Task name: RELATIONAL, TR=0.72, totally 232 volumes, 10 waves, 6 copes BACK
|
Cope: 01:MATCH BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
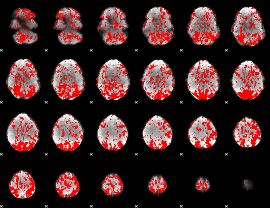 |
GLM binary result thresholded by 1.5
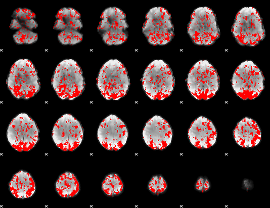 |
GLM binary result thresholded by 2
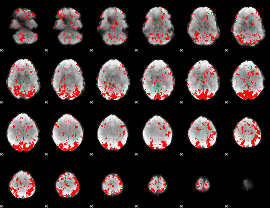 |
GLM binary result thresholded by 2.5
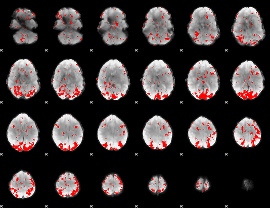 |
GLM binary result thresholded by 3
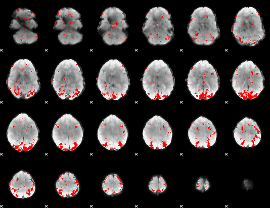 |
GLM binary result thresholded by 3.5
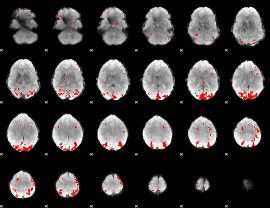 |
GLM binary result thresholded by 4
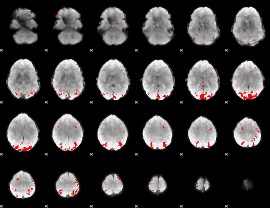 |
GLM Z-score result thresholded by 1
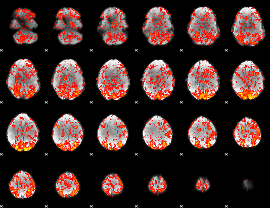 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:331; Jaccard:0.13122
 |
Component:331; Jaccard:0.16184
 |
Component:331; Jaccard:0.19585
 |
Component:331; Jaccard:0.2221
 |
Component:043; Jaccard:0.24212
 |
Component:203; Jaccard:0.23804
 |
Component:203; Jaccard:0.2145
 |
Correlation:0.04877
  |
Correlation:0.04877
  |
Correlation:0.04877
  |
Correlation:0.04877
  |
Correlation:0.11466
  |
Correlation:0.10881
  |
Correlation:0.10881
  |
Component:172; Jaccard:0.11964
 |
Component:172; Jaccard:0.14752
 |
Component:203; Jaccard:0.18244
 |
Component:043; Jaccard:0.2174
 |
Component:203; Jaccard:0.23459
 |
Component:043; Jaccard:0.23588
 |
Component:043; Jaccard:0.21251
 |
Correlation:0.15077
  |
Correlation:0.15077
  |
Correlation:0.10881
  |
Correlation:0.11466
  |
Correlation:0.10881
  |
Correlation:0.11466
  |
Correlation:0.11466
  |
Component:203; Jaccard:0.11492
 |
Component:203; Jaccard:0.14639
 |
Component:043; Jaccard:0.17961
 |
Component:203; Jaccard:0.21584
 |
Component:331; Jaccard:0.22914
 |
Component:331; Jaccard:0.21071
 |
Component:172; Jaccard:0.17921
 |
Correlation:0.10881
  |
Correlation:0.10881
  |
Correlation:0.11466
  |
Correlation:0.10881
  |
Correlation:0.04877
  |
Correlation:0.04877
  |
Correlation:0.15077
  |
Component:043; Jaccard:0.11027
 |
Component:043; Jaccard:0.14183
 |
Component:172; Jaccard:0.17679
 |
Component:172; Jaccard:0.19633
 |
Component:172; Jaccard:0.2051
 |
Component:172; Jaccard:0.19915
 |
Component:331; Jaccard:0.17661
 |
Correlation:0.11466
  |
Correlation:0.11466
  |
Correlation:0.15077
  |
Correlation:0.15077
  |
Correlation:0.15077
  |
Correlation:0.15077
  |
Correlation:0.04877
  |
Component:102; Jaccard:0.1034
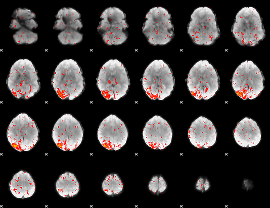 |
Component:102; Jaccard:0.11859
 |
Component:327; Jaccard:0.14142
 |
Component:327; Jaccard:0.1617
 |
Component:327; Jaccard:0.17253
 |
Component:327; Jaccard:0.1679
 |
Component:327; Jaccard:0.1489
 |
Correlation:0.14592
  |
Correlation:0.14592
  |
Correlation:0.11355
  |
Correlation:0.11355
  |
Correlation:0.11355
  |
Correlation:0.11355
  |
Correlation:0.11355
  |
Component:244; Jaccard:0.10297
 |
Component:327; Jaccard:0.11806
 |
Component:086; Jaccard:0.13417
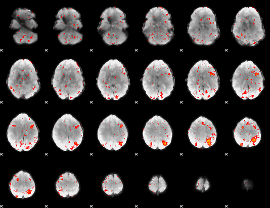 |
Component:020; Jaccard:0.14606
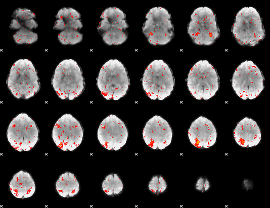 |
Component:086; Jaccard:0.14698
 |
Component:020; Jaccard:0.13148
 |
Component:086; Jaccard:0.11072
 |
Correlation:-0.086121
  |
Correlation:0.11355
  |
Correlation:0.14466
  |
Correlation:0.11124
  |
Correlation:0.14466
  |
Correlation:0.11124
  |
Correlation:0.14466
  |
Component:115; Jaccard:0.10023
 |
Component:086; Jaccard:0.11401
 |
Component:020; Jaccard:0.13328
 |
Component:086; Jaccard:0.14485
 |
Component:020; Jaccard:0.1431
 |
Component:086; Jaccard:0.13145
 |
Component:020; Jaccard:0.11009
 |
Correlation:-0.11178
  |
Correlation:0.14466
  |
Correlation:0.11124
  |
Correlation:0.14466
  |
Correlation:0.11124
  |
Correlation:0.14466
  |
Correlation:0.11124
  |
Component:327; Jaccard:0.097116
 |
Component:020; Jaccard:0.11385
 |
Component:102; Jaccard:0.13171
 |
Component:102; Jaccard:0.13722
 |
Component:102; Jaccard:0.13037
 |
Component:102; Jaccard:0.11674
 |
Component:102; Jaccard:0.098444
 |
Correlation:0.11355
  |
Correlation:0.11124
  |
Correlation:0.14592
  |
Correlation:0.14592
  |
Correlation:0.14592
  |
Correlation:0.14592
  |
Correlation:0.14592
  |
Component:355; Jaccard:0.096125
 |
Component:244; Jaccard:0.11092
 |
Component:244; Jaccard:0.11748
 |
Component:244; Jaccard:0.1154
 |
Component:349; Jaccard:0.11091
 |
Component:263; Jaccard:0.10104
 |
Component:263; Jaccard:0.085521
 |
Correlation:-0.23712
  |
Correlation:-0.086121
  |
Correlation:-0.086121
  |
Correlation:-0.086121
  |
Correlation:-0.11914
  |
Correlation:0.094294
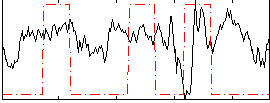 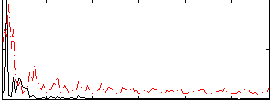 |
Correlation:0.094294
  |
Component:020; Jaccard:0.09525
 |
Component:115; Jaccard:0.10871
 |
Component:349; Jaccard:0.11071
 |
Component:349; Jaccard:0.11346
 |
Component:263; Jaccard:0.11059
 |
Component:349; Jaccard:0.1001
 |
Component:349; Jaccard:0.082736
 |
Correlation:0.11124
  |
Correlation:-0.11178
  |
Correlation:-0.11914
  |
Correlation:-0.11914
  |
Correlation:0.094294
  |
Correlation:-0.11914
  |
Correlation:-0.11914
  |
Cope: 02:REL BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
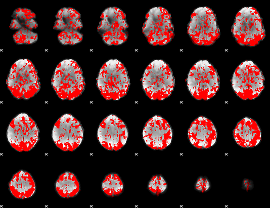 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
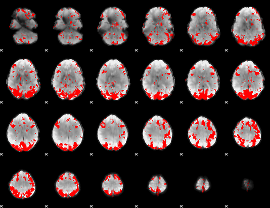 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:244; Jaccard:0.13417
 |
Component:244; Jaccard:0.15731
 |
Component:244; Jaccard:0.18013
 |
Component:043; Jaccard:0.21195
 |
Component:043; Jaccard:0.25383
 |
Component:043; Jaccard:0.2903
 |
Component:043; Jaccard:0.31578
 |
Correlation:0.3562
  |
Correlation:0.3562
  |
Correlation:0.3562
  |
Correlation:0.4519
  |
Correlation:0.4519
  |
Correlation:0.4519
  |
Correlation:0.4519
  |
Component:331; Jaccard:0.12441
 |
Component:331; Jaccard:0.15008
 |
Component:331; Jaccard:0.17843
 |
Component:331; Jaccard:0.20103
 |
Component:203; Jaccard:0.22977
 |
Component:203; Jaccard:0.25698
 |
Component:203; Jaccard:0.27402
 |
Correlation:0.34943
  |
Correlation:0.34943
  |
Correlation:0.34943
  |
Correlation:0.34943
  |
Correlation:0.30155
  |
Correlation:0.30155
  |
Correlation:0.30155
  |
Component:355; Jaccard:0.12036
 |
Component:355; Jaccard:0.13867
 |
Component:043; Jaccard:0.1712
 |
Component:203; Jaccard:0.19898
 |
Component:331; Jaccard:0.21783
 |
Component:331; Jaccard:0.22992
 |
Component:331; Jaccard:0.23233
 |
Correlation:0.30291
  |
Correlation:0.30291
  |
Correlation:0.4519
  |
Correlation:0.30155
  |
Correlation:0.34943
  |
Correlation:0.34943
  |
Correlation:0.34943
  |
Component:203; Jaccard:0.10991
 |
Component:203; Jaccard:0.13592
 |
Component:203; Jaccard:0.1663
 |
Component:244; Jaccard:0.19569
 |
Component:244; Jaccard:0.20424
 |
Component:223; Jaccard:0.20657
 |
Component:327; Jaccard:0.21122
 |
Correlation:0.30155
  |
Correlation:0.30155
  |
Correlation:0.30155
  |
Correlation:0.3562
  |
Correlation:0.3562
  |
Correlation:0.35992
  |
Correlation:0.29039
  |
Component:172; Jaccard:0.10801
 |
Component:043; Jaccard:0.13552
 |
Component:223; Jaccard:0.15701
 |
Component:223; Jaccard:0.18018
 |
Component:223; Jaccard:0.19842
 |
Component:327; Jaccard:0.20414
 |
Component:223; Jaccard:0.20181
 |
Correlation:0.073203
  |
Correlation:0.4519
  |
Correlation:0.35992
  |
Correlation:0.35992
  |
Correlation:0.35992
  |
Correlation:0.29039
  |
Correlation:0.35992
  |
Component:043; Jaccard:0.10738
 |
Component:223; Jaccard:0.13028
 |
Component:355; Jaccard:0.15602
 |
Component:327; Jaccard:0.16878
 |
Component:327; Jaccard:0.18983
 |
Component:244; Jaccard:0.20188
 |
Component:244; Jaccard:0.19022
 |
Correlation:0.4519
  |
Correlation:0.35992
  |
Correlation:0.30291
  |
Correlation:0.29039
  |
Correlation:0.29039
  |
Correlation:0.3562
  |
Correlation:0.3562
  |
Component:223; Jaccard:0.10709
 |
Component:172; Jaccard:0.1266
 |
Component:172; Jaccard:0.14738
 |
Component:355; Jaccard:0.16787
 |
Component:172; Jaccard:0.17528
 |
Component:172; Jaccard:0.18325
 |
Component:172; Jaccard:0.18516
 |
Correlation:0.35992
  |
Correlation:0.073203
  |
Correlation:0.073203
  |
Correlation:0.30291
  |
Correlation:0.073203
  |
Correlation:0.073203
  |
Correlation:0.073203
  |
Component:048; Jaccard:0.10308
 |
Component:327; Jaccard:0.12126
 |
Component:327; Jaccard:0.14534
 |
Component:172; Jaccard:0.16389
 |
Component:355; Jaccard:0.17402
 |
Component:355; Jaccard:0.1683
 |
Component:349; Jaccard:0.15804
 |
Correlation:0.27383
  |
Correlation:0.29039
  |
Correlation:0.29039
  |
Correlation:0.073203
  |
Correlation:0.30291
  |
Correlation:0.30291
  |
Correlation:0.40337
  |
Component:142; Jaccard:0.10227
 |
Component:349; Jaccard:0.117
 |
Component:349; Jaccard:0.13267
 |
Component:349; Jaccard:0.14533
 |
Component:349; Jaccard:0.15211
 |
Component:349; Jaccard:0.15845
 |
Component:355; Jaccard:0.1552
 |
Correlation:0.11138
  |
Correlation:0.40337
  |
Correlation:0.40337
  |
Correlation:0.40337
  |
Correlation:0.40337
  |
Correlation:0.40337
  |
Correlation:0.30291
  |
Component:349; Jaccard:0.10133
 |
Component:211; Jaccard:0.11547
 |
Component:016; Jaccard:0.1312
 |
Component:016; Jaccard:0.14424
 |
Component:016; Jaccard:0.15126
 |
Component:016; Jaccard:0.15338
 |
Component:211; Jaccard:0.15127
 |
Correlation:0.40337
  |
Correlation:0.17528
  |
Correlation:0.39501
  |
Correlation:0.39501
  |
Correlation:0.39501
  |
Correlation:0.39501
  |
Correlation:0.17528
  |
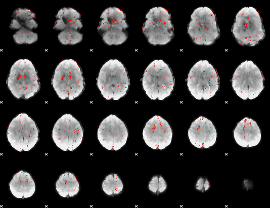
Cope: 03:MATCH-REL BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
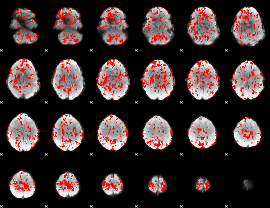 |
GLM binary result thresholded by 1.5
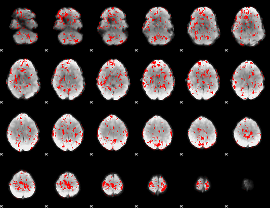 |
GLM binary result thresholded by 2
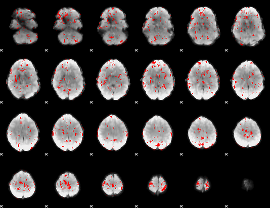 |
GLM binary result thresholded by 2.5
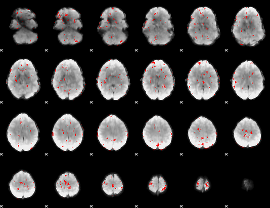 |
GLM binary result thresholded by 3
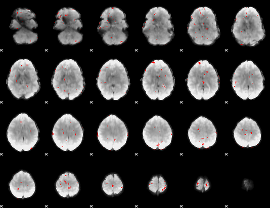 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:060; Jaccard:0.080652
 |
Component:399; Jaccard:0.084528
 |
Component:399; Jaccard:0.077815
 |
Component:399; Jaccard:0.056057
 |
Component:090; Jaccard:0.027399
 |
Component:189; Jaccard:0.011852
 |
Component:189; Jaccard:0.00399
 |
Correlation:-0.075804
  |
Correlation:-0.2673
  |
Correlation:-0.2673
  |
Correlation:-0.2673
  |
Correlation:0.17638
  |
Correlation:0.4976
  |
Correlation:0.4976
  |
Component:399; Jaccard:0.079898
 |
Component:060; Jaccard:0.082096
 |
Component:060; Jaccard:0.071884
 |
Component:379; Jaccard:0.049287
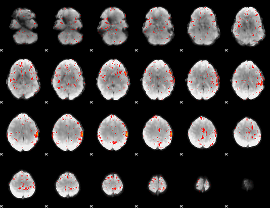 |
Component:399; Jaccard:0.027318
 |
Component:090; Jaccard:0.011362
 |
Component:281; Jaccard:0.003951
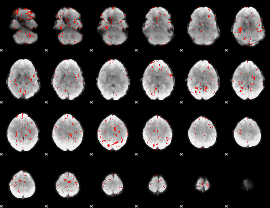 |
Correlation:-0.2673
  |
Correlation:-0.075804
  |
Correlation:-0.075804
  |
Correlation:-0.0087984
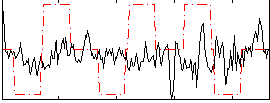 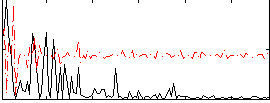 |
Correlation:-0.2673
  |
Correlation:0.17638
  |
Correlation:0.30504
  |
Component:079; Jaccard:0.079049
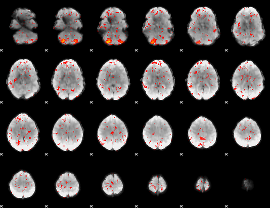 |
Component:079; Jaccard:0.07884
 |
Component:090; Jaccard:0.069009
 |
Component:090; Jaccard:0.049136
 |
Component:308; Jaccard:0.026063
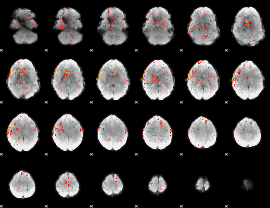 |
Component:308; Jaccard:0.011323
 |
Component:090; Jaccard:0.003744
 |
Correlation:-0.017111
  |
Correlation:-0.017111
  |
Correlation:0.17638
  |
Correlation:0.17638
  |
Correlation:0.16115
  |
Correlation:0.16115
  |
Correlation:0.17638
  |
Component:042; Jaccard:0.078999
 |
Component:379; Jaccard:0.076597
 |
Component:079; Jaccard:0.067525
 |
Component:060; Jaccard:0.046882
 |
Component:189; Jaccard:0.024708
 |
Component:281; Jaccard:0.010638
 |
Component:148; Jaccard:0.003025
 |
Correlation:0.0066737
  |
Correlation:-0.0087984
  |
Correlation:-0.017111
  |
Correlation:-0.075804
  |
Correlation:0.4976
  |
Correlation:0.30504
  |
Correlation:0.14587
  |
Component:379; Jaccard:0.075608
 |
Component:042; Jaccard:0.075342
 |
Component:379; Jaccard:0.065806
 |
Component:308; Jaccard:0.045314
 |
Component:379; Jaccard:0.024031
 |
Component:379; Jaccard:0.010251
 |
Component:207; Jaccard:0.002931
 |
Correlation:-0.0087984
  |
Correlation:0.0066737
  |
Correlation:-0.0087984
  |
Correlation:0.16115
  |
Correlation:-0.0087984
  |
Correlation:-0.0087984
  |
Correlation:0.3075
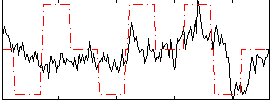 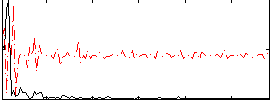 |
Component:090; Jaccard:0.072732
 |
Component:090; Jaccard:0.075243
 |
Component:078; Jaccard:0.063915
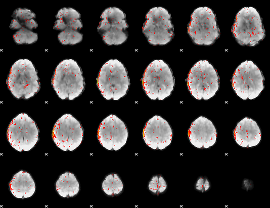 |
Component:079; Jaccard:0.044155
 |
Component:207; Jaccard:0.023036
 |
Component:228; Jaccard:0.010088
 |
Component:308; Jaccard:0.002878
 |
Correlation:0.17638
  |
Correlation:0.17638
  |
Correlation:0.10404
  |
Correlation:-0.017111
  |
Correlation:0.3075
  |
Correlation:0.20332
  |
Correlation:0.16115
  |
Component:257; Jaccard:0.072443
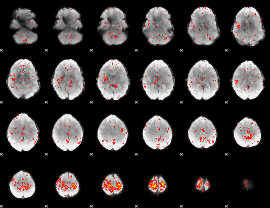 |
Component:308; Jaccard:0.071452
 |
Component:308; Jaccard:0.061448
 |
Component:189; Jaccard:0.044055
 |
Component:281; Jaccard:0.022274
 |
Component:148; Jaccard:0.009598
 |
Component:304; Jaccard:0.002858
 |
Correlation:-0.15715
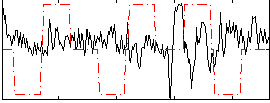 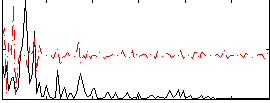 |
Correlation:0.16115
  |
Correlation:0.16115
  |
Correlation:0.4976
  |
Correlation:0.30504
  |
Correlation:0.14587
  |
Correlation:0.078681
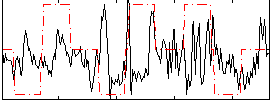  |
Component:310; Jaccard:0.071431
 |
Component:078; Jaccard:0.067093
 |
Component:042; Jaccard:0.061147
 |
Component:078; Jaccard:0.043631
 |
Component:079; Jaccard:0.022168
 |
Component:192; Jaccard:0.009265
 |
Component:228; Jaccard:0.002729
 |
Correlation:-0.07438
  |
Correlation:0.10404
  |
Correlation:0.0066737
  |
Correlation:0.10404
  |
Correlation:-0.017111
  |
Correlation:-0.01381
  |
Correlation:0.20332
  |
Component:308; Jaccard:0.071414
 |
Component:148; Jaccard:0.066628
 |
Component:189; Jaccard:0.060079
 |
Component:206; Jaccard:0.042345
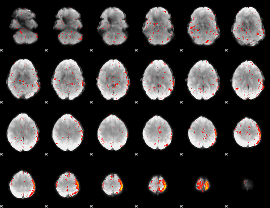 |
Component:025; Jaccard:0.022063
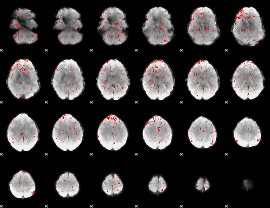 |
Component:025; Jaccard:0.009121
 |
Component:177; Jaccard:0.002625
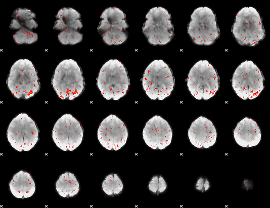 |
Correlation:0.16115
  |
Correlation:0.14587
  |
Correlation:0.4976
  |
Correlation:0.06929
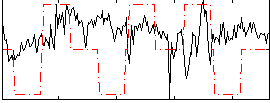 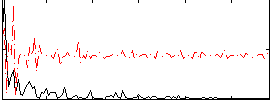 |
Correlation:0.065403
  |
Correlation:0.065403
  |
Correlation:0.11497
  |
Component:306; Jaccard:0.067287
 |
Component:310; Jaccard:0.066332
 |
Component:148; Jaccard:0.058641
 |
Component:042; Jaccard:0.042303
 |
Component:081; Jaccard:0.021263
 |
Component:078; Jaccard:0.008949
 |
Component:025; Jaccard:0.002542
 |
Correlation:0.098744
  |
Correlation:-0.07438
  |
Correlation:0.14587
  |
Correlation:0.0066737
  |
Correlation:-0.043513
  |
Correlation:0.10404
  |
Correlation:0.065403
  |
Cope: 04:REL-MATCH BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
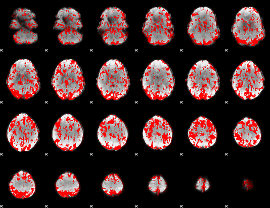 |
GLM binary result thresholded by 1.5
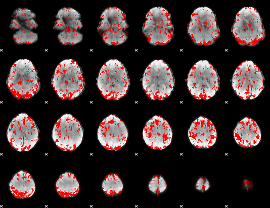 |
GLM binary result thresholded by 2
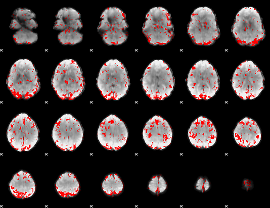 |
GLM binary result thresholded by 2.5
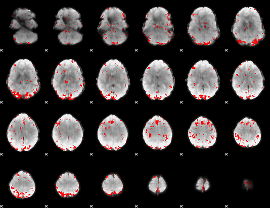 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
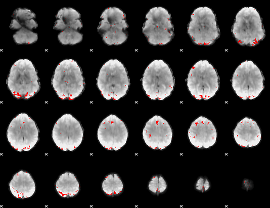 |
GLM binary result thresholded by 4
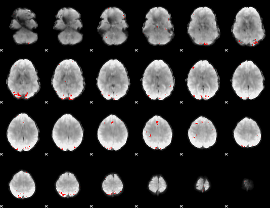 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:244; Jaccard:0.11324
 |
Component:244; Jaccard:0.12758
 |
Component:142; Jaccard:0.13373
 |
Component:142; Jaccard:0.13084
 |
Component:212; Jaccard:0.11874
 |
Component:212; Jaccard:0.090934
 |
Component:212; Jaccard:0.057733
 |
Correlation:0.26217
  |
Correlation:0.26217
  |
Correlation:0.2211
  |
Correlation:0.2211
  |
Correlation:0.42292
  |
Correlation:0.42292
  |
Correlation:0.42292
  |
Component:142; Jaccard:0.10693
 |
Component:142; Jaccard:0.12254
 |
Component:223; Jaccard:0.13159
 |
Component:223; Jaccard:0.13069
 |
Component:043; Jaccard:0.11038
 |
Component:043; Jaccard:0.088596
 |
Component:043; Jaccard:0.056707
 |
Correlation:0.2211
  |
Correlation:0.2211
  |
Correlation:0.27531
  |
Correlation:0.27531
  |
Correlation:0.19988
  |
Correlation:0.19988
  |
Correlation:0.19988
  |
Component:355; Jaccard:0.097923
 |
Component:223; Jaccard:0.1158
 |
Component:244; Jaccard:0.13041
 |
Component:212; Jaccard:0.12558
 |
Component:223; Jaccard:0.10995
 |
Component:211; Jaccard:0.077082
 |
Component:211; Jaccard:0.045495
 |
Correlation:0.32008
  |
Correlation:0.27531
  |
Correlation:0.26217
  |
Correlation:0.42292
  |
Correlation:0.27531
  |
Correlation:0.066003
  |
Correlation:0.066003
  |
Component:223; Jaccard:0.096907
 |
Component:211; Jaccard:0.1067
 |
Component:211; Jaccard:0.11698
 |
Component:211; Jaccard:0.11909
 |
Component:142; Jaccard:0.10625
 |
Component:223; Jaccard:0.076264
 |
Component:016; Jaccard:0.042527
 |
Correlation:0.27531
  |
Correlation:0.066003
  |
Correlation:0.066003
  |
Correlation:0.066003
  |
Correlation:0.2211
  |
Correlation:0.27531
  |
Correlation:0.37746
  |
Component:048; Jaccard:0.095997
 |
Component:355; Jaccard:0.10502
 |
Component:043; Jaccard:0.11446
 |
Component:043; Jaccard:0.11786
 |
Component:211; Jaccard:0.10241
 |
Component:016; Jaccard:0.073226
 |
Component:223; Jaccard:0.040416
 |
Correlation:0.26101
  |
Correlation:0.32008
  |
Correlation:0.19988
  |
Correlation:0.19988
  |
Correlation:0.066003
  |
Correlation:0.37746
  |
Correlation:0.27531
  |
Component:211; Jaccard:0.093456
 |
Component:016; Jaccard:0.10225
 |
Component:016; Jaccard:0.11273
 |
Component:244; Jaccard:0.11352
 |
Component:016; Jaccard:0.098786
 |
Component:142; Jaccard:0.070754
 |
Component:142; Jaccard:0.037019
 |
Correlation:0.066003
  |
Correlation:0.37746
  |
Correlation:0.37746
  |
Correlation:0.26217
  |
Correlation:0.37746
  |
Correlation:0.2211
  |
Correlation:0.2211
  |
Component:047; Jaccard:0.090563
 |
Component:043; Jaccard:0.10061
 |
Component:212; Jaccard:0.11072
 |
Component:016; Jaccard:0.11317
 |
Component:244; Jaccard:0.084288
 |
Component:203; Jaccard:0.061751
 |
Component:203; Jaccard:0.034681
 |
Correlation:0.18431
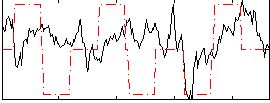 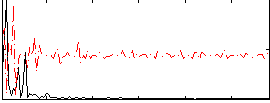 |
Correlation:0.19988
  |
Correlation:0.42292
  |
Correlation:0.37746
  |
Correlation:0.26217
  |
Correlation:0.11424
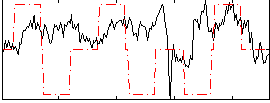 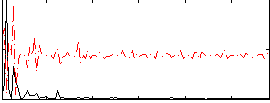 |
Correlation:0.11424
  |
Component:016; Jaccard:0.088795
 |
Component:048; Jaccard:0.099538
 |
Component:355; Jaccard:0.10504
 |
Component:126; Jaccard:0.09427
 |
Component:203; Jaccard:0.081476
 |
Component:126; Jaccard:0.053706
 |
Component:126; Jaccard:0.033349
 |
Correlation:0.37746
  |
Correlation:0.26101
  |
Correlation:0.32008
  |
Correlation:0.26189
  |
Correlation:0.11424
  |
Correlation:0.26189
  |
Correlation:0.26189
  |
Component:043; Jaccard:0.085205
 |
Component:047; Jaccard:0.098783
 |
Component:047; Jaccard:0.10036
 |
Component:203; Jaccard:0.091623
 |
Component:126; Jaccard:0.077727
 |
Component:365; Jaccard:0.053573
 |
Component:080; Jaccard:0.03237
 |
Correlation:0.19988
  |
Correlation:0.18431
  |
Correlation:0.18431
  |
Correlation:0.11424
  |
Correlation:0.26189
  |
Correlation:0.38494
  |
Correlation:-0.018332
 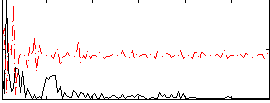 |
Component:126; Jaccard:0.082834
 |
Component:126; Jaccard:0.092673
 |
Component:126; Jaccard:0.098327
 |
Component:355; Jaccard:0.089845
 |
Component:365; Jaccard:0.076956
 |
Component:244; Jaccard:0.053039
 |
Component:349; Jaccard:0.030818
 |
Correlation:0.26189
  |
Correlation:0.26189
  |
Correlation:0.26189
  |
Correlation:0.32008
  |
Correlation:0.38494
  |
Correlation:0.26217
  |
Correlation:0.30969
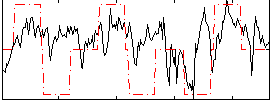  |
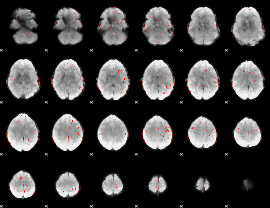
Cope: 05:neg_MATCH BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
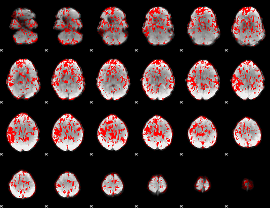 |
GLM binary result thresholded by 1.5
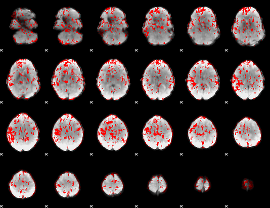 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:279; Jaccard:0.10463
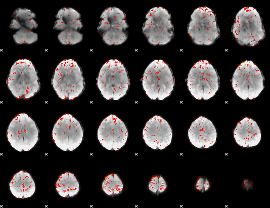 |
Component:279; Jaccard:0.114
 |
Component:279; Jaccard:0.11581
 |
Component:279; Jaccard:0.10767
 |
Component:334; Jaccard:0.088328
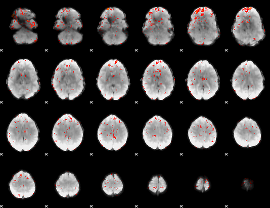 |
Component:334; Jaccard:0.062081
 |
Component:334; Jaccard:0.035263
 |
Correlation:0.34157
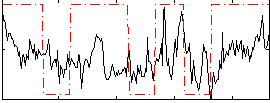 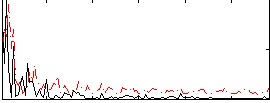 |
Correlation:0.34157
  |
Correlation:0.34157
  |
Correlation:0.34157
  |
Correlation:0.50422
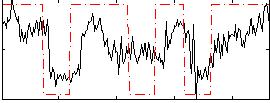 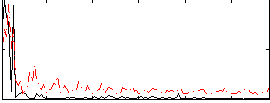 |
Correlation:0.50422
  |
Correlation:0.50422
  |
Component:191; Jaccard:0.097829
 |
Component:191; Jaccard:0.10466
 |
Component:191; Jaccard:0.10552
 |
Component:334; Jaccard:0.10326
 |
Component:032; Jaccard:0.088074
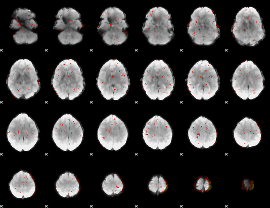 |
Component:032; Jaccard:0.061545
 |
Component:272; Jaccard:0.033398
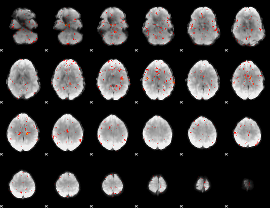 |
Correlation:0.26512
  |
Correlation:0.26512
  |
Correlation:0.26512
  |
Correlation:0.50422
  |
Correlation:0.18251
  |
Correlation:0.18251
  |
Correlation:-0.030229
  |
Component:395; Jaccard:0.087721
 |
Component:334; Jaccard:0.097344
 |
Component:334; Jaccard:0.10459
 |
Component:032; Jaccard:0.097304
 |
Component:374; Jaccard:0.086339
 |
Component:374; Jaccard:0.058314
 |
Component:170; Jaccard:0.032042
 |
Correlation:0.33671
  |
Correlation:0.50422
  |
Correlation:0.50422
  |
Correlation:0.18251
  |
Correlation:0.041837
  |
Correlation:0.041837
  |
Correlation:0.093906
  |
Component:334; Jaccard:0.086095
 |
Component:395; Jaccard:0.089449
 |
Component:374; Jaccard:0.092999
 |
Component:374; Jaccard:0.095554
 |
Component:279; Jaccard:0.085544
 |
Component:170; Jaccard:0.058227
 |
Component:279; Jaccard:0.029277
 |
Correlation:0.50422
  |
Correlation:0.33671
  |
Correlation:0.041837
  |
Correlation:0.041837
  |
Correlation:0.34157
  |
Correlation:0.093906
  |
Correlation:0.34157
  |
Component:253; Jaccard:0.072277
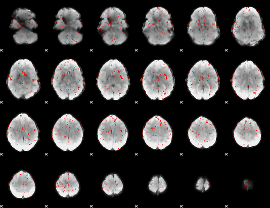 |
Component:374; Jaccard:0.082557
 |
Component:170; Jaccard:0.090673
 |
Component:191; Jaccard:0.094578
 |
Component:170; Jaccard:0.0813
 |
Component:279; Jaccard:0.055006
 |
Component:249; Jaccard:0.028882
 |
Correlation:0.031591
  |
Correlation:0.041837
  |
Correlation:0.093906
  |
Correlation:0.26512
  |
Correlation:0.093906
  |
Correlation:0.34157
  |
Correlation:0.10034
  |
Component:374; Jaccard:0.069924
 |
Component:170; Jaccard:0.080254
 |
Component:032; Jaccard:0.088395
 |
Component:170; Jaccard:0.091959
 |
Component:282; Jaccard:0.07721
 |
Component:282; Jaccard:0.052388
 |
Component:032; Jaccard:0.028617
 |
Correlation:0.041837
  |
Correlation:0.093906
  |
Correlation:0.18251
  |
Correlation:0.093906
  |
Correlation:0.12297
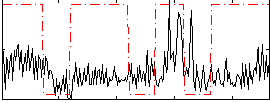 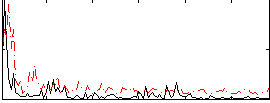 |
Correlation:0.12297
  |
Correlation:0.18251
  |
Component:170; Jaccard:0.068979
 |
Component:253; Jaccard:0.079799
 |
Component:282; Jaccard:0.084943
 |
Component:282; Jaccard:0.087796
 |
Component:191; Jaccard:0.073626
 |
Component:180; Jaccard:0.050056
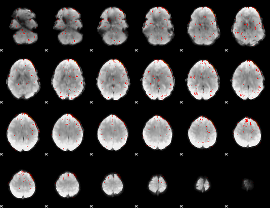 |
Component:374; Jaccard:0.028576
 |
Correlation:0.093906
  |
Correlation:0.031591
  |
Correlation:0.12297
  |
Correlation:0.12297
  |
Correlation:0.26512
  |
Correlation:-0.10016
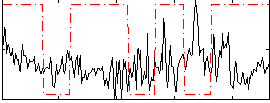 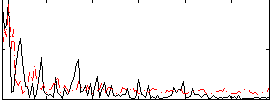 |
Correlation:0.041837
  |
Component:353; Jaccard:0.068872
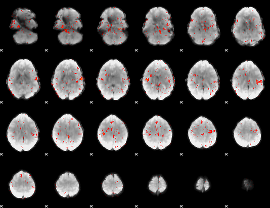 |
Component:032; Jaccard:0.074984
 |
Component:253; Jaccard:0.083895
 |
Component:180; Jaccard:0.079572
 |
Component:180; Jaccard:0.069448
 |
Component:272; Jaccard:0.048426
 |
Component:352; Jaccard:0.027212
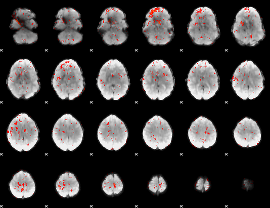 |
Correlation:0.17909
  |
Correlation:0.18251
  |
Correlation:0.031591
  |
Correlation:-0.10016
  |
Correlation:-0.10016
  |
Correlation:-0.030229
  |
Correlation:0.29241
  |
Component:226; Jaccard:0.067969
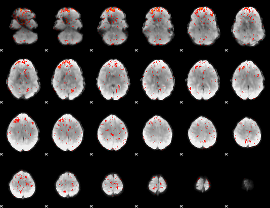 |
Component:269; Jaccard:0.074364
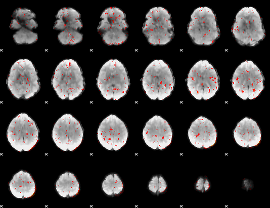 |
Component:395; Jaccard:0.083777
 |
Component:253; Jaccard:0.078474
 |
Component:352; Jaccard:0.064563
 |
Component:191; Jaccard:0.045599
 |
Component:180; Jaccard:0.026339
 |
Correlation:0.34239
  |
Correlation:0.087308
  |
Correlation:0.33671
  |
Correlation:0.031591
  |
Correlation:0.29241
  |
Correlation:0.26512
  |
Correlation:-0.10016
  |
Component:269; Jaccard:0.067759
 |
Component:180; Jaccard:0.074278
 |
Component:180; Jaccard:0.079538
 |
Component:395; Jaccard:0.07603
 |
Component:253; Jaccard:0.063491
 |
Component:352; Jaccard:0.045136
 |
Component:253; Jaccard:0.026202
 |
Correlation:0.087308
  |
Correlation:-0.10016
  |
Correlation:-0.10016
  |
Correlation:0.33671
  |
Correlation:0.031591
  |
Correlation:0.29241
  |
Correlation:0.031591
  |
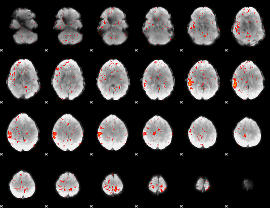
Cope: 06:neg_REL BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
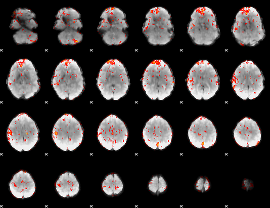 |
GLM Z-score result thresholded by 3
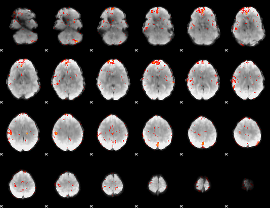 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
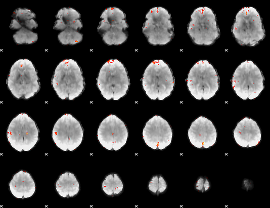 |
Component:395; Jaccard:0.10594
 |
Component:395; Jaccard:0.11532
 |
Component:395; Jaccard:0.12093
 |
Component:228; Jaccard:0.13172
 |
Component:228; Jaccard:0.13149
 |
Component:228; Jaccard:0.11169
 |
Component:228; Jaccard:0.074831
 |
Correlation:0.19074
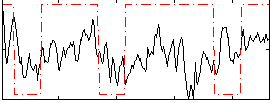  |
Correlation:0.19074
  |
Correlation:0.19074
  |
Correlation:0.35268
  |
Correlation:0.35268
  |
Correlation:0.35268
  |
Correlation:0.35268
  |
Component:191; Jaccard:0.10409
 |
Component:191; Jaccard:0.11248
 |
Component:228; Jaccard:0.11906
 |
Component:395; Jaccard:0.11759
 |
Component:395; Jaccard:0.10508
 |
Component:395; Jaccard:0.083348
 |
Component:395; Jaccard:0.058175
 |
Correlation:0.061966
  |
Correlation:0.061966
  |
Correlation:0.35268
  |
Correlation:0.19074
  |
Correlation:0.19074
  |
Correlation:0.19074
  |
Correlation:0.19074
  |
Component:228; Jaccard:0.088548
 |
Component:228; Jaccard:0.10492
 |
Component:191; Jaccard:0.11654
 |
Component:191; Jaccard:0.10905
 |
Component:149; Jaccard:0.096249
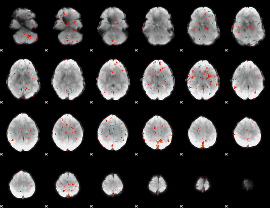 |
Component:149; Jaccard:0.081613
 |
Component:149; Jaccard:0.056057
 |
Correlation:0.35268
  |
Correlation:0.35268
  |
Correlation:0.061966
  |
Correlation:0.061966
  |
Correlation:0.23689
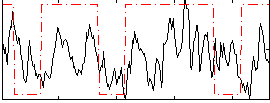 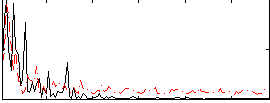 |
Correlation:0.23689
  |
Correlation:0.23689
  |
Component:313; Jaccard:0.084723
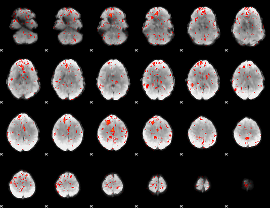 |
Component:381; Jaccard:0.092485
 |
Component:381; Jaccard:0.099065
 |
Component:381; Jaccard:0.09907
 |
Component:381; Jaccard:0.093364
 |
Component:179; Jaccard:0.072656
 |
Component:381; Jaccard:0.045561
 |
Correlation:0.062161
  |
Correlation:0.0049083
  |
Correlation:0.0049083
  |
Correlation:0.0049083
  |
Correlation:0.0049083
  |
Correlation:0.18172
  |
Correlation:0.0049083
  |
Component:381; Jaccard:0.084154
 |
Component:256; Jaccard:0.089078
 |
Component:179; Jaccard:0.093898
 |
Component:179; Jaccard:0.097633
 |
Component:191; Jaccard:0.089617
 |
Component:381; Jaccard:0.070277
 |
Component:179; Jaccard:0.042597
 |
Correlation:0.0049083
  |
Correlation:0.24178
  |
Correlation:0.18172
  |
Correlation:0.18172
  |
Correlation:0.061966
  |
Correlation:0.0049083
  |
Correlation:0.18172
  |
Component:279; Jaccard:0.083547
 |
Component:313; Jaccard:0.086581
 |
Component:256; Jaccard:0.090913
 |
Component:149; Jaccard:0.093974
 |
Component:179; Jaccard:0.088887
 |
Component:011; Jaccard:0.065963
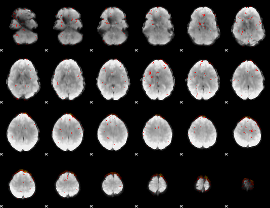 |
Component:191; Jaccard:0.042209
 |
Correlation:0.0673
  |
Correlation:0.062161
  |
Correlation:0.24178
  |
Correlation:0.23689
  |
Correlation:0.18172
  |
Correlation:0.27822
  |
Correlation:0.061966
  |
Component:256; Jaccard:0.081276
 |
Component:179; Jaccard:0.08512
 |
Component:170; Jaccard:0.088416
 |
Component:011; Jaccard:0.089907
 |
Component:011; Jaccard:0.080878
 |
Component:191; Jaccard:0.064195
 |
Component:027; Jaccard:0.041439
 |
Correlation:0.24178
  |
Correlation:0.18172
  |
Correlation:0.10635
  |
Correlation:0.27822
  |
Correlation:0.27822
  |
Correlation:0.061966
  |
Correlation:0.12772
  |
Component:027; Jaccard:0.080413
 |
Component:027; Jaccard:0.08507
 |
Component:027; Jaccard:0.086024
 |
Component:170; Jaccard:0.088536
 |
Component:354; Jaccard:0.076066
 |
Component:027; Jaccard:0.058154
 |
Component:011; Jaccard:0.039705
 |
Correlation:0.12772
  |
Correlation:0.12772
  |
Correlation:0.12772
  |
Correlation:0.10635
  |
Correlation:0.11917
  |
Correlation:0.12772
  |
Correlation:0.27822
  |
Component:079; Jaccard:0.077956
 |
Component:279; Jaccard:0.084939
 |
Component:354; Jaccard:0.085733
 |
Component:256; Jaccard:0.087379
 |
Component:352; Jaccard:0.075643
 |
Component:256; Jaccard:0.057682
 |
Component:352; Jaccard:0.038868
 |
Correlation:0.10026
  |
Correlation:0.0673
  |
Correlation:0.11917
  |
Correlation:0.24178
  |
Correlation:0.01401
  |
Correlation:0.24178
  |
Correlation:0.01401
  |
Component:033; Jaccard:0.077482
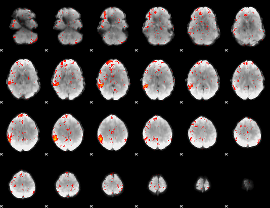 |
Component:170; Jaccard:0.080739
 |
Component:149; Jaccard:0.085323
 |
Component:354; Jaccard:0.086661
 |
Component:256; Jaccard:0.074605
 |
Component:352; Jaccard:0.057255
 |
Component:256; Jaccard:0.038162
 |
Correlation:0.39759
  |
Correlation:0.10635
  |
Correlation:0.23689
  |
Correlation:0.11917
  |
Correlation:0.24178
  |
Correlation:0.01401
  |
Correlation:0.24178
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































