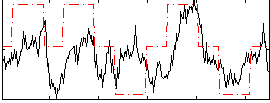 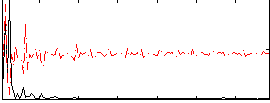 |
Subject: 37 |
GLM result cope: 01, threshold: 3.5
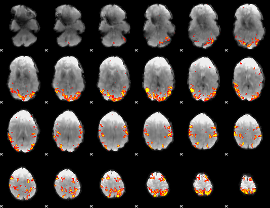 |
GLM result cope: 02, threshold: 4
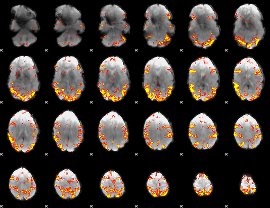 |
GLM result cope: 06, threshold: 3
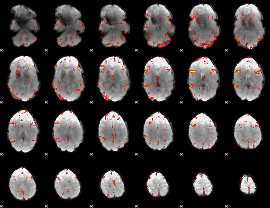 |
HAFNI result component: 163
Overlap rate: 0.46338
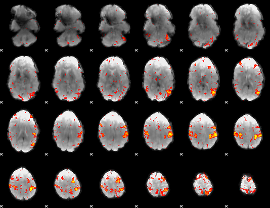 |
HAFNI result component: 356
Overlap rate: 0.27703
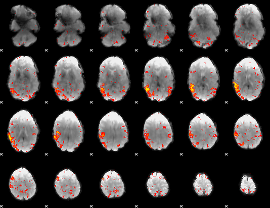 |
HAFNI result component: 213
Overlap rate: 0.48677
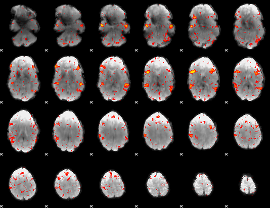 |
Correlation: 0.1789
  |
Correlation: 0.44247
  |
Correlation: 0.38696
  |
Subject: 38 |
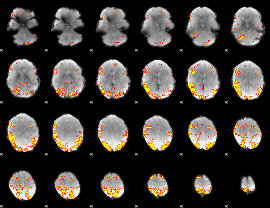
GLM result cope: 01, threshold: 4
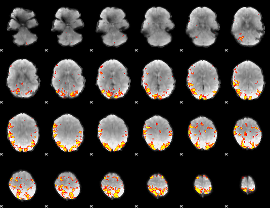 |
GLM result cope: 02, threshold: 4
 |
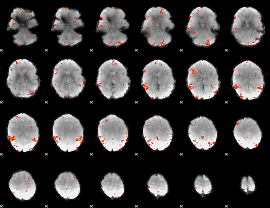
GLM result cope: 06, threshold: 3
 |
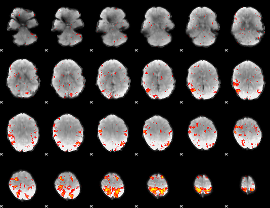
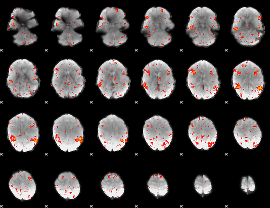
HAFNI result component: 145
Overlap rate: 0.43262
 |
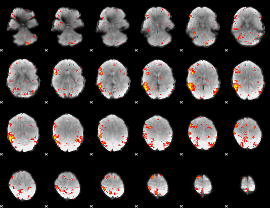
HAFNI result component: 243
Overlap rate: 0.38146
 |
HAFNI result component: 171
Overlap rate: 0.56338
 |
Correlation: 0.32454
  |
Correlation: 0.30604
  |
Correlation: 0.44589
  |
Subject: 39 |
GLM result cope: 01, threshold: 4
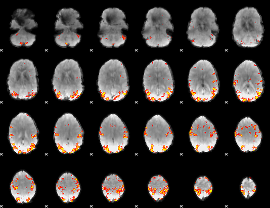 |
GLM result cope: 02, threshold: 4
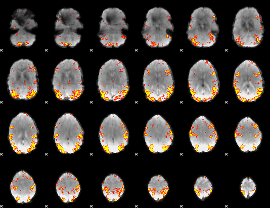 |
GLM result cope: 06, threshold: 2.5
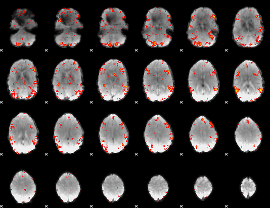 |
HAFNI result component: 243
Overlap rate: 0.38104
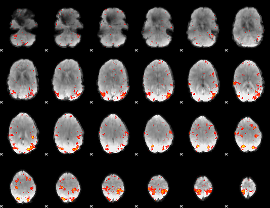 |
HAFNI result component: 367
Overlap rate: 0.23393
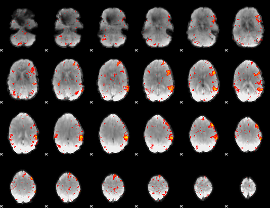 |
HAFNI result component: 367
Overlap rate: 0.44018
 |
Correlation: 0.30004
  |
Correlation: 0.36983
  |
Correlation: 0.34275
  |
Subject: 40 |
GLM result cope: 01, threshold: 3.5
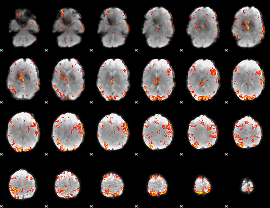 |
GLM result cope: 02, threshold: 3.5
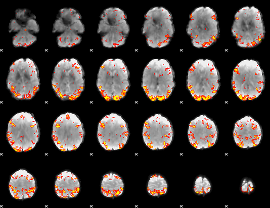 |
GLM result cope: 06, threshold: 3.5
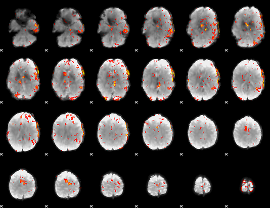 |
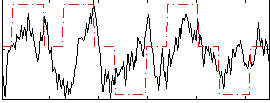
HAFNI result component: 203
Overlap rate: 0.27845
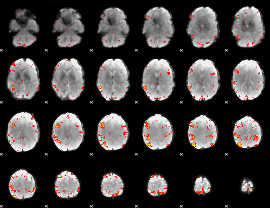 |
HAFNI result component: 203
Overlap rate: 0.35924
 |
HAFNI result component: 125
Overlap rate: 0.21595
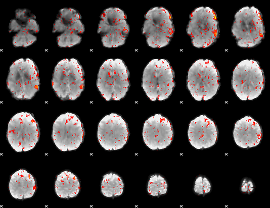 |
Correlation: 0.11703
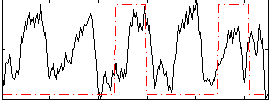 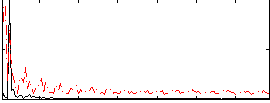 |
Correlation: 0.38917
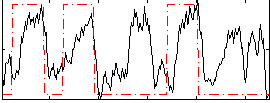 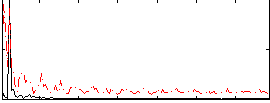 |
Correlation: 0.24807
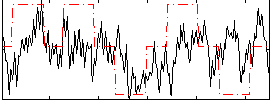  |
Subject: 41 |
GLM result cope: 01, threshold: 4
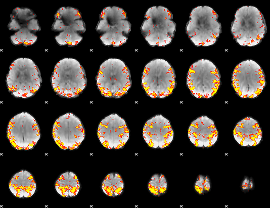 |
GLM result cope: 02, threshold: 4
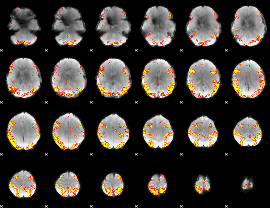 |
GLM result cope: 06, threshold: 1.5
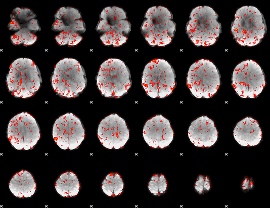 |
HAFNI result component: 180
Overlap rate: 0.24979
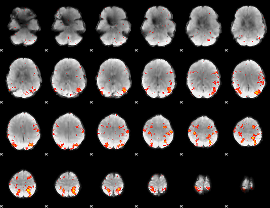 |
HAFNI result component: 274
Overlap rate: 0.23594
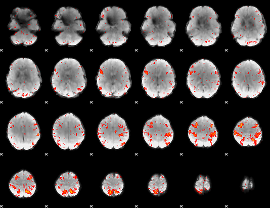 |
HAFNI result component: 398
Overlap rate: 0.49492
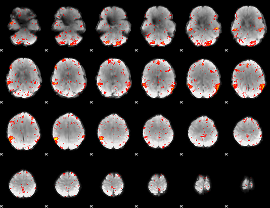 |
Correlation: 0.36566
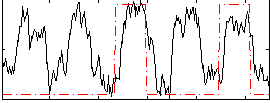  |
Correlation: 0.25148
  |
Correlation: 0.36133
  |
Subject: 42 |
GLM result cope: 01, threshold: 4
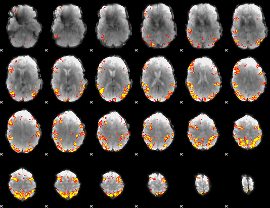 |
GLM result cope: 02, threshold: 4
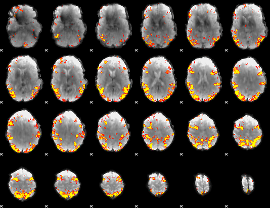 |
GLM result cope: 06, threshold: 3.5
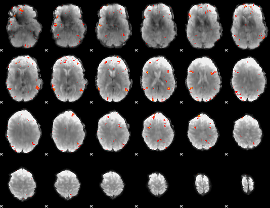 |
HAFNI result component: 168
Overlap rate: 0.38774
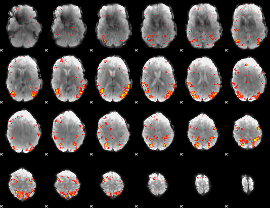 |
HAFNI result component: 168
Overlap rate: 0.32447
 |
HAFNI result component: 205
Overlap rate: 0.48228
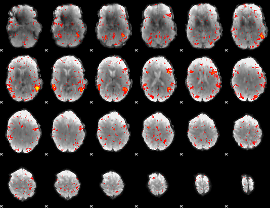 |
Correlation: 0.27268
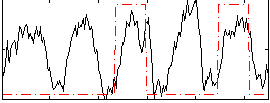  |
Correlation: 0.45376
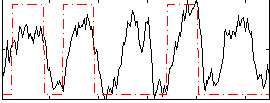 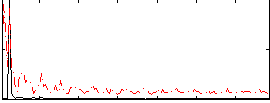 |
Correlation: 0.4432
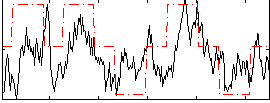 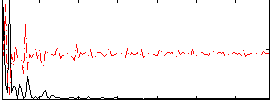 |
Subject: 43 |
GLM result cope: 01, threshold: 4
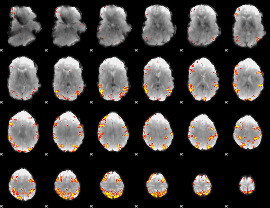 |
GLM result cope: 02, threshold: 4
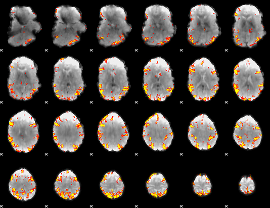 |
GLM result cope: 06, threshold: 2.5
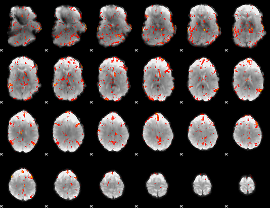 |
HAFNI result component: 378
Overlap rate: 0.548
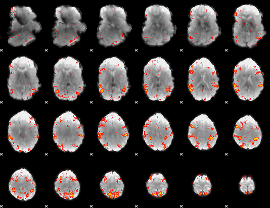 |
HAFNI result component: 378
Overlap rate: 0.38766
 |
HAFNI result component: 022
Overlap rate: 0.36659
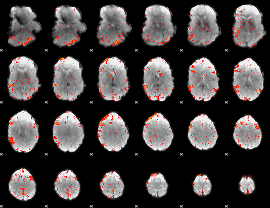 |
Correlation: 0.24038
  |
Correlation: 0.22138
  |
Correlation: 0.35014
  |
Subject: 44 |
GLM result cope: 01, threshold: 3
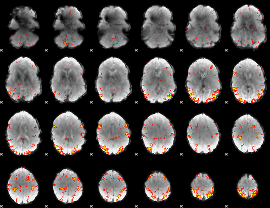 |
GLM result cope: 02, threshold: 4
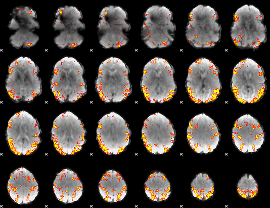 |
GLM result cope: 06, threshold: 4
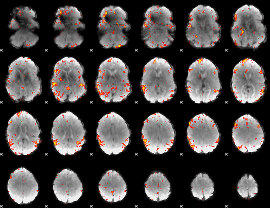 |
HAFNI result component: 348
Overlap rate: 0.59317
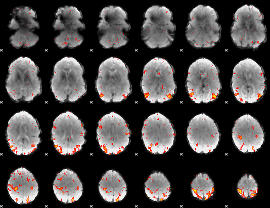 |
HAFNI result component: 358
Overlap rate: 0.25966
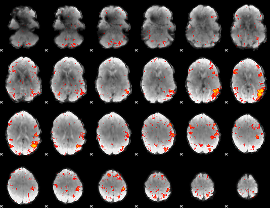 |
HAFNI result component: 317
Overlap rate: 0.37606
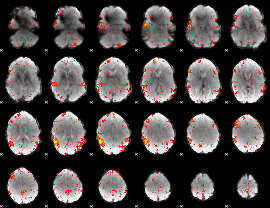 |
Correlation: 0.33977
  |
Correlation: 0.48755
  |
Correlation: 0.40974
  |
Subject: 45 |
GLM result cope: 01, threshold: 3.5
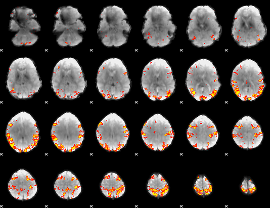 |
GLM result cope: 02, threshold: 4
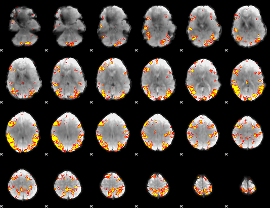 |
GLM result cope: 06, threshold: 3
 |
HAFNI result component: 295
Overlap rate: 0.44626
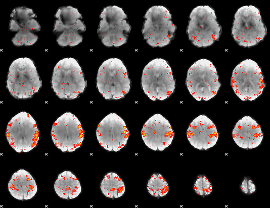 |
HAFNI result component: 255
Overlap rate: 0.34682
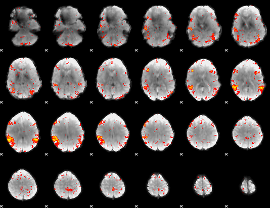 |
HAFNI result component: 224
Overlap rate: 0.51274
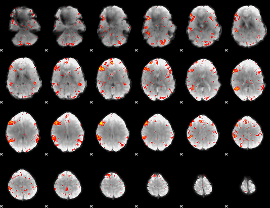 |
Correlation: 0.242
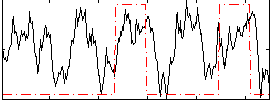 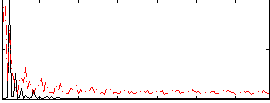 |
Correlation: 0.37237
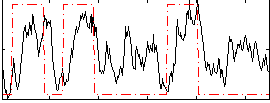 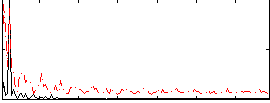 |
Correlation: 0.32404
  |
Subject: 46 |
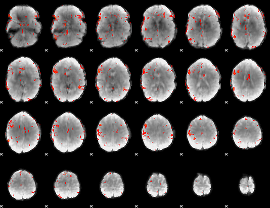
GLM result cope: 01, threshold: 3
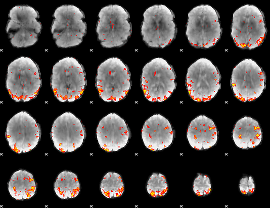 |
GLM result cope: 02, threshold: 3
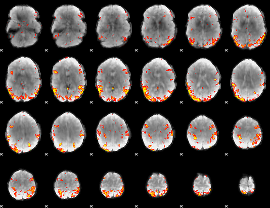 |
GLM result cope: 06, threshold: 2.5
 |
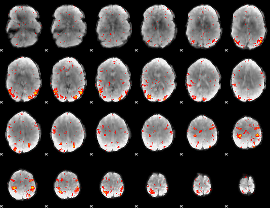
HAFNI result component: 039
Overlap rate: 0.47775
 |
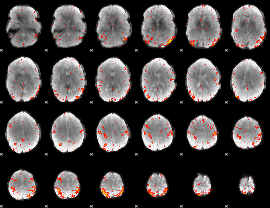
HAFNI result component: 223
Overlap rate: 0.48081
 |
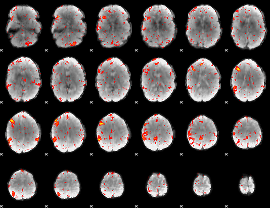
HAFNI result component: 317
Overlap rate: 0.33583
 |
Correlation: 0.27774
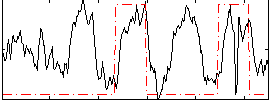 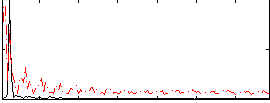 |
Correlation: 0.24161
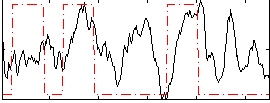 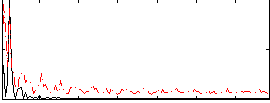 |
Correlation: 0.28133
  |
Subject: 47 |
GLM result cope: 01, threshold: 4
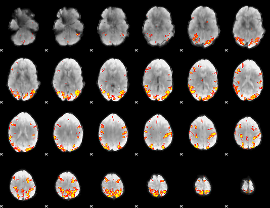 |
GLM result cope: 02, threshold: 4
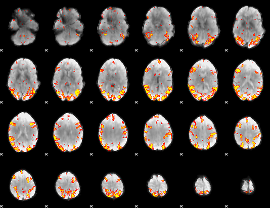 |
GLM result cope: 06, threshold: 2
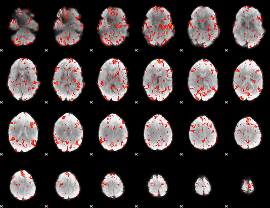 |
HAFNI result component: 226
Overlap rate: 0.37508
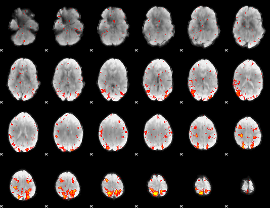 |
HAFNI result component: 226
Overlap rate: 0.23234
 |
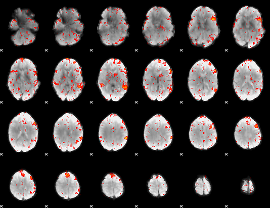
HAFNI result component: 375
Overlap rate: 0.31222
 |
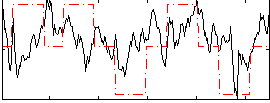
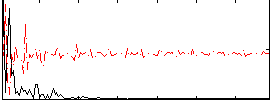
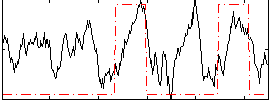
Correlation: 0.35704
  |
Correlation: 0.16997
  |
Correlation: 0.29573
  |
Subject: 48 |
GLM result cope: 01, threshold: 3.5
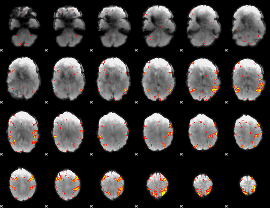 |
GLM result cope: 02, threshold: 3.5
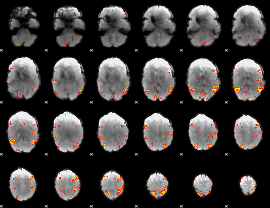 |
GLM result cope: 06, threshold: 2
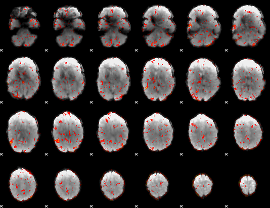 |
HAFNI result component: 189
Overlap rate: 0.61036
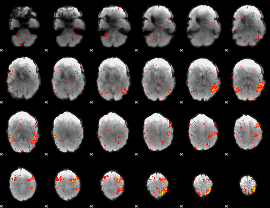 |
HAFNI result component: 189
Overlap rate: 0.36418
 |
HAFNI result component: 320
Overlap rate: 0.29246
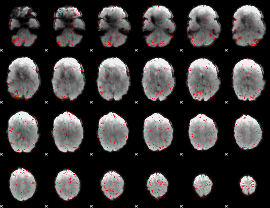 |
Correlation: 0.44278
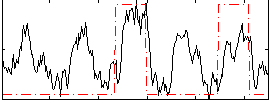 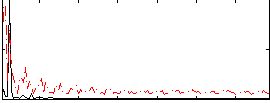 |
Correlation: 0.20061
  |
Correlation: 0.25007
  |
Subject: 49 |
GLM result cope: 01, threshold: 4
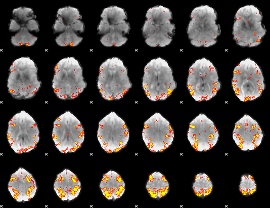 |
GLM result cope: 02, threshold: 4
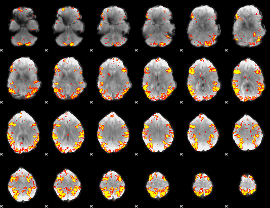 |
GLM result cope: 06, threshold: 3
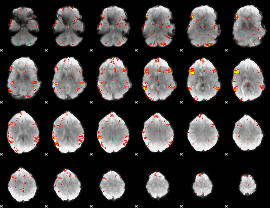 |
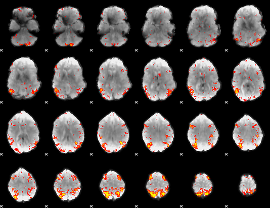
HAFNI result component: 329
Overlap rate: 0.40856
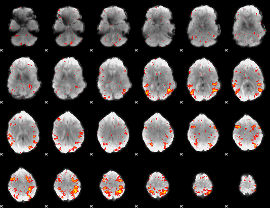 |
HAFNI result component: 310
Overlap rate: 0.35403
 |
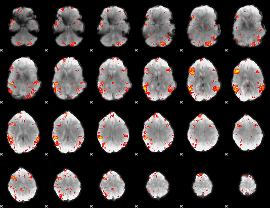
HAFNI result component: 325
Overlap rate: 0.7532
 |
Correlation: 0.3203
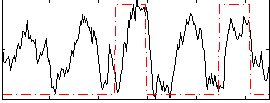  |
Correlation: 0.17449
  |
Correlation: 0.39083
  |
Subject: 50 |
GLM result cope: 01, threshold: 4
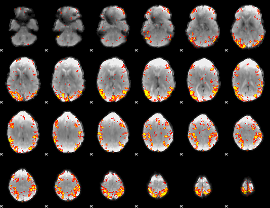 |
GLM result cope: 02, threshold: 4
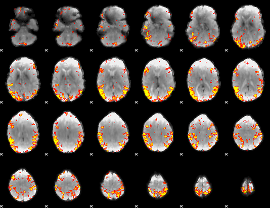 |
GLM result cope: 06, threshold: 2.5
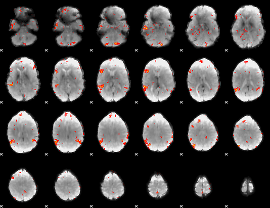 |
HAFNI result component: 077
Overlap rate: 0.25197
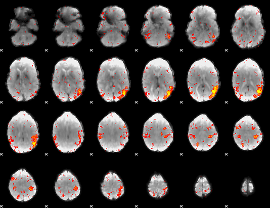 |
HAFNI result component: 077
Overlap rate: 0.26357
 |
HAFNI result component: 296
Overlap rate: 0.81781
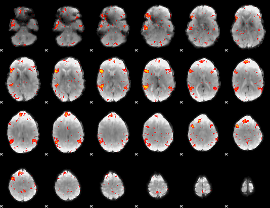 |
Correlation: 0.26039
  |
Correlation: 0.42891
  |
Correlation: 0.44395
  |
Subject: 51 |
GLM result cope: 01, threshold: 4
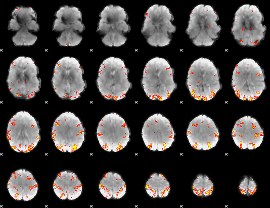 |
GLM result cope: 02, threshold: 3
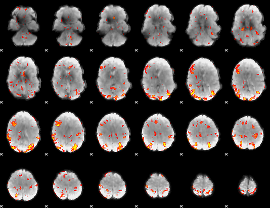 |
GLM result cope: 06, threshold: 1
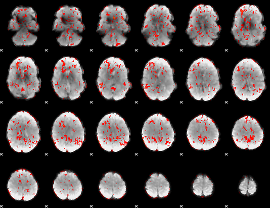 |
HAFNI result component: 127
Overlap rate: 0.55118
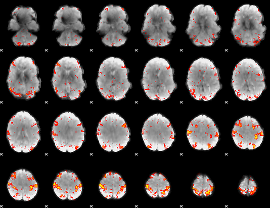 |
HAFNI result component: 008
Overlap rate: 0.46827
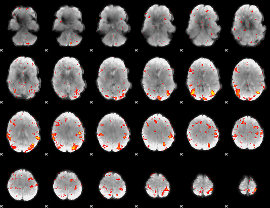 |
HAFNI result component: 367
Overlap rate: 0.52459
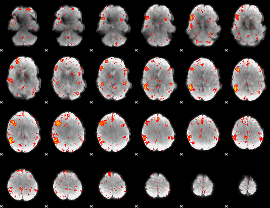 |
Correlation: 0.35178
  |
Correlation: 0.20552
 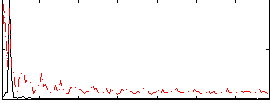 |
Correlation: 0.04069
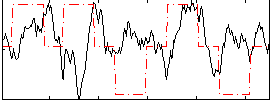 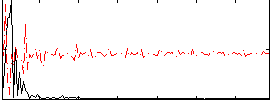 |
Subject: 52 |
GLM result cope: 01, threshold: 4
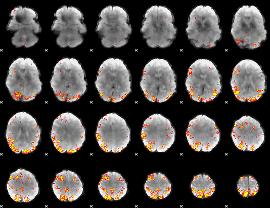 |
GLM result cope: 02, threshold: 3.5
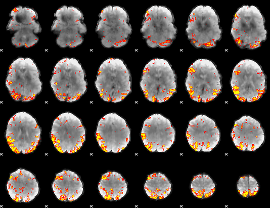 |
GLM result cope: 06, threshold: 2
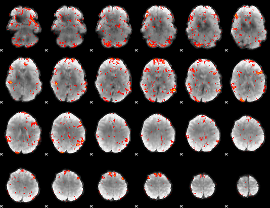 |
HAFNI result component: 125
Overlap rate: 0.50495
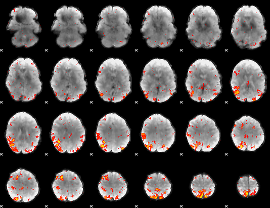 |
HAFNI result component: 321
Overlap rate: 0.32684
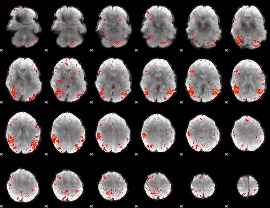 |
HAFNI result component: 133
Overlap rate: 0.5131
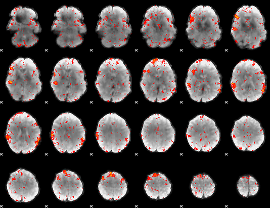 |
Correlation: 0.31702
  |
Correlation: 0.50804
  |
Correlation: 0.43198
  |
Subject: 53 |
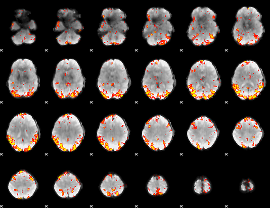
GLM result cope: 01, threshold: 3.5
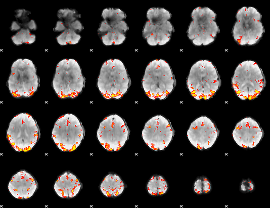 |
GLM result cope: 02, threshold: 3.5
 |
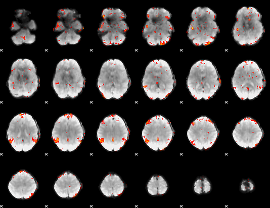
GLM result cope: 06, threshold: 3
 |
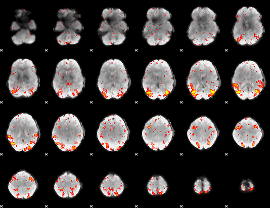
HAFNI result component: 396
Overlap rate: 0.51613
 |
HAFNI result component: 396
Overlap rate: 0.39532
 |
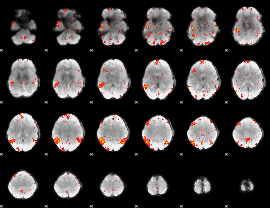
HAFNI result component: 293
Overlap rate: 0.44201
 |
Correlation: 0.12781
  |
Correlation: 0.42491
  |
Correlation: 0.33
  |
Subject: 54 |
GLM result cope: 01, threshold: 3.5
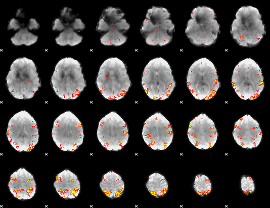 |
GLM result cope: 02, threshold: 4
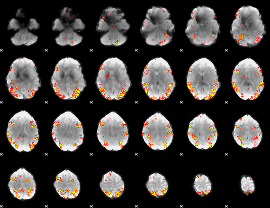 |
GLM result cope: 06, threshold: 2.5
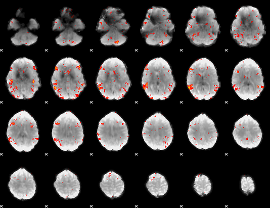 |
HAFNI result component: 339
Overlap rate: 0.56955
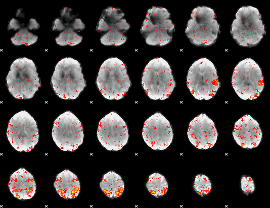 |
HAFNI result component: 327
Overlap rate: 0.42349
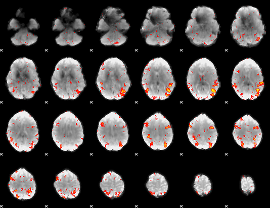 |
HAFNI result component: 099
Overlap rate: 0.83904
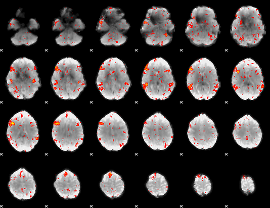 |
Correlation: 0.20678
  |
Correlation: 0.27234
  |
Correlation: 0.35412
  |
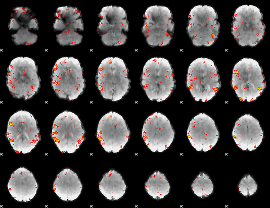
Subject: 55 |
GLM result cope: 01, threshold: 3.5
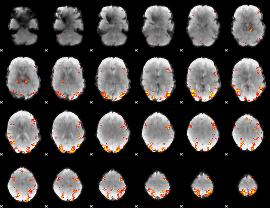 |
GLM result cope: 02, threshold: 3.5
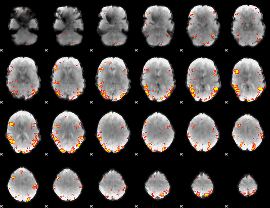 |
GLM result cope: 06, threshold: 3
 |
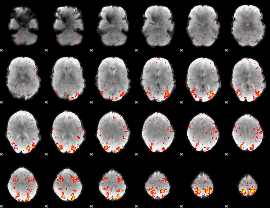
HAFNI result component: 297
Overlap rate: 0.50355
 |
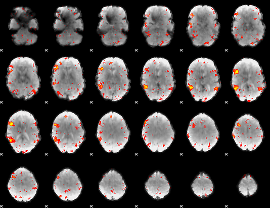
HAFNI result component: 278
Overlap rate: 0.44151
 |
HAFNI result component: 278
Overlap rate: 0.55413
 |
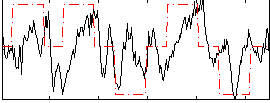
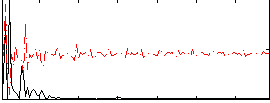
Correlation: 0.44968
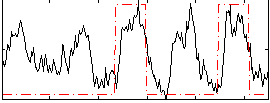  |
Correlation: 0.4135
  |
Correlation: 0.40601
  |
Subject: 56 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 06, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 57 |
GLM result cope: 01, threshold: 4
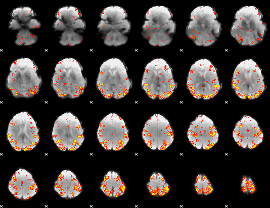 |
GLM result cope: 02, threshold: 4
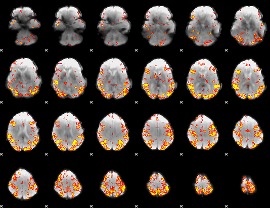 |
GLM result cope: 06, threshold: 2
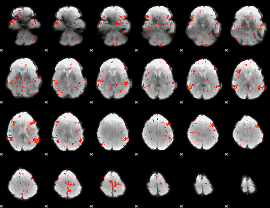 |
HAFNI result component: 261
Overlap rate: 0.31714
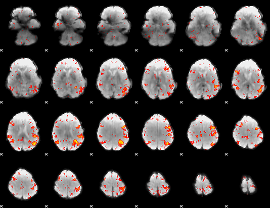 |
HAFNI result component: 261
Overlap rate: 0.26705
 |
HAFNI result component: 300
Overlap rate: 0.65132
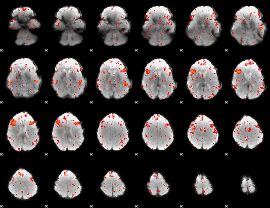 |
Correlation: 0.14685
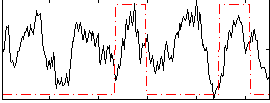 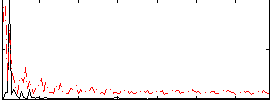 |
Correlation: 0.34264
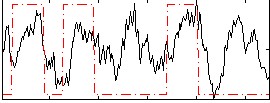 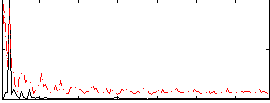 |
Correlation: 0.31315
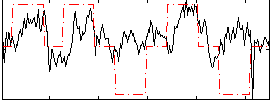 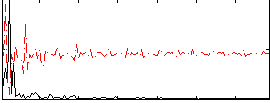 |
Subject: 58 |
GLM result cope: 01, threshold: 4
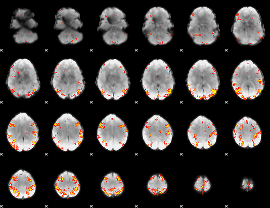 |
GLM result cope: 02, threshold: 3.5
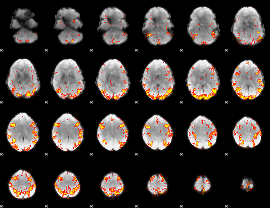 |
GLM result cope: 06, threshold: 3
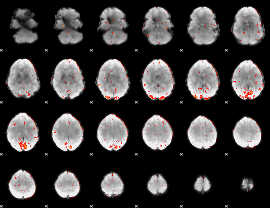 |
HAFNI result component: 273
Overlap rate: 0.24173
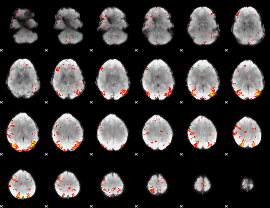 |
HAFNI result component: 350
Overlap rate: 0.32233
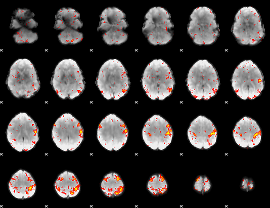 |
HAFNI result component: 058
Overlap rate: 0.32809
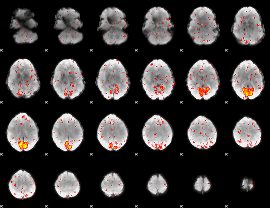 |
Correlation: 0.36981
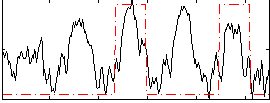 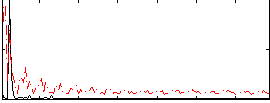 |
Correlation: 0.19376
  |
Correlation: 0.24893
  |
Subject: 59 |
GLM result cope: 01, threshold: 3
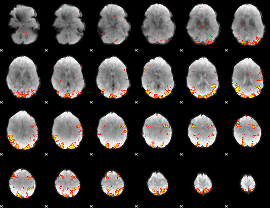 |
GLM result cope: 02, threshold: 4
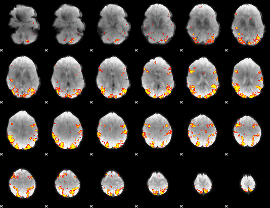 |
GLM result cope: 06, threshold: 2.5
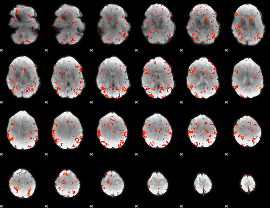 |
HAFNI result component: 054
Overlap rate: 0.35761
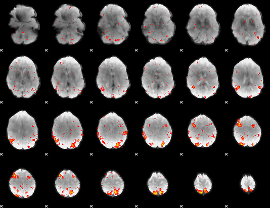 |
HAFNI result component: 309
Overlap rate: 0.26446
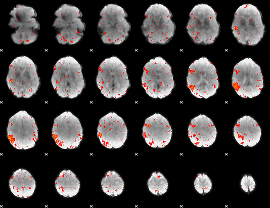 |
HAFNI result component: 400
Overlap rate: 0.49852
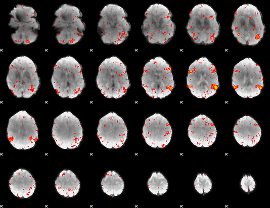 |
Correlation: 0.20694
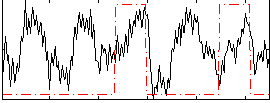 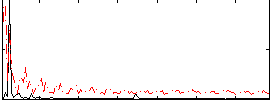 |
Correlation: 0.34361
  |
Correlation: 0.41444
  |
Subject: 60 |
GLM result cope: 01, threshold: 4
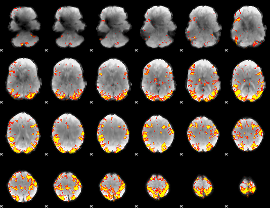 |
GLM result cope: 02, threshold: 4
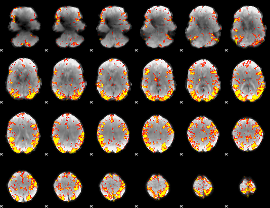 |
GLM result cope: 06, threshold: 3.5
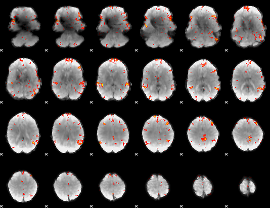 |
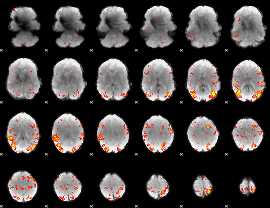
HAFNI result component: 286
Overlap rate: 0.38524
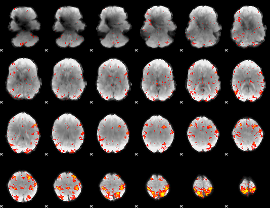 |
HAFNI result component: 363
Overlap rate: 0.21361
 |
HAFNI result component: 069
Overlap rate: 0.36507
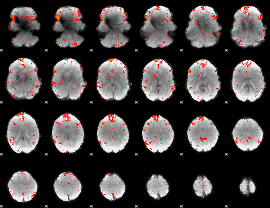 |
Correlation: 0.34451
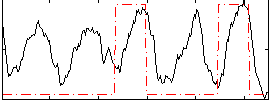 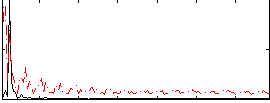 |
Correlation: 0.34162
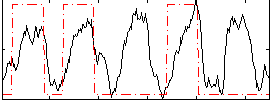 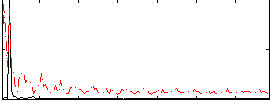 |
Correlation: 0.2665
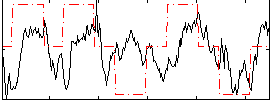 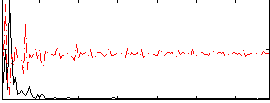 |
Subject: 61 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 06, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 62 |
GLM result cope: 01, threshold: 3.5
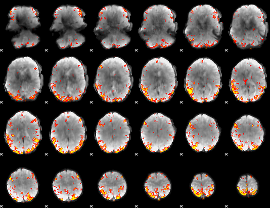 |
GLM result cope: 02, threshold: 3.5
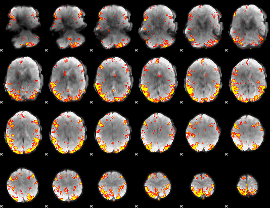 |
GLM result cope: 06, threshold: 2
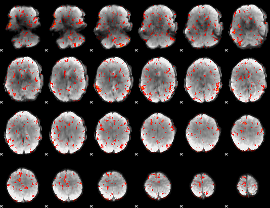 |
HAFNI result component: 220
Overlap rate: 0.39694
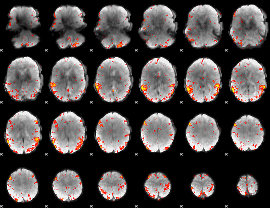 |
HAFNI result component: 220
Overlap rate: 0.42365
 |
HAFNI result component: 220
Overlap rate: 0.39659
 |
Correlation: -0.020305
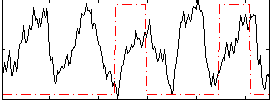  |
Correlation: 0.32583
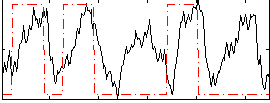  |
Correlation: 0.20646
  |
Subject: 63 |
GLM result cope: 01, threshold: 3.5
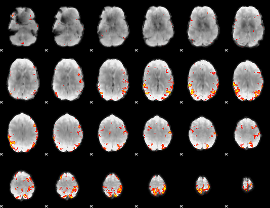 |
GLM result cope: 02, threshold: 4
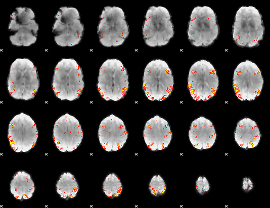 |
GLM result cope: 06, threshold: 2
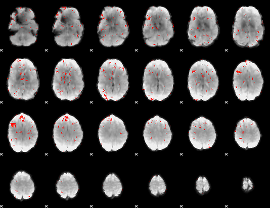 |
HAFNI result component: 174
Overlap rate: 0.43351
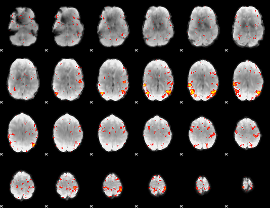 |
HAFNI result component: 148
Overlap rate: 0.47771
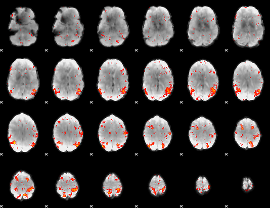 |
HAFNI result component: 321
Overlap rate: 0.57143
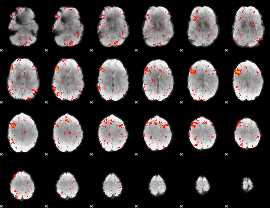 |
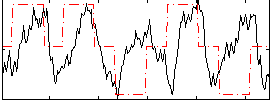
Correlation: 0.43023
  |
Correlation: 0.23642
  |
Correlation: -0.014666
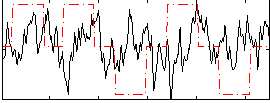 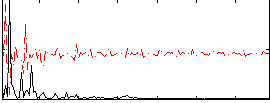 |
Subject: 64 |
GLM result cope: 01, threshold: 3
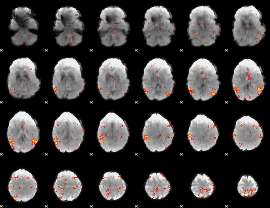 |
GLM result cope: 02, threshold: 4
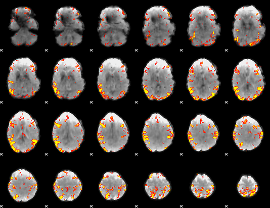 |
GLM result cope: 06, threshold: 4
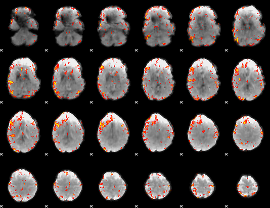 |
HAFNI result component: 328
Overlap rate: 0.60423
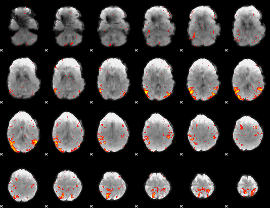 |
HAFNI result component: 328
Overlap rate: 0.29667
 |
HAFNI result component: 387
Overlap rate: 0.17162
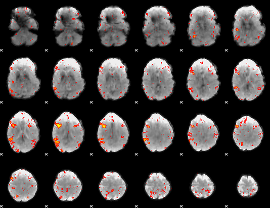 |
Correlation: 0.27267
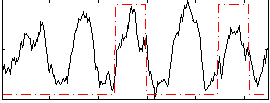  |
Correlation: 0.4669
  |
Correlation: 0.15931
  |
Subject: 65 |
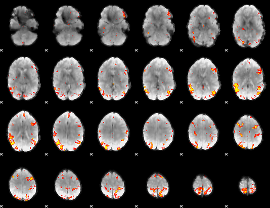
GLM result cope: 01, threshold: 4
 |
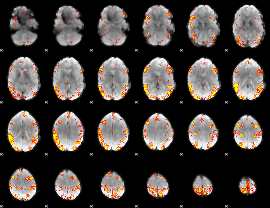
GLM result cope: 02, threshold: 4
 |
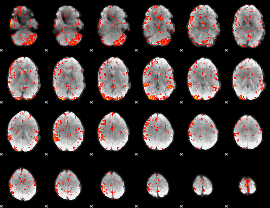
GLM result cope: 06, threshold: 2
 |
HAFNI result component: 359
Overlap rate: 0.44247
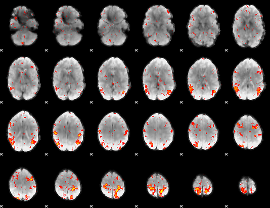 |
HAFNI result component: 383
Overlap rate: 0.29598
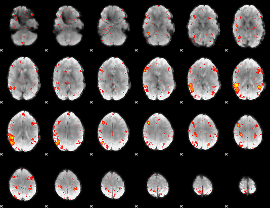 |
HAFNI result component: 352
Overlap rate: 0.42058
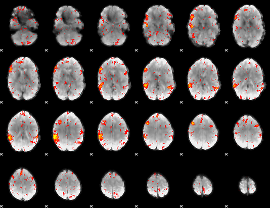 |
Correlation: 0.27792
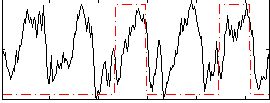  |
Correlation: 0.42574
  |
Correlation: 0.46101
  |
Subject: 66 |
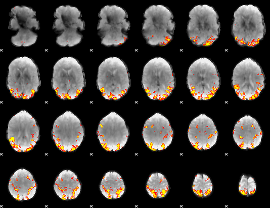
GLM result cope: 01, threshold: 4
 |
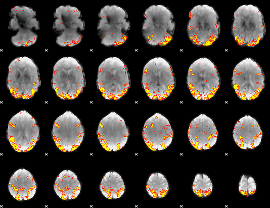
GLM result cope: 02, threshold: 4
 |
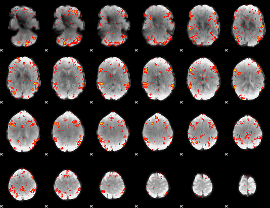
GLM result cope: 06, threshold: 2.5
 |
HAFNI result component: 352
Overlap rate: 0.51537
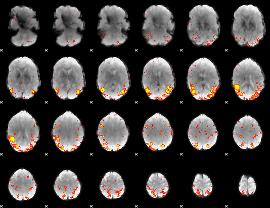 |
HAFNI result component: 352
Overlap rate: 0.36697
 |
HAFNI result component: 274
Overlap rate: 0.4492
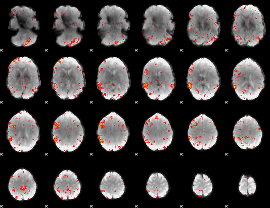 |
Correlation: 0.29709
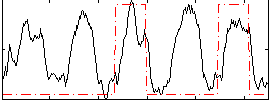  |
Correlation: 0.45197
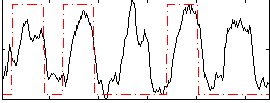 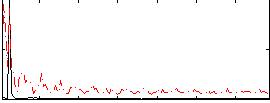 |
Correlation: 0.36791
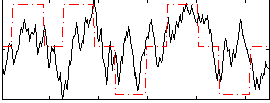 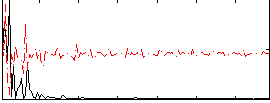 |
Subject: 67 |
GLM result cope: 01, threshold: 3.5
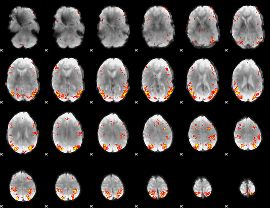 |
GLM result cope: 02, threshold: 4
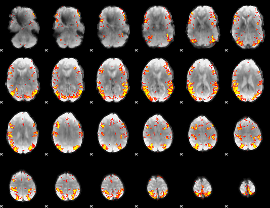 |
GLM result cope: 06, threshold: 2.5
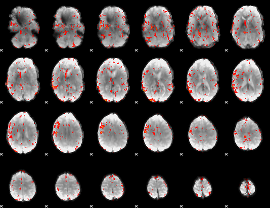 |
HAFNI result component: 320
Overlap rate: 0.50734
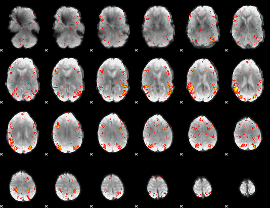 |
HAFNI result component: 320
Overlap rate: 0.35089
 |
HAFNI result component: 070
Overlap rate: 0.27024
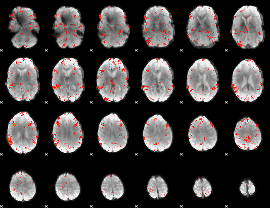 |
Correlation: 0.11796
  |
Correlation: 0.525
  |
Correlation: 0.4345
  |
Subject: 68 |
GLM result cope: 01, threshold: 4
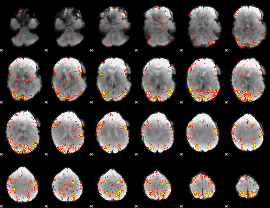 |
GLM result cope: 02, threshold: 4
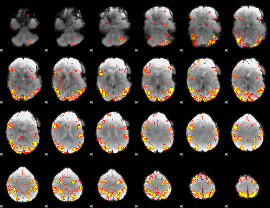 |
GLM result cope: 06, threshold: 2
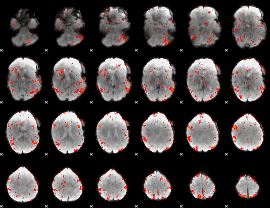 |
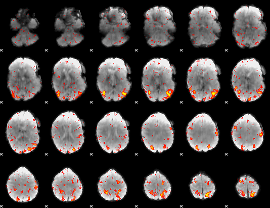
HAFNI result component: 382
Overlap rate: 0.35807
 |
HAFNI result component: 113
Overlap rate: 0.26414
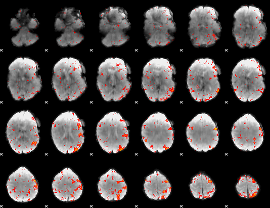 |
HAFNI result component: 113
Overlap rate: 0.35192
 |
Correlation: 0.18977
  |
Correlation: 0.36687
  |
Correlation: 0.36423
  |















































































































































































































































































































































































































































































































































































































































































































































































































































