Task name: RELATIONAL, TR=0.72, totally 232 volumes, 10 waves, 6 copes BACK
|
Cope: 01:MATCH BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
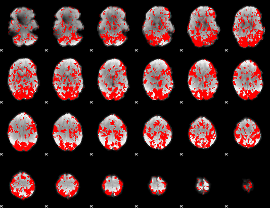 |
GLM binary result thresholded by 1.5
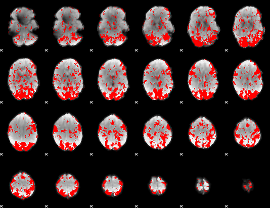 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
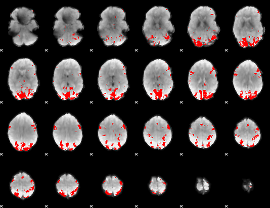 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
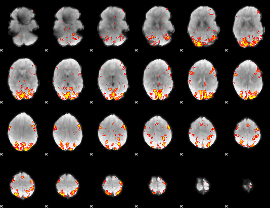 |
GLM Z-score result thresholded by 3.5
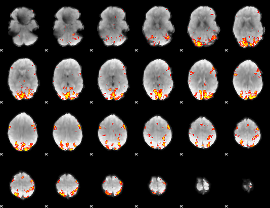 |
GLM Z-score result thresholded by 4
 |
Component:010; Jaccard:0.13592
 |
Component:010; Jaccard:0.17021
 |
Component:010; Jaccard:0.20679
 |
Component:010; Jaccard:0.24024
 |
Component:010; Jaccard:0.26034
 |
Component:010; Jaccard:0.26584
 |
Component:010; Jaccard:0.25932
 |
Correlation:0.2946
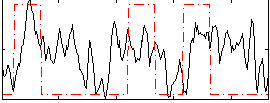 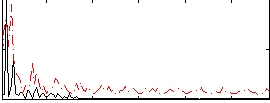 |
Correlation:0.2946
  |
Correlation:0.2946
  |
Correlation:0.2946
  |
Correlation:0.2946
  |
Correlation:0.2946
  |
Correlation:0.2946
  |
Component:260; Jaccard:0.13376
 |
Component:260; Jaccard:0.15686
 |
Component:257; Jaccard:0.18425
 |
Component:257; Jaccard:0.21017
 |
Component:257; Jaccard:0.225
 |
Component:257; Jaccard:0.22668
 |
Component:303; Jaccard:0.22275
 |
Correlation:0.13116
  |
Correlation:0.13116
  |
Correlation:0.15775
  |
Correlation:0.15775
  |
Correlation:0.15775
  |
Correlation:0.15775
  |
Correlation:0.26508
  |
Component:171; Jaccard:0.12545
 |
Component:257; Jaccard:0.15462
 |
Component:395; Jaccard:0.17974
 |
Component:395; Jaccard:0.20399
 |
Component:395; Jaccard:0.21795
 |
Component:303; Jaccard:0.22548
 |
Component:171; Jaccard:0.22181
 |
Correlation:0.1216
  |
Correlation:0.15775
  |
Correlation:0.23733
 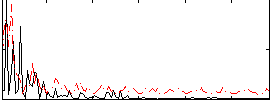 |
Correlation:0.23733
  |
Correlation:0.23733
  |
Correlation:0.26508
  |
Correlation:0.1216
  |
Component:257; Jaccard:0.12497
 |
Component:171; Jaccard:0.1518
 |
Component:171; Jaccard:0.17891
 |
Component:171; Jaccard:0.20302
 |
Component:378; Jaccard:0.21742
 |
Component:171; Jaccard:0.22483
 |
Component:257; Jaccard:0.21964
 |
Correlation:0.15775
  |
Correlation:0.1216
  |
Correlation:0.1216
  |
Correlation:0.1216
  |
Correlation:0.006221
  |
Correlation:0.1216
  |
Correlation:0.15775
  |
Component:395; Jaccard:0.1249
 |
Component:395; Jaccard:0.15106
 |
Component:260; Jaccard:0.17661
 |
Component:303; Jaccard:0.19954
 |
Component:171; Jaccard:0.21717
 |
Component:395; Jaccard:0.22294
 |
Component:378; Jaccard:0.2177
 |
Correlation:0.23733
  |
Correlation:0.23733
  |
Correlation:0.13116
  |
Correlation:0.26508
  |
Correlation:0.1216
  |
Correlation:0.23733
  |
Correlation:0.006221
  |
Component:303; Jaccard:0.12075
 |
Component:303; Jaccard:0.14642
 |
Component:303; Jaccard:0.17375
 |
Component:378; Jaccard:0.1987
 |
Component:303; Jaccard:0.21677
 |
Component:378; Jaccard:0.22246
 |
Component:342; Jaccard:0.2118
 |
Correlation:0.26508
  |
Correlation:0.26508
  |
Correlation:0.26508
  |
Correlation:0.006221
  |
Correlation:0.26508
  |
Correlation:0.006221
  |
Correlation:0.23577
  |
Component:378; Jaccard:0.11756
 |
Component:378; Jaccard:0.14406
 |
Component:378; Jaccard:0.17256
 |
Component:342; Jaccard:0.19549
 |
Component:342; Jaccard:0.21247
 |
Component:342; Jaccard:0.21681
 |
Component:395; Jaccard:0.21074
 |
Correlation:0.006221
  |
Correlation:0.006221
  |
Correlation:0.006221
  |
Correlation:0.23577
  |
Correlation:0.23577
  |
Correlation:0.23577
  |
Correlation:0.23733
  |
Component:342; Jaccard:0.11701
 |
Component:342; Jaccard:0.14294
 |
Component:342; Jaccard:0.17003
 |
Component:260; Jaccard:0.18832
 |
Component:260; Jaccard:0.19116
 |
Component:260; Jaccard:0.18419
 |
Component:260; Jaccard:0.17097
 |
Correlation:0.23577
  |
Correlation:0.23577
  |
Correlation:0.23577
  |
Correlation:0.13116
  |
Correlation:0.13116
  |
Correlation:0.13116
  |
Correlation:0.13116
  |
Component:029; Jaccard:0.10903
 |
Component:029; Jaccard:0.12705
 |
Component:029; Jaccard:0.13891
 |
Component:097; Jaccard:0.14339
 |
Component:097; Jaccard:0.1507
 |
Component:097; Jaccard:0.15426
 |
Component:097; Jaccard:0.14733
 |
Correlation:0.19918
  |
Correlation:0.19918
  |
Correlation:0.19918
  |
Correlation:0.22361
  |
Correlation:0.22361
  |
Correlation:0.22361
  |
Correlation:0.22361
  |
Component:097; Jaccard:0.09508
 |
Component:097; Jaccard:0.11149
 |
Component:097; Jaccard:0.12793
 |
Component:029; Jaccard:0.14181
 |
Component:029; Jaccard:0.13709
 |
Component:231; Jaccard:0.12937
 |
Component:231; Jaccard:0.12397
 |
Correlation:0.22361
  |
Correlation:0.22361
  |
Correlation:0.22361
  |
Correlation:0.19918
  |
Correlation:0.19918
  |
Correlation:0.16009
  |
Correlation:0.16009
  |
Cope: 02:REL BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
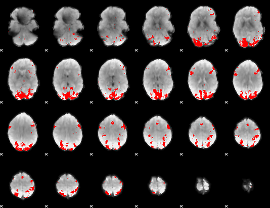 |
GLM binary result thresholded by 4
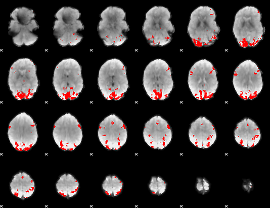 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:171; Jaccard:0.14252
 |
Component:257; Jaccard:0.177
 |
Component:257; Jaccard:0.21185
 |
Component:378; Jaccard:0.23967
 |
Component:378; Jaccard:0.25817
 |
Component:378; Jaccard:0.26394
 |
Component:378; Jaccard:0.26205
 |
Correlation:0.17917
  |
Correlation:0.16487
  |
Correlation:0.16487
  |
Correlation:0.19494
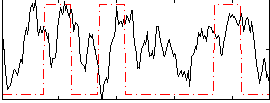 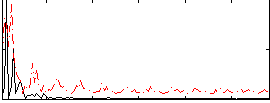 |
Correlation:0.19494
  |
Correlation:0.19494
  |
Correlation:0.19494
  |
Component:378; Jaccard:0.14166
 |
Component:378; Jaccard:0.17668
 |
Component:378; Jaccard:0.21053
 |
Component:257; Jaccard:0.23427
 |
Component:257; Jaccard:0.24832
 |
Component:257; Jaccard:0.24699
 |
Component:171; Jaccard:0.24515
 |
Correlation:0.19494
  |
Correlation:0.19494
  |
Correlation:0.19494
  |
Correlation:0.16487
  |
Correlation:0.16487
  |
Correlation:0.16487
  |
Correlation:0.17917
  |
Component:257; Jaccard:0.14117
 |
Component:171; Jaccard:0.17385
 |
Component:171; Jaccard:0.20363
 |
Component:171; Jaccard:0.22744
 |
Component:171; Jaccard:0.24106
 |
Component:171; Jaccard:0.24658
 |
Component:257; Jaccard:0.2357
 |
Correlation:0.16487
  |
Correlation:0.17917
  |
Correlation:0.17917
  |
Correlation:0.17917
  |
Correlation:0.17917
  |
Correlation:0.17917
  |
Correlation:0.16487
  |
Component:010; Jaccard:0.13694
 |
Component:010; Jaccard:0.1672
 |
Component:010; Jaccard:0.19725
 |
Component:010; Jaccard:0.22168
 |
Component:010; Jaccard:0.23297
 |
Component:010; Jaccard:0.23107
 |
Component:342; Jaccard:0.22818
 |
Correlation:0.075278
  |
Correlation:0.075278
  |
Correlation:0.075278
  |
Correlation:0.075278
  |
Correlation:0.075278
  |
Correlation:0.075278
  |
Correlation:0.18814
 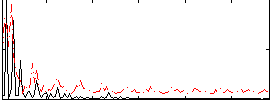 |
Component:260; Jaccard:0.13327
 |
Component:395; Jaccard:0.15959
 |
Component:342; Jaccard:0.18518
 |
Component:342; Jaccard:0.20791
 |
Component:342; Jaccard:0.22284
 |
Component:342; Jaccard:0.22985
 |
Component:010; Jaccard:0.21977
 |
Correlation:0.040894
  |
Correlation:0.022831
 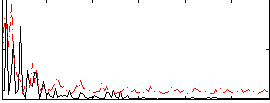 |
Correlation:0.18814
  |
Correlation:0.18814
  |
Correlation:0.18814
  |
Correlation:0.18814
  |
Correlation:0.075278
  |
Component:395; Jaccard:0.13196
 |
Component:342; Jaccard:0.15865
 |
Component:395; Jaccard:0.18467
 |
Component:395; Jaccard:0.20675
 |
Component:395; Jaccard:0.21685
 |
Component:395; Jaccard:0.21945
 |
Component:303; Jaccard:0.21452
 |
Correlation:0.022831
  |
Correlation:0.18814
  |
Correlation:0.022831
  |
Correlation:0.022831
  |
Correlation:0.022831
  |
Correlation:0.022831
  |
Correlation:0.04693
  |
Component:342; Jaccard:0.12901
 |
Component:303; Jaccard:0.15582
 |
Component:303; Jaccard:0.1822
 |
Component:303; Jaccard:0.20275
 |
Component:303; Jaccard:0.21615
 |
Component:303; Jaccard:0.21916
 |
Component:395; Jaccard:0.20966
 |
Correlation:0.18814
  |
Correlation:0.04693
  |
Correlation:0.04693
  |
Correlation:0.04693
  |
Correlation:0.04693
  |
Correlation:0.04693
  |
Correlation:0.022831
  |
Component:303; Jaccard:0.12776
 |
Component:260; Jaccard:0.153
 |
Component:260; Jaccard:0.16907
 |
Component:097; Jaccard:0.17791
 |
Component:097; Jaccard:0.19073
 |
Component:097; Jaccard:0.19225
 |
Component:097; Jaccard:0.18425
 |
Correlation:0.04693
  |
Correlation:0.040894
  |
Correlation:0.040894
  |
Correlation:0.3582
  |
Correlation:0.3582
  |
Correlation:0.3582
  |
Correlation:0.3582
  |
Component:029; Jaccard:0.1212
 |
Component:029; Jaccard:0.14392
 |
Component:097; Jaccard:0.15777
 |
Component:260; Jaccard:0.17562
 |
Component:260; Jaccard:0.17404
 |
Component:260; Jaccard:0.16394
 |
Component:260; Jaccard:0.15161
 |
Correlation:0.19644
 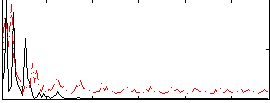 |
Correlation:0.19644
  |
Correlation:0.3582
  |
Correlation:0.040894
  |
Correlation:0.040894
  |
Correlation:0.040894
  |
Correlation:0.040894
  |
Component:097; Jaccard:0.11289
 |
Component:097; Jaccard:0.13485
 |
Component:029; Jaccard:0.15764
 |
Component:029; Jaccard:0.16075
 |
Component:029; Jaccard:0.15296
 |
Component:029; Jaccard:0.13857
 |
Component:231; Jaccard:0.12985
 |
Correlation:0.3582
  |
Correlation:0.3582
  |
Correlation:0.19644
  |
Correlation:0.19644
  |
Correlation:0.19644
  |
Correlation:0.19644
  |
Correlation:0.12362
  |
Cope: 03:MATCH-REL BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
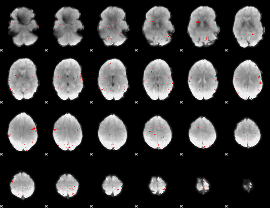 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
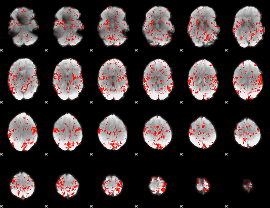 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
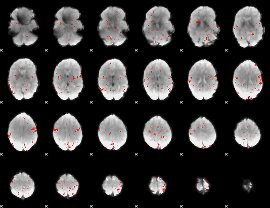 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:359; Jaccard:0.099136
 |
Component:359; Jaccard:0.10654
 |
Component:359; Jaccard:0.097611
 |
Component:359; Jaccard:0.066561
 |
Component:359; Jaccard:0.032933
 |
Component:359; Jaccard:0.012928
 |
Component:359; Jaccard:0.003705
 |
Correlation:0.30621
  |
Correlation:0.30621
  |
Correlation:0.30621
  |
Correlation:0.30621
  |
Correlation:0.30621
  |
Correlation:0.30621
  |
Correlation:0.30621
  |
Component:327; Jaccard:0.09075
 |
Component:327; Jaccard:0.099171
 |
Component:327; Jaccard:0.09202
 |
Component:327; Jaccard:0.062218
 |
Component:327; Jaccard:0.030486
 |
Component:327; Jaccard:0.010082
 |
Component:327; Jaccard:0.003411
 |
Correlation:0.16747
  |
Correlation:0.16747
  |
Correlation:0.16747
  |
Correlation:0.16747
  |
Correlation:0.16747
  |
Correlation:0.16747
  |
Correlation:0.16747
  |
Component:373; Jaccard:0.08779
 |
Component:199; Jaccard:0.090466
 |
Component:199; Jaccard:0.085111
 |
Component:199; Jaccard:0.053726
 |
Component:008; Jaccard:0.026213
 |
Component:199; Jaccard:0.007951
 |
Component:176; Jaccard:0.002595
 |
Correlation:0.06218
  |
Correlation:0.077308
  |
Correlation:0.077308
  |
Correlation:0.077308
  |
Correlation:-0.006142
  |
Correlation:0.077308
  |
Correlation:0.15859
  |
Component:199; Jaccard:0.08281
 |
Component:373; Jaccard:0.089173
 |
Component:176; Jaccard:0.081612
 |
Component:013; Jaccard:0.052701
 |
Component:176; Jaccard:0.024368
 |
Component:176; Jaccard:0.007623
 |
Component:325; Jaccard:0.002439
 |
Correlation:0.077308
  |
Correlation:0.06218
  |
Correlation:0.15859
  |
Correlation:0.30167
  |
Correlation:0.15859
  |
Correlation:0.15859
  |
Correlation:0.10489
  |
Component:211; Jaccard:0.080405
 |
Component:013; Jaccard:0.088166
 |
Component:013; Jaccard:0.080125
 |
Component:176; Jaccard:0.052057
 |
Component:199; Jaccard:0.022606
 |
Component:013; Jaccard:0.007069
 |
Component:010; Jaccard:0.002387
 |
Correlation:0.026024
  |
Correlation:0.30167
  |
Correlation:0.30167
  |
Correlation:0.15859
  |
Correlation:0.077308
  |
Correlation:0.30167
  |
Correlation:0.12999
  |
Component:013; Jaccard:0.080234
 |
Component:176; Jaccard:0.08812
 |
Component:008; Jaccard:0.078035
 |
Component:008; Jaccard:0.051552
 |
Component:013; Jaccard:0.022226
 |
Component:008; Jaccard:0.006723
 |
Component:199; Jaccard:0.002257
 |
Correlation:0.30167
  |
Correlation:0.15859
  |
Correlation:-0.006142
  |
Correlation:-0.006142
  |
Correlation:0.30167
  |
Correlation:-0.006142
  |
Correlation:0.077308
  |
Component:176; Jaccard:0.077984
 |
Component:211; Jaccard:0.080914
 |
Component:315; Jaccard:0.072217
 |
Component:315; Jaccard:0.044649
 |
Component:315; Jaccard:0.018227
 |
Component:345; Jaccard:0.006324
 |
Component:008; Jaccard:0.002146
 |
Correlation:0.15859
  |
Correlation:0.026024
  |
Correlation:0.096062
  |
Correlation:0.096062
  |
Correlation:0.096062
  |
Correlation:0.1155
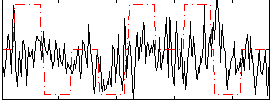  |
Correlation:-0.006142
  |
Component:136; Jaccard:0.074992
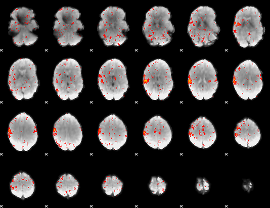 |
Component:315; Jaccard:0.079471
 |
Component:136; Jaccard:0.070569
 |
Component:211; Jaccard:0.043881
 |
Component:211; Jaccard:0.017181
 |
Component:211; Jaccard:0.006032
 |
Component:118; Jaccard:0.002036
 |
Correlation:0.11716
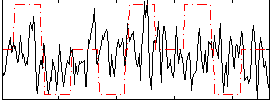  |
Correlation:0.096062
  |
Correlation:0.11716
  |
Correlation:0.026024
  |
Correlation:0.026024
  |
Correlation:0.026024
  |
Correlation:0.099996
  |
Component:234; Jaccard:0.074543
 |
Component:136; Jaccard:0.07922
 |
Component:211; Jaccard:0.069217
 |
Component:136; Jaccard:0.039772
 |
Component:136; Jaccard:0.016849
 |
Component:010; Jaccard:0.00586
 |
Component:272; Jaccard:0.001885
 |
Correlation:0.016077
  |
Correlation:0.11716
  |
Correlation:0.026024
  |
Correlation:0.11716
  |
Correlation:0.11716
  |
Correlation:0.12999
  |
Correlation:0.0016531
  |
Component:315; Jaccard:0.073661
 |
Component:008; Jaccard:0.079013
 |
Component:373; Jaccard:0.067059
 |
Component:373; Jaccard:0.035035
 |
Component:069; Jaccard:0.016109
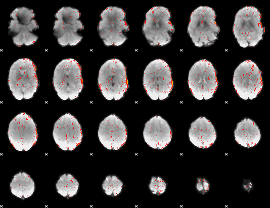 |
Component:118; Jaccard:0.005672
 |
Component:013; Jaccard:0.001855
 |
Correlation:0.096062
  |
Correlation:-0.006142
  |
Correlation:0.06218
  |
Correlation:0.06218
  |
Correlation:0.18584
  |
Correlation:0.099996
  |
Correlation:0.30167
  |
Cope: 04:REL-MATCH BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
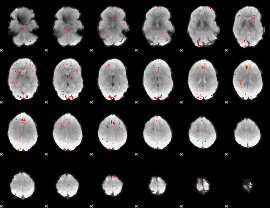 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
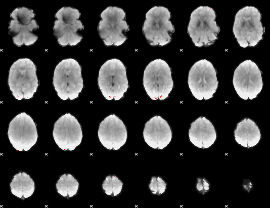 |
GLM Z-score result thresholded by 1
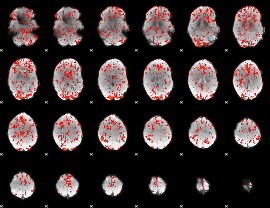 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
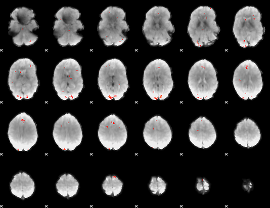 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:091; Jaccard:0.097415
 |
Component:378; Jaccard:0.099201
 |
Component:378; Jaccard:0.086691
 |
Component:378; Jaccard:0.063135
 |
Component:378; Jaccard:0.037452
 |
Component:378; Jaccard:0.021671
 |
Component:378; Jaccard:0.010016
 |
Correlation:0.21405
  |
Correlation:0.11186
  |
Correlation:0.11186
  |
Correlation:0.11186
  |
Correlation:0.11186
  |
Correlation:0.11186
  |
Correlation:0.11186
  |
Component:378; Jaccard:0.095014
 |
Component:091; Jaccard:0.092909
 |
Component:097; Jaccard:0.072901
 |
Component:097; Jaccard:0.05607
 |
Component:097; Jaccard:0.035481
 |
Component:097; Jaccard:0.020928
 |
Component:097; Jaccard:0.009808
 |
Correlation:0.11186
  |
Correlation:0.21405
  |
Correlation:0.079769
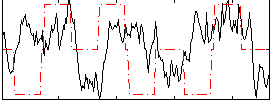 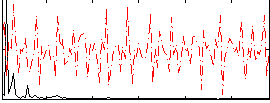 |
Correlation:0.079769
  |
Correlation:0.079769
  |
Correlation:0.079769
  |
Correlation:0.079769
  |
Component:207; Jaccard:0.088814
 |
Component:097; Jaccard:0.087355
 |
Component:207; Jaccard:0.066282
 |
Component:363; Jaccard:0.046953
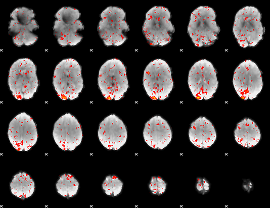 |
Component:363; Jaccard:0.027
 |
Component:363; Jaccard:0.01476
 |
Component:363; Jaccard:0.007506
 |
Correlation:0.1582
  |
Correlation:0.079769
  |
Correlation:0.1582
  |
Correlation:0.077142
 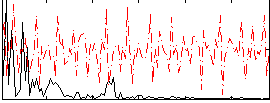 |
Correlation:0.077142
  |
Correlation:0.077142
  |
Correlation:0.077142
  |
Component:097; Jaccard:0.082825
 |
Component:207; Jaccard:0.084269
 |
Component:363; Jaccard:0.065368
 |
Component:171; Jaccard:0.043255
 |
Component:171; Jaccard:0.025092
 |
Component:171; Jaccard:0.012508
 |
Component:171; Jaccard:0.005993
 |
Correlation:0.079769
  |
Correlation:0.1582
  |
Correlation:0.077142
  |
Correlation:0.034119
  |
Correlation:0.034119
  |
Correlation:0.034119
  |
Correlation:0.034119
  |
Component:363; Jaccard:0.081243
 |
Component:363; Jaccard:0.07922
 |
Component:171; Jaccard:0.063078
 |
Component:207; Jaccard:0.037029
 |
Component:273; Jaccard:0.019109
 |
Component:273; Jaccard:0.008601
 |
Component:257; Jaccard:0.004104
 |
Correlation:0.077142
  |
Correlation:0.077142
  |
Correlation:0.034119
  |
Correlation:0.1582
  |
Correlation:0.28021
  |
Correlation:0.28021
  |
Correlation:0.0042201
  |
Component:250; Jaccard:0.078201
 |
Component:171; Jaccard:0.076657
 |
Component:091; Jaccard:0.055902
 |
Component:273; Jaccard:0.034014
 |
Component:207; Jaccard:0.017423
 |
Component:257; Jaccard:0.007825
 |
Component:273; Jaccard:0.003468
 |
Correlation:0.1967
  |
Correlation:0.034119
  |
Correlation:0.21405
  |
Correlation:0.28021
  |
Correlation:0.1582
  |
Correlation:0.0042201
  |
Correlation:0.28021
  |
Component:171; Jaccard:0.074394
 |
Component:250; Jaccard:0.073141
 |
Component:250; Jaccard:0.054889
 |
Component:216; Jaccard:0.027774
 |
Component:257; Jaccard:0.014584
 |
Component:342; Jaccard:0.006092
 |
Component:303; Jaccard:0.002853
 |
Correlation:0.034119
  |
Correlation:0.1967
  |
Correlation:0.1967
  |
Correlation:0.10123
  |
Correlation:0.0042201
  |
Correlation:-0.028233
  |
Correlation:-0.1293
  |
Component:273; Jaccard:0.074116
 |
Component:281; Jaccard:0.072197
 |
Component:273; Jaccard:0.053876
 |
Component:257; Jaccard:0.027657
 |
Component:231; Jaccard:0.014075
 |
Component:231; Jaccard:0.006059
 |
Component:331; Jaccard:0.002785
 |
Correlation:0.28021
  |
Correlation:-0.022715
  |
Correlation:0.28021
  |
Correlation:0.0042201
  |
Correlation:-0.021613
  |
Correlation:-0.021613
  |
Correlation:0.051229
  |
Component:281; Jaccard:0.074042
 |
Component:273; Jaccard:0.069294
 |
Component:281; Jaccard:0.051606
 |
Component:250; Jaccard:0.027066
 |
Component:342; Jaccard:0.01326
 |
Component:207; Jaccard:0.005851
 |
Component:037; Jaccard:0.002068
 |
Correlation:-0.022715
  |
Correlation:0.28021
  |
Correlation:-0.022715
  |
Correlation:0.1967
  |
Correlation:-0.028233
  |
Correlation:0.1582
  |
Correlation:0.13578
  |
Component:100; Jaccard:0.070823
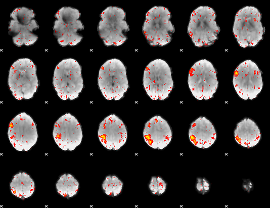 |
Component:100; Jaccard:0.061944
 |
Component:190; Jaccard:0.045668
 |
Component:342; Jaccard:0.025923
 |
Component:250; Jaccard:0.011221
 |
Component:303; Jaccard:0.005843
 |
Component:342; Jaccard:0.001858
 |
Correlation:-0.010962
  |
Correlation:-0.010962
  |
Correlation:0.30295
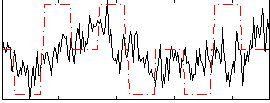 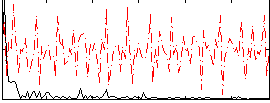 |
Correlation:-0.028233
  |
Correlation:0.1967
  |
Correlation:-0.1293
  |
Correlation:-0.028233
  |
Cope: 05:neg_MATCH BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
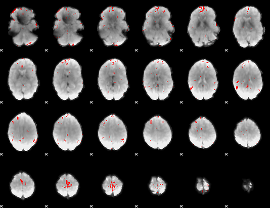 |
GLM binary result thresholded by 4
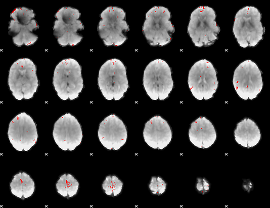 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:313; Jaccard:0.10055
 |
Component:313; Jaccard:0.11558
 |
Component:313; Jaccard:0.12381
 |
Component:113; Jaccard:0.12752
 |
Component:113; Jaccard:0.11178
 |
Component:113; Jaccard:0.081179
 |
Component:113; Jaccard:0.049357
 |
Correlation:0.031642
  |
Correlation:0.031642
  |
Correlation:0.031642
  |
Correlation:0.17624
  |
Correlation:0.17624
  |
Correlation:0.17624
  |
Correlation:0.17624
  |
Component:397; Jaccard:0.095287
 |
Component:397; Jaccard:0.10492
 |
Component:113; Jaccard:0.11999
 |
Component:313; Jaccard:0.11912
 |
Component:063; Jaccard:0.096226
 |
Component:063; Jaccard:0.067287
 |
Component:063; Jaccard:0.040767
 |
Correlation:0.1686
  |
Correlation:0.1686
  |
Correlation:0.17624
  |
Correlation:0.031642
  |
Correlation:0.22258
  |
Correlation:0.22258
  |
Correlation:0.22258
  |
Component:054; Jaccard:0.087458
 |
Component:054; Jaccard:0.10347
 |
Component:054; Jaccard:0.1143
 |
Component:054; Jaccard:0.11518
 |
Component:313; Jaccard:0.095419
 |
Component:313; Jaccard:0.064491
 |
Component:313; Jaccard:0.035596
 |
Correlation:-0.019148
  |
Correlation:-0.019148
  |
Correlation:-0.019148
  |
Correlation:-0.019148
  |
Correlation:0.031642
  |
Correlation:0.031642
  |
Correlation:0.031642
  |
Component:027; Jaccard:0.087265
 |
Component:113; Jaccard:0.10326
 |
Component:397; Jaccard:0.10526
 |
Component:063; Jaccard:0.10785
 |
Component:054; Jaccard:0.094674
 |
Component:054; Jaccard:0.061381
 |
Component:344; Jaccard:0.033469
 |
Correlation:0.24559
  |
Correlation:0.17624
  |
Correlation:0.1686
  |
Correlation:0.22258
  |
Correlation:-0.019148
  |
Correlation:-0.019148
  |
Correlation:0.16334
 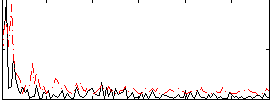 |
Component:150; Jaccard:0.085762
 |
Component:150; Jaccard:0.095044
 |
Component:063; Jaccard:0.1033
 |
Component:150; Jaccard:0.096112
 |
Component:150; Jaccard:0.076188
 |
Component:372; Jaccard:0.053913
 |
Component:372; Jaccard:0.033464
 |
Correlation:0.13003
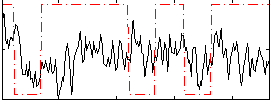  |
Correlation:0.13003
  |
Correlation:0.22258
  |
Correlation:0.13003
  |
Correlation:0.13003
  |
Correlation:-0.043654
  |
Correlation:-0.043654
  |
Component:113; Jaccard:0.084951
 |
Component:027; Jaccard:0.093922
 |
Component:150; Jaccard:0.098729
 |
Component:397; Jaccard:0.095798
 |
Component:372; Jaccard:0.074688
 |
Component:007; Jaccard:0.049593
 |
Component:007; Jaccard:0.0299
 |
Correlation:0.17624
  |
Correlation:0.24559
  |
Correlation:0.13003
  |
Correlation:0.1686
  |
Correlation:-0.043654
  |
Correlation:0.1034
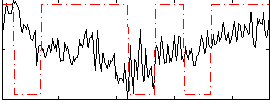  |
Correlation:0.1034
  |
Component:007; Jaccard:0.08387
 |
Component:007; Jaccard:0.091482
 |
Component:027; Jaccard:0.09599
 |
Component:007; Jaccard:0.088933
 |
Component:397; Jaccard:0.074041
 |
Component:110; Jaccard:0.049474
 |
Component:054; Jaccard:0.029489
 |
Correlation:0.1034
  |
Correlation:0.1034
  |
Correlation:0.24559
  |
Correlation:0.1034
  |
Correlation:0.1686
  |
Correlation:0.056484
 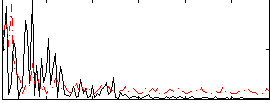 |
Correlation:-0.019148
  |
Component:261; Jaccard:0.082908
 |
Component:063; Jaccard:0.090814
 |
Component:007; Jaccard:0.094978
 |
Component:110; Jaccard:0.087605
 |
Component:110; Jaccard:0.072108
 |
Component:383; Jaccard:0.048918
 |
Component:110; Jaccard:0.028878
 |
Correlation:0.15685
  |
Correlation:0.22258
  |
Correlation:0.1034
  |
Correlation:0.056484
  |
Correlation:0.056484
  |
Correlation:0.16744
  |
Correlation:0.056484
  |
Component:121; Jaccard:0.081769
 |
Component:110; Jaccard:0.086974
 |
Component:110; Jaccard:0.092042
 |
Component:372; Jaccard:0.086788
 |
Component:007; Jaccard:0.071573
 |
Component:344; Jaccard:0.048128
 |
Component:354; Jaccard:0.027113
 |
Correlation:0.067447
  |
Correlation:0.056484
  |
Correlation:0.056484
  |
Correlation:-0.043654
  |
Correlation:0.1034
  |
Correlation:0.16334
  |
Correlation:-0.00088659
  |
Component:398; Jaccard:0.08126
 |
Component:398; Jaccard:0.086421
 |
Component:084; Jaccard:0.089005
 |
Component:084; Jaccard:0.085665
 |
Component:383; Jaccard:0.070968
 |
Component:397; Jaccard:0.047884
 |
Component:305; Jaccard:0.026944
 |
Correlation:-0.10041
  |
Correlation:-0.10041
  |
Correlation:0.070969
  |
Correlation:0.070969
  |
Correlation:0.16744
  |
Correlation:0.1686
  |
Correlation:0.13023
  |
Cope: 06:neg_REL BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:027; Jaccard:0.098904
 |
Component:027; Jaccard:0.11266
 |
Component:027; Jaccard:0.12143
 |
Component:027; Jaccard:0.11794
 |
Component:027; Jaccard:0.095993
 |
Component:105; Jaccard:0.068299
 |
Component:105; Jaccard:0.048825
 |
Correlation:0.17305
  |
Correlation:0.17305
  |
Correlation:0.17305
  |
Correlation:0.17305
  |
Correlation:0.17305
  |
Correlation:0.25908
  |
Correlation:0.25908
  |
Component:313; Jaccard:0.096326
 |
Component:313; Jaccard:0.10892
 |
Component:313; Jaccard:0.11579
 |
Component:313; Jaccard:0.11235
 |
Component:313; Jaccard:0.094237
 |
Component:027; Jaccard:0.066419
 |
Component:027; Jaccard:0.040202
 |
Correlation:-0.064755
  |
Correlation:-0.064755
  |
Correlation:-0.064755
  |
Correlation:-0.064755
  |
Correlation:-0.064755
  |
Correlation:0.17305
  |
Correlation:0.17305
  |
Component:110; Jaccard:0.090502
 |
Component:110; Jaccard:0.10221
 |
Component:110; Jaccard:0.10964
 |
Component:110; Jaccard:0.10803
 |
Component:110; Jaccard:0.092257
 |
Component:110; Jaccard:0.065123
 |
Component:355; Jaccard:0.039911
 |
Correlation:0.16193
  |
Correlation:0.16193
  |
Correlation:0.16193
  |
Correlation:0.16193
  |
Correlation:0.16193
  |
Correlation:0.16193
  |
Correlation:0.12499
  |
Component:246; Jaccard:0.087596
 |
Component:150; Jaccard:0.097861
 |
Component:150; Jaccard:0.1052
 |
Component:150; Jaccard:0.098691
 |
Component:150; Jaccard:0.082516
 |
Component:313; Jaccard:0.062756
 |
Component:110; Jaccard:0.039254
 |
Correlation:0.23695
  |
Correlation:0.083559
  |
Correlation:0.083559
  |
Correlation:0.083559
  |
Correlation:0.083559
  |
Correlation:-0.064755
  |
Correlation:0.16193
  |
Component:150; Jaccard:0.086252
 |
Component:246; Jaccard:0.096433
 |
Component:246; Jaccard:0.10028
 |
Component:246; Jaccard:0.095487
 |
Component:084; Jaccard:0.078631
 |
Component:355; Jaccard:0.055417
 |
Component:222; Jaccard:0.034203
 |
Correlation:0.083559
  |
Correlation:0.23695
  |
Correlation:0.23695
  |
Correlation:0.23695
  |
Correlation:0.0040992
  |
Correlation:0.12499
  |
Correlation:0.066581
  |
Component:319; Jaccard:0.084168
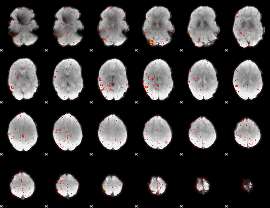 |
Component:398; Jaccard:0.090028
 |
Component:113; Jaccard:0.094972
 |
Component:113; Jaccard:0.091456
 |
Component:246; Jaccard:0.078546
 |
Component:222; Jaccard:0.055244
 |
Component:113; Jaccard:0.031888
 |
Correlation:0.21397
  |
Correlation:-0.03568
  |
Correlation:0.21629
  |
Correlation:0.21629
  |
Correlation:0.23695
  |
Correlation:0.066581
  |
Correlation:0.21629
  |
Component:108; Jaccard:0.082912
 |
Component:319; Jaccard:0.089541
 |
Component:354; Jaccard:0.09309
 |
Component:084; Jaccard:0.090479
 |
Component:113; Jaccard:0.078217
 |
Component:150; Jaccard:0.054696
 |
Component:313; Jaccard:0.031852
 |
Correlation:0.069302
  |
Correlation:0.21397
  |
Correlation:0.052176
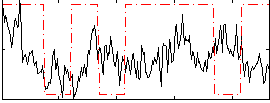  |
Correlation:0.0040992
  |
Correlation:0.21629
  |
Correlation:0.083559
  |
Correlation:-0.064755
  |
Component:398; Jaccard:0.081982
 |
Component:354; Jaccard:0.087292
 |
Component:398; Jaccard:0.091339
 |
Component:354; Jaccard:0.08992
 |
Component:354; Jaccard:0.07783
 |
Component:084; Jaccard:0.054085
 |
Component:007; Jaccard:0.031733
 |
Correlation:-0.03568
  |
Correlation:0.052176
  |
Correlation:-0.03568
  |
Correlation:0.052176
  |
Correlation:0.052176
  |
Correlation:0.0040992
  |
Correlation:0.16098
  |
Component:397; Jaccard:0.081029
 |
Component:121; Jaccard:0.085896
 |
Component:084; Jaccard:0.090006
 |
Component:007; Jaccard:0.086875
 |
Component:105; Jaccard:0.077422
 |
Component:246; Jaccard:0.053397
 |
Component:372; Jaccard:0.031127
 |
Correlation:0.058477
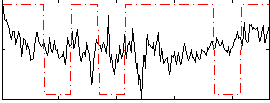  |
Correlation:0.14755
 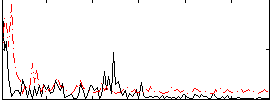 |
Correlation:0.0040992
  |
Correlation:0.16098
  |
Correlation:0.25908
  |
Correlation:0.23695
  |
Correlation:0.32728
  |
Component:121; Jaccard:0.080761
 |
Component:113; Jaccard:0.085541
 |
Component:007; Jaccard:0.089205
 |
Component:398; Jaccard:0.08633
 |
Component:222; Jaccard:0.075112
 |
Component:113; Jaccard:0.053054
 |
Component:017; Jaccard:0.03101
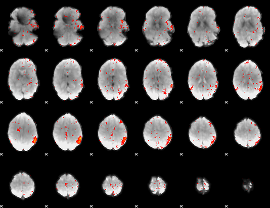 |
Correlation:0.14755
  |
Correlation:0.21629
  |
Correlation:0.16098
  |
Correlation:-0.03568
  |
Correlation:0.066581
  |
Correlation:0.21629
  |
Correlation:0.10323
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































