Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:FACES BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
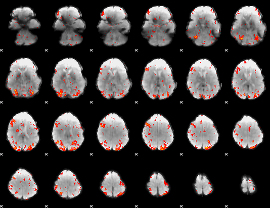 |
GLM Z-score result thresholded by 3
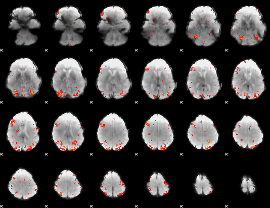 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:026; Jaccard:0.13848
 |
Component:026; Jaccard:0.17878
 |
Component:026; Jaccard:0.21635
 |
Component:026; Jaccard:0.23256
 |
Component:026; Jaccard:0.21296
 |
Component:026; Jaccard:0.16142
 |
Component:026; Jaccard:0.10588
 |
Correlation:0.18118
  |
Correlation:0.18118
  |
Correlation:0.18118
  |
Correlation:0.18118
  |
Correlation:0.18118
  |
Correlation:0.18118
  |
Correlation:0.18118
  |
Component:259; Jaccard:0.10506
 |
Component:259; Jaccard:0.12987
 |
Component:259; Jaccard:0.14666
 |
Component:244; Jaccard:0.14848
 |
Component:244; Jaccard:0.13974
 |
Component:244; Jaccard:0.11369
 |
Component:244; Jaccard:0.074671
 |
Correlation:0.18548
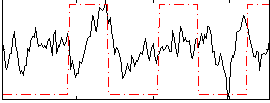  |
Correlation:0.18548
  |
Correlation:0.18548
  |
Correlation:-0.12833
  |
Correlation:-0.12833
  |
Correlation:-0.12833
  |
Correlation:-0.12833
  |
Component:314; Jaccard:0.10187
 |
Component:224; Jaccard:0.11922
 |
Component:224; Jaccard:0.13542
 |
Component:259; Jaccard:0.14446
 |
Component:224; Jaccard:0.12699
 |
Component:224; Jaccard:0.095098
 |
Component:224; Jaccard:0.066591
 |
Correlation:0.033706
 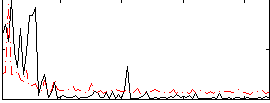 |
Correlation:0.46273
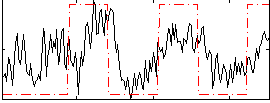 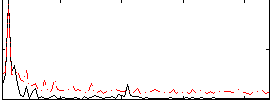 |
Correlation:0.46273
  |
Correlation:0.18548
  |
Correlation:0.46273
  |
Correlation:0.46273
  |
Correlation:0.46273
  |
Component:224; Jaccard:0.10021
 |
Component:159; Jaccard:0.11785
 |
Component:159; Jaccard:0.13504
 |
Component:159; Jaccard:0.13937
 |
Component:159; Jaccard:0.12415
 |
Component:159; Jaccard:0.090597
 |
Component:159; Jaccard:0.058083
 |
Correlation:0.46273
  |
Correlation:0.071596
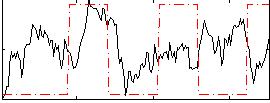 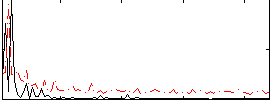 |
Correlation:0.071596
  |
Correlation:0.071596
  |
Correlation:0.071596
  |
Correlation:0.071596
  |
Correlation:0.071596
  |
Component:229; Jaccard:0.097502
 |
Component:314; Jaccard:0.11637
 |
Component:244; Jaccard:0.13333
 |
Component:224; Jaccard:0.13875
 |
Component:259; Jaccard:0.1216
 |
Component:259; Jaccard:0.087
 |
Component:259; Jaccard:0.05578
 |
Correlation:-0.084688
  |
Correlation:0.033706
  |
Correlation:-0.12833
  |
Correlation:0.46273
  |
Correlation:0.18548
  |
Correlation:0.18548
  |
Correlation:0.18548
  |
Component:159; Jaccard:0.095882
 |
Component:229; Jaccard:0.11292
 |
Component:314; Jaccard:0.12359
 |
Component:314; Jaccard:0.12078
 |
Component:314; Jaccard:0.10178
 |
Component:229; Jaccard:0.076354
 |
Component:229; Jaccard:0.05164
 |
Correlation:0.071596
  |
Correlation:-0.084688
  |
Correlation:0.033706
  |
Correlation:0.033706
  |
Correlation:0.033706
  |
Correlation:-0.084688
  |
Correlation:-0.084688
  |
Component:260; Jaccard:0.091268
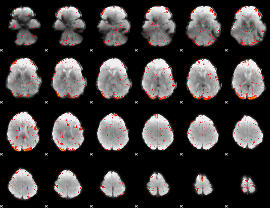 |
Component:244; Jaccard:0.11001
 |
Component:229; Jaccard:0.1207
 |
Component:229; Jaccard:0.11707
 |
Component:018; Jaccard:0.098075
 |
Component:314; Jaccard:0.07356
 |
Component:018; Jaccard:0.048294
 |
Correlation:0.31652
  |
Correlation:-0.12833
  |
Correlation:-0.084688
  |
Correlation:-0.084688
  |
Correlation:0.063895
  |
Correlation:0.033706
  |
Correlation:0.063895
  |
Component:255; Jaccard:0.090976
 |
Component:255; Jaccard:0.10444
 |
Component:255; Jaccard:0.11489
 |
Component:018; Jaccard:0.11277
 |
Component:229; Jaccard:0.096423
 |
Component:018; Jaccard:0.071793
 |
Component:255; Jaccard:0.047177
 |
Correlation:-0.014403
  |
Correlation:-0.014403
  |
Correlation:-0.014403
  |
Correlation:0.063895
  |
Correlation:-0.084688
  |
Correlation:0.063895
  |
Correlation:-0.014403
  |
Component:172; Jaccard:0.090894
 |
Component:018; Jaccard:0.10145
 |
Component:018; Jaccard:0.11386
 |
Component:255; Jaccard:0.10944
 |
Component:255; Jaccard:0.093378
 |
Component:255; Jaccard:0.070825
 |
Component:314; Jaccard:0.046857
 |
Correlation:0.2339
  |
Correlation:0.063895
  |
Correlation:0.063895
  |
Correlation:-0.014403
  |
Correlation:-0.014403
  |
Correlation:-0.014403
  |
Correlation:0.033706
  |
Component:362; Jaccard:0.089001
 |
Component:260; Jaccard:0.099732
 |
Component:362; Jaccard:0.10345
 |
Component:362; Jaccard:0.097854
 |
Component:362; Jaccard:0.083548
 |
Component:362; Jaccard:0.060173
 |
Component:362; Jaccard:0.036037
 |
Correlation:0.081104
  |
Correlation:0.31652
  |
Correlation:0.081104
  |
Correlation:0.081104
  |
Correlation:0.081104
  |
Correlation:0.081104
  |
Correlation:0.081104
  |
Cope: 02:SHAPES BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
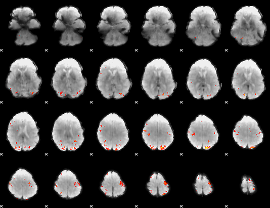 |
Component:026; Jaccard:0.13443
 |
Component:337; Jaccard:0.16813
 |
Component:244; Jaccard:0.21272
 |
Component:244; Jaccard:0.24302
 |
Component:244; Jaccard:0.24152
 |
Component:244; Jaccard:0.2015
 |
Component:244; Jaccard:0.1504
 |
Correlation:-0.026358
  |
Correlation:0.34266
  |
Correlation:0.3924
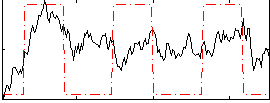  |
Correlation:0.3924
  |
Correlation:0.3924
  |
Correlation:0.3924
  |
Correlation:0.3924
  |
Component:337; Jaccard:0.12923
 |
Component:244; Jaccard:0.1659
 |
Component:337; Jaccard:0.20242
 |
Component:337; Jaccard:0.21389
 |
Component:337; Jaccard:0.19613
 |
Component:337; Jaccard:0.15228
 |
Component:337; Jaccard:0.10404
 |
Correlation:0.34266
  |
Correlation:0.3924
  |
Correlation:0.34266
  |
Correlation:0.34266
  |
Correlation:0.34266
  |
Correlation:0.34266
  |
Correlation:0.34266
  |
Component:244; Jaccard:0.12029
 |
Component:026; Jaccard:0.1621
 |
Component:026; Jaccard:0.17721
 |
Component:026; Jaccard:0.17365
 |
Component:026; Jaccard:0.15251
 |
Component:026; Jaccard:0.11433
 |
Component:026; Jaccard:0.079189
 |
Correlation:0.3924
  |
Correlation:-0.026358
  |
Correlation:-0.026358
  |
Correlation:-0.026358
  |
Correlation:-0.026358
  |
Correlation:-0.026358
  |
Correlation:-0.026358
  |
Component:229; Jaccard:0.11529
 |
Component:229; Jaccard:0.13335
 |
Component:137; Jaccard:0.14384
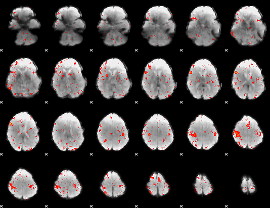 |
Component:018; Jaccard:0.1411
 |
Component:018; Jaccard:0.12033
 |
Component:018; Jaccard:0.094306
 |
Component:137; Jaccard:0.062187
 |
Correlation:0.20157
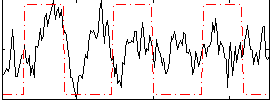  |
Correlation:0.20157
  |
Correlation:0.11162
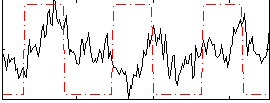  |
Correlation:0.037946
 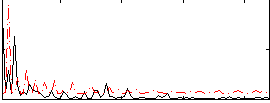 |
Correlation:0.037946
  |
Correlation:0.037946
  |
Correlation:0.11162
  |
Component:314; Jaccard:0.11518
 |
Component:314; Jaccard:0.1321
 |
Component:229; Jaccard:0.142
 |
Component:137; Jaccard:0.1378
 |
Component:137; Jaccard:0.11937
 |
Component:137; Jaccard:0.089092
 |
Component:018; Jaccard:0.061042
 |
Correlation:0.044803
  |
Correlation:0.044803
  |
Correlation:0.20157
  |
Correlation:0.11162
  |
Correlation:0.11162
  |
Correlation:0.11162
  |
Correlation:0.037946
  |
Component:137; Jaccard:0.10778
 |
Component:137; Jaccard:0.13189
 |
Component:018; Jaccard:0.13972
 |
Component:229; Jaccard:0.13648
 |
Component:229; Jaccard:0.11385
 |
Component:314; Jaccard:0.087756
 |
Component:314; Jaccard:0.060944
 |
Correlation:0.11162
  |
Correlation:0.11162
  |
Correlation:0.037946
  |
Correlation:0.20157
  |
Correlation:0.20157
  |
Correlation:0.044803
  |
Correlation:0.044803
  |
Component:255; Jaccard:0.10576
 |
Component:018; Jaccard:0.12273
 |
Component:314; Jaccard:0.13848
 |
Component:314; Jaccard:0.13117
 |
Component:314; Jaccard:0.11187
 |
Component:229; Jaccard:0.08737
 |
Component:229; Jaccard:0.060241
 |
Correlation:0.25041
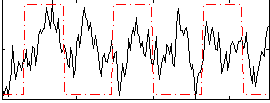 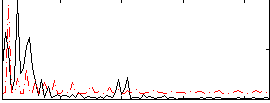 |
Correlation:0.037946
  |
Correlation:0.044803
  |
Correlation:0.044803
  |
Correlation:0.044803
  |
Correlation:0.20157
  |
Correlation:0.20157
  |
Component:018; Jaccard:0.098325
 |
Component:255; Jaccard:0.12098
 |
Component:255; Jaccard:0.12807
 |
Component:255; Jaccard:0.11983
 |
Component:255; Jaccard:0.099868
 |
Component:255; Jaccard:0.074734
 |
Component:255; Jaccard:0.051533
 |
Correlation:0.037946
  |
Correlation:0.25041
  |
Correlation:0.25041
  |
Correlation:0.25041
  |
Correlation:0.25041
  |
Correlation:0.25041
  |
Correlation:0.25041
  |
Component:349; Jaccard:0.08334
 |
Component:389; Jaccard:0.089427
 |
Component:159; Jaccard:0.093454
 |
Component:194; Jaccard:0.090486
 |
Component:194; Jaccard:0.08049
 |
Component:194; Jaccard:0.062599
 |
Component:194; Jaccard:0.045307
 |
Correlation:0.023806
  |
Correlation:0.25428
  |
Correlation:0.039195
 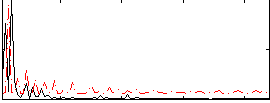 |
Correlation:0.37459
  |
Correlation:0.37459
  |
Correlation:0.37459
  |
Correlation:0.37459
  |
Component:159; Jaccard:0.080189
 |
Component:159; Jaccard:0.089388
 |
Component:389; Jaccard:0.092409
 |
Component:389; Jaccard:0.088608
 |
Component:389; Jaccard:0.079391
 |
Component:389; Jaccard:0.060812
 |
Component:389; Jaccard:0.044317
 |
Correlation:0.039195
  |
Correlation:0.039195
  |
Correlation:0.25428
  |
Correlation:0.25428
  |
Correlation:0.25428
  |
Correlation:0.25428
  |
Correlation:0.25428
  |
Cope: 03:FACES-SHAPES BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
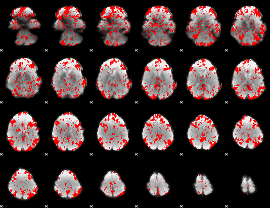 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:266; Jaccard:0.11296
 |
Component:060; Jaccard:0.13675
 |
Component:060; Jaccard:0.16348
 |
Component:217; Jaccard:0.17861
 |
Component:217; Jaccard:0.18029
 |
Component:217; Jaccard:0.15807
 |
Component:217; Jaccard:0.12322
 |
Correlation:0.39491
  |
Correlation:0.27478
  |
Correlation:0.27478
  |
Correlation:0.32197
  |
Correlation:0.32197
  |
Correlation:0.32197
  |
Correlation:0.32197
  |
Component:060; Jaccard:0.10891
 |
Component:266; Jaccard:0.13504
 |
Component:217; Jaccard:0.15648
 |
Component:060; Jaccard:0.17659
 |
Component:060; Jaccard:0.16681
 |
Component:224; Jaccard:0.13996
 |
Component:224; Jaccard:0.10534
 |
Correlation:0.27478
  |
Correlation:0.39491
  |
Correlation:0.32197
  |
Correlation:0.27478
  |
Correlation:0.27478
  |
Correlation:0.49631
  |
Correlation:0.49631
  |
Component:224; Jaccard:0.107
 |
Component:224; Jaccard:0.13313
 |
Component:224; Jaccard:0.15585
 |
Component:224; Jaccard:0.16928
 |
Component:224; Jaccard:0.16497
 |
Component:060; Jaccard:0.13634
 |
Component:060; Jaccard:0.099528
 |
Correlation:0.49631
  |
Correlation:0.49631
  |
Correlation:0.49631
  |
Correlation:0.49631
  |
Correlation:0.49631
  |
Correlation:0.27478
  |
Correlation:0.27478
  |
Component:217; Jaccard:0.099338
 |
Component:217; Jaccard:0.12667
 |
Component:266; Jaccard:0.14807
 |
Component:266; Jaccard:0.14856
 |
Component:266; Jaccard:0.13115
 |
Component:266; Jaccard:0.103
 |
Component:384; Jaccard:0.077241
 |
Correlation:0.32197
  |
Correlation:0.32197
  |
Correlation:0.39491
  |
Correlation:0.39491
  |
Correlation:0.39491
  |
Correlation:0.39491
  |
Correlation:0.29123
  |
Component:260; Jaccard:0.094832
 |
Component:260; Jaccard:0.10948
 |
Component:384; Jaccard:0.12288
 |
Component:384; Jaccard:0.12922
 |
Component:384; Jaccard:0.12233
 |
Component:384; Jaccard:0.1015
 |
Component:266; Jaccard:0.075813
 |
Correlation:0.25427
  |
Correlation:0.25427
  |
Correlation:0.29123
  |
Correlation:0.29123
  |
Correlation:0.29123
  |
Correlation:0.29123
  |
Correlation:0.39491
  |
Component:384; Jaccard:0.088271
 |
Component:384; Jaccard:0.10681
 |
Component:260; Jaccard:0.12134
 |
Component:260; Jaccard:0.12254
 |
Component:260; Jaccard:0.10818
 |
Component:382; Jaccard:0.082907
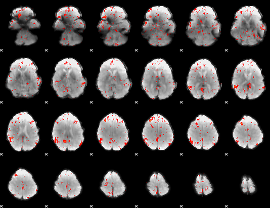 |
Component:382; Jaccard:0.061773
 |
Correlation:0.29123
  |
Correlation:0.29123
  |
Correlation:0.25427
  |
Correlation:0.25427
  |
Correlation:0.25427
  |
Correlation:0.44642
 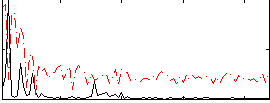 |
Correlation:0.44642
  |
Component:259; Jaccard:0.08391
 |
Component:383; Jaccard:0.098937
 |
Component:383; Jaccard:0.11205
 |
Component:383; Jaccard:0.1125
 |
Component:383; Jaccard:0.099971
 |
Component:260; Jaccard:0.08109
 |
Component:071; Jaccard:0.053549
 |
Correlation:0.21659
  |
Correlation:0.37639
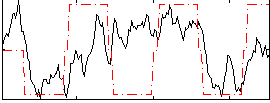 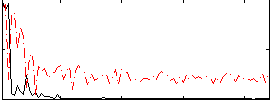 |
Correlation:0.37639
  |
Correlation:0.37639
  |
Correlation:0.37639
  |
Correlation:0.25427
  |
Correlation:0.27718
  |
Component:071; Jaccard:0.082905
 |
Component:071; Jaccard:0.097757
 |
Component:071; Jaccard:0.1077
 |
Component:071; Jaccard:0.10866
 |
Component:382; Jaccard:0.098193
 |
Component:071; Jaccard:0.077975
 |
Component:383; Jaccard:0.051609
 |
Correlation:0.27718
  |
Correlation:0.27718
  |
Correlation:0.27718
  |
Correlation:0.27718
  |
Correlation:0.44642
  |
Correlation:0.27718
  |
Correlation:0.37639
  |
Component:383; Jaccard:0.082249
 |
Component:259; Jaccard:0.094647
 |
Component:382; Jaccard:0.10198
 |
Component:382; Jaccard:0.10697
 |
Component:071; Jaccard:0.096106
 |
Component:383; Jaccard:0.075005
 |
Component:260; Jaccard:0.051418
 |
Correlation:0.37639
  |
Correlation:0.21659
  |
Correlation:0.44642
  |
Correlation:0.44642
  |
Correlation:0.27718
  |
Correlation:0.37639
  |
Correlation:0.25427
  |
Component:276; Jaccard:0.080558
 |
Component:382; Jaccard:0.09274
 |
Component:259; Jaccard:0.099908
 |
Component:259; Jaccard:0.098624
 |
Component:259; Jaccard:0.086503
 |
Component:259; Jaccard:0.068172
 |
Component:259; Jaccard:0.049771
 |
Correlation:-0.0037074
  |
Correlation:0.44642
  |
Correlation:0.21659
  |
Correlation:0.21659
  |
Correlation:0.21659
  |
Correlation:0.21659
  |
Correlation:0.21659
  |
Cope: 04:neg_FACES BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:132; Jaccard:0.099524
 |
Component:132; Jaccard:0.12244
 |
Component:132; Jaccard:0.14175
 |
Component:132; Jaccard:0.13795
 |
Component:132; Jaccard:0.11552
 |
Component:132; Jaccard:0.07871
 |
Component:132; Jaccard:0.045905
 |
Correlation:0.46725
  |
Correlation:0.46725
  |
Correlation:0.46725
  |
Correlation:0.46725
  |
Correlation:0.46725
  |
Correlation:0.46725
  |
Correlation:0.46725
  |
Component:286; Jaccard:0.097835
 |
Component:286; Jaccard:0.11314
 |
Component:286; Jaccard:0.11786
 |
Component:286; Jaccard:0.10691
 |
Component:286; Jaccard:0.075277
 |
Component:286; Jaccard:0.047661
 |
Component:374; Jaccard:0.026186
 |
Correlation:0.27769
 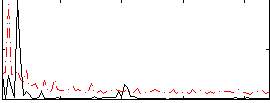 |
Correlation:0.27769
  |
Correlation:0.27769
  |
Correlation:0.27769
  |
Correlation:0.27769
  |
Correlation:0.27769
  |
Correlation:0.17078
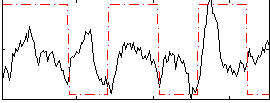  |
Component:374; Jaccard:0.08615
 |
Component:374; Jaccard:0.096504
 |
Component:374; Jaccard:0.098779
 |
Component:374; Jaccard:0.086933
 |
Component:374; Jaccard:0.067139
 |
Component:374; Jaccard:0.046524
 |
Component:286; Jaccard:0.025962
 |
Correlation:0.17078
  |
Correlation:0.17078
  |
Correlation:0.17078
  |
Correlation:0.17078
  |
Correlation:0.17078
  |
Correlation:0.17078
  |
Correlation:0.27769
  |
Component:222; Jaccard:0.083204
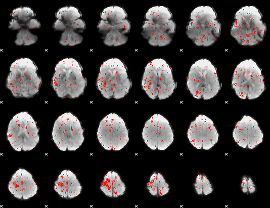 |
Component:222; Jaccard:0.089198
 |
Component:198; Jaccard:0.088129
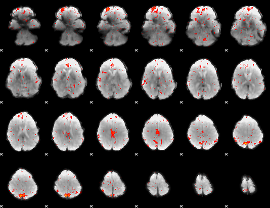 |
Component:198; Jaccard:0.080668
 |
Component:198; Jaccard:0.059115
 |
Component:198; Jaccard:0.036822
 |
Component:198; Jaccard:0.02082
 |
Correlation:0.11152
  |
Correlation:0.11152
  |
Correlation:0.067287
  |
Correlation:0.067287
  |
Correlation:0.067287
  |
Correlation:0.067287
  |
Correlation:0.067287
  |
Component:089; Jaccard:0.082852
 |
Component:089; Jaccard:0.087801
 |
Component:006; Jaccard:0.083107
 |
Component:006; Jaccard:0.075515
 |
Component:006; Jaccard:0.055346
 |
Component:006; Jaccard:0.035887
 |
Component:006; Jaccard:0.020106
 |
Correlation:-0.022229
  |
Correlation:-0.022229
  |
Correlation:0.27095
  |
Correlation:0.27095
  |
Correlation:0.27095
  |
Correlation:0.27095
  |
Correlation:0.27095
  |
Component:324; Jaccard:0.07795
 |
Component:198; Jaccard:0.081722
 |
Component:089; Jaccard:0.083084
 |
Component:089; Jaccard:0.065562
 |
Component:095; Jaccard:0.048617
 |
Component:324; Jaccard:0.02907
 |
Component:324; Jaccard:0.01698
 |
Correlation:0.18766
  |
Correlation:0.067287
  |
Correlation:-0.022229
  |
Correlation:-0.022229
  |
Correlation:0.034755
  |
Correlation:0.18766
  |
Correlation:0.18766
  |
Component:348; Jaccard:0.076888
 |
Component:324; Jaccard:0.081559
 |
Component:222; Jaccard:0.081702
 |
Component:095; Jaccard:0.064259
 |
Component:306; Jaccard:0.047926
 |
Component:095; Jaccard:0.02807
 |
Component:095; Jaccard:0.014958
 |
Correlation:0.14017
  |
Correlation:0.18766
  |
Correlation:0.11152
  |
Correlation:0.034755
  |
Correlation:0.25495
  |
Correlation:0.034755
  |
Correlation:0.034755
  |
Component:006; Jaccard:0.072701
 |
Component:006; Jaccard:0.081502
 |
Component:324; Jaccard:0.078778
 |
Component:306; Jaccard:0.063482
 |
Component:324; Jaccard:0.04461
 |
Component:306; Jaccard:0.025785
 |
Component:306; Jaccard:0.013994
 |
Correlation:0.27095
  |
Correlation:0.27095
  |
Correlation:0.18766
  |
Correlation:0.25495
  |
Correlation:0.18766
  |
Correlation:0.25495
  |
Correlation:0.25495
  |
Component:306; Jaccard:0.071633
 |
Component:348; Jaccard:0.078898
 |
Component:306; Jaccard:0.076632
 |
Component:324; Jaccard:0.062267
 |
Component:089; Jaccard:0.040999
 |
Component:331; Jaccard:0.023885
 |
Component:331; Jaccard:0.013856
 |
Correlation:0.25495
  |
Correlation:0.14017
  |
Correlation:0.25495
  |
Correlation:0.18766
  |
Correlation:-0.022229
  |
Correlation:0.29343
  |
Correlation:0.29343
  |
Component:315; Jaccard:0.071012
 |
Component:306; Jaccard:0.075658
 |
Component:348; Jaccard:0.075729
 |
Component:222; Jaccard:0.062028
 |
Component:331; Jaccard:0.040978
 |
Component:089; Jaccard:0.023469
 |
Component:089; Jaccard:0.013293
 |
Correlation:-0.1376
  |
Correlation:0.25495
  |
Correlation:0.14017
  |
Correlation:0.11152
  |
Correlation:0.29343
  |
Correlation:-0.022229
  |
Correlation:-0.022229
  |
Cope: 05:neg_SHAPES BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:217; Jaccard:0.088942
 |
Component:217; Jaccard:0.10683
 |
Component:217; Jaccard:0.12
 |
Component:095; Jaccard:0.13231
 |
Component:095; Jaccard:0.12855
 |
Component:095; Jaccard:0.096925
 |
Component:095; Jaccard:0.056353
 |
Correlation:0.36464
  |
Correlation:0.36464
  |
Correlation:0.36464
  |
Correlation:0.08697
  |
Correlation:0.08697
  |
Correlation:0.08697
  |
Correlation:0.08697
  |
Component:383; Jaccard:0.086401
 |
Component:383; Jaccard:0.10252
 |
Component:383; Jaccard:0.11557
 |
Component:217; Jaccard:0.11651
 |
Component:217; Jaccard:0.094133
 |
Component:217; Jaccard:0.064139
 |
Component:217; Jaccard:0.037891
 |
Correlation:0.39791
  |
Correlation:0.39791
  |
Correlation:0.39791
  |
Correlation:0.36464
  |
Correlation:0.36464
  |
Correlation:0.36464
  |
Correlation:0.36464
  |
Component:305; Jaccard:0.084668
 |
Component:071; Jaccard:0.097658
 |
Component:095; Jaccard:0.11543
 |
Component:383; Jaccard:0.11229
 |
Component:383; Jaccard:0.093972
 |
Component:383; Jaccard:0.061156
 |
Component:383; Jaccard:0.035494
 |
Correlation:0.12094
  |
Correlation:0.24166
  |
Correlation:0.08697
  |
Correlation:0.39791
  |
Correlation:0.39791
  |
Correlation:0.39791
  |
Correlation:0.39791
  |
Component:071; Jaccard:0.084139
 |
Component:089; Jaccard:0.094336
 |
Component:071; Jaccard:0.10774
 |
Component:071; Jaccard:0.10612
 |
Component:071; Jaccard:0.090578
 |
Component:071; Jaccard:0.058576
 |
Component:071; Jaccard:0.0341
 |
Correlation:0.24166
  |
Correlation:0.12941
  |
Correlation:0.24166
  |
Correlation:0.24166
  |
Correlation:0.24166
  |
Correlation:0.24166
  |
Correlation:0.24166
  |
Component:089; Jaccard:0.082794
 |
Component:095; Jaccard:0.093564
 |
Component:089; Jaccard:0.10042
 |
Component:168; Jaccard:0.095674
 |
Component:089; Jaccard:0.075091
 |
Component:089; Jaccard:0.049122
 |
Component:089; Jaccard:0.027844
 |
Correlation:0.12941
  |
Correlation:0.08697
  |
Correlation:0.12941
  |
Correlation:0.18192
  |
Correlation:0.12941
  |
Correlation:0.12941
  |
Correlation:0.12941
  |
Component:168; Jaccard:0.079915
 |
Component:305; Jaccard:0.091808
 |
Component:168; Jaccard:0.098989
 |
Component:089; Jaccard:0.095654
 |
Component:168; Jaccard:0.072863
 |
Component:168; Jaccard:0.047352
 |
Component:168; Jaccard:0.02658
 |
Correlation:0.18192
  |
Correlation:0.12094
  |
Correlation:0.18192
  |
Correlation:0.12941
  |
Correlation:0.18192
  |
Correlation:0.18192
  |
Correlation:0.18192
  |
Component:315; Jaccard:0.079607
 |
Component:168; Jaccard:0.091547
 |
Component:305; Jaccard:0.092704
 |
Component:090; Jaccard:0.086489
 |
Component:090; Jaccard:0.071352
 |
Component:090; Jaccard:0.046163
 |
Component:090; Jaccard:0.025613
 |
Correlation:0.096414
  |
Correlation:0.18192
  |
Correlation:0.12094
  |
Correlation:0.22049
  |
Correlation:0.22049
  |
Correlation:0.22049
  |
Correlation:0.22049
  |
Component:286; Jaccard:0.076073
 |
Component:115; Jaccard:0.085255
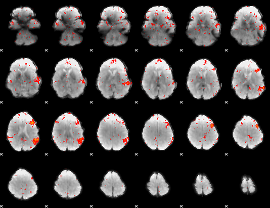 |
Component:115; Jaccard:0.08879
 |
Component:073; Jaccard:0.080506
 |
Component:073; Jaccard:0.067835
 |
Component:073; Jaccard:0.043136
 |
Component:374; Jaccard:0.024898
 |
Correlation:-0.18718
  |
Correlation:0.40378
  |
Correlation:0.40378
  |
Correlation:0.07493
  |
Correlation:0.07493
  |
Correlation:0.07493
  |
Correlation:-0.35625
  |
Component:266; Jaccard:0.075579
 |
Component:315; Jaccard:0.084756
 |
Component:090; Jaccard:0.088414
 |
Component:286; Jaccard:0.079757
 |
Component:198; Jaccard:0.062173
 |
Component:374; Jaccard:0.041025
 |
Component:006; Jaccard:0.023505
 |
Correlation:0.34018
  |
Correlation:0.096414
  |
Correlation:0.22049
  |
Correlation:-0.18718
  |
Correlation:-0.12749
  |
Correlation:-0.35625
  |
Correlation:-0.11864
  |
Component:374; Jaccard:0.075231
 |
Component:286; Jaccard:0.08385
 |
Component:374; Jaccard:0.086696
 |
Component:374; Jaccard:0.079079
 |
Component:286; Jaccard:0.06199
 |
Component:198; Jaccard:0.040979
 |
Component:198; Jaccard:0.023304
 |
Correlation:-0.35625
  |
Correlation:-0.18718
  |
Correlation:-0.35625
  |
Correlation:-0.35625
  |
Correlation:-0.18718
  |
Correlation:-0.12749
  |
Correlation:-0.12749
  |
Cope: 06:SHAPES-FACES BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:337; Jaccard:0.12473
 |
Component:337; Jaccard:0.15482
 |
Component:337; Jaccard:0.17327
 |
Component:337; Jaccard:0.16007
 |
Component:337; Jaccard:0.11711
 |
Component:337; Jaccard:0.070597
 |
Component:337; Jaccard:0.038058
 |
Correlation:0.35734
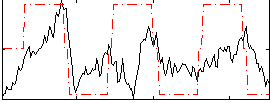  |
Correlation:0.35734
  |
Correlation:0.35734
  |
Correlation:0.35734
  |
Correlation:0.35734
  |
Correlation:0.35734
  |
Correlation:0.35734
  |
Component:078; Jaccard:0.10227
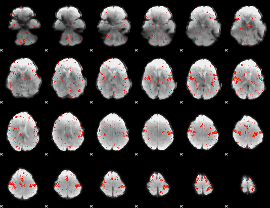 |
Component:389; Jaccard:0.11773
 |
Component:389; Jaccard:0.12401
 |
Component:389; Jaccard:0.10346
 |
Component:078; Jaccard:0.075251
 |
Component:078; Jaccard:0.04745
 |
Component:078; Jaccard:0.027357
 |
Correlation:0.23318
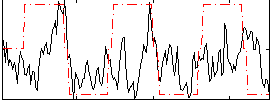 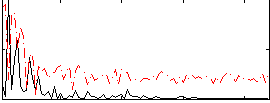 |
Correlation:0.19659
  |
Correlation:0.19659
  |
Correlation:0.19659
  |
Correlation:0.23318
  |
Correlation:0.23318
  |
Correlation:0.23318
  |
Component:389; Jaccard:0.10182
 |
Component:078; Jaccard:0.11456
 |
Component:078; Jaccard:0.11744
 |
Component:078; Jaccard:0.10269
 |
Component:132; Jaccard:0.073638
 |
Component:132; Jaccard:0.044999
 |
Component:389; Jaccard:0.021764
 |
Correlation:0.19659
  |
Correlation:0.23318
  |
Correlation:0.23318
  |
Correlation:0.23318
  |
Correlation:0.51089
  |
Correlation:0.51089
  |
Correlation:0.19659
  |
Component:132; Jaccard:0.098564
 |
Component:132; Jaccard:0.11263
 |
Component:132; Jaccard:0.11738
 |
Component:132; Jaccard:0.10163
 |
Component:389; Jaccard:0.073218
 |
Component:389; Jaccard:0.043389
 |
Component:132; Jaccard:0.02173
 |
Correlation:0.51089
  |
Correlation:0.51089
  |
Correlation:0.51089
  |
Correlation:0.51089
  |
Correlation:0.19659
  |
Correlation:0.19659
  |
Correlation:0.51089
  |
Component:118; Jaccard:0.09012
 |
Component:118; Jaccard:0.1007
 |
Component:118; Jaccard:0.10209
 |
Component:118; Jaccard:0.087899
 |
Component:118; Jaccard:0.061401
 |
Component:118; Jaccard:0.036551
 |
Component:118; Jaccard:0.019696
 |
Correlation:0.35991
  |
Correlation:0.35991
  |
Correlation:0.35991
  |
Correlation:0.35991
  |
Correlation:0.35991
  |
Correlation:0.35991
  |
Correlation:0.35991
  |
Component:348; Jaccard:0.085457
 |
Component:322; Jaccard:0.091348
 |
Component:322; Jaccard:0.091656
 |
Component:322; Jaccard:0.077011
 |
Component:322; Jaccard:0.057221
 |
Component:322; Jaccard:0.035808
 |
Component:358; Jaccard:0.01847
 |
Correlation:0.20786
  |
Correlation:0.23171
  |
Correlation:0.23171
  |
Correlation:0.23171
  |
Correlation:0.23171
  |
Correlation:0.23171
  |
Correlation:0.31382
  |
Component:322; Jaccard:0.083075
 |
Component:348; Jaccard:0.087862
 |
Component:348; Jaccard:0.081974
 |
Component:358; Jaccard:0.074026
 |
Component:358; Jaccard:0.052676
 |
Component:358; Jaccard:0.03315
 |
Component:137; Jaccard:0.017223
 |
Correlation:0.23171
  |
Correlation:0.20786
  |
Correlation:0.20786
  |
Correlation:0.31382
  |
Correlation:0.31382
  |
Correlation:0.31382
  |
Correlation:0.13277
  |
Component:307; Jaccard:0.076945
 |
Component:358; Jaccard:0.083434
 |
Component:358; Jaccard:0.08057
 |
Component:348; Jaccard:0.068013
 |
Component:348; Jaccard:0.043812
 |
Component:137; Jaccard:0.028348
 |
Component:322; Jaccard:0.016632
 |
Correlation:0.32812
  |
Correlation:0.31382
  |
Correlation:0.31382
  |
Correlation:0.20786
  |
Correlation:0.20786
  |
Correlation:0.13277
  |
Correlation:0.23171
  |
Component:358; Jaccard:0.07479
 |
Component:307; Jaccard:0.081503
 |
Component:307; Jaccard:0.078601
 |
Component:307; Jaccard:0.063511
 |
Component:307; Jaccard:0.042786
 |
Component:348; Jaccard:0.025789
 |
Component:242; Jaccard:0.015515
 |
Correlation:0.31382
  |
Correlation:0.32812
  |
Correlation:0.32812
  |
Correlation:0.32812
  |
Correlation:0.32812
  |
Correlation:0.20786
  |
Correlation:0.030278
  |
Component:286; Jaccard:0.073555
 |
Component:286; Jaccard:0.078578
 |
Component:286; Jaccard:0.075758
 |
Component:331; Jaccard:0.060268
 |
Component:331; Jaccard:0.04
 |
Component:242; Jaccard:0.02443
 |
Component:324; Jaccard:0.01367
 |
Correlation:0.25215
  |
Correlation:0.25215
  |
Correlation:0.25215
  |
Correlation:0.2561
  |
Correlation:0.2561
  |
Correlation:0.030278
  |
Correlation:0.066953
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































