Task name: RELATIONAL, TR=0.72, totally 232 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"MATCH" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
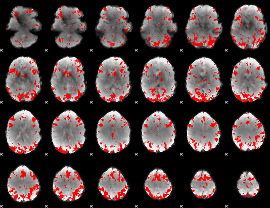 |
GLM binary result thresholded by 3.5
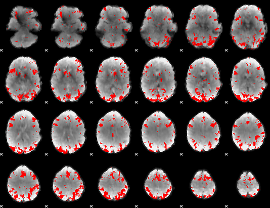 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
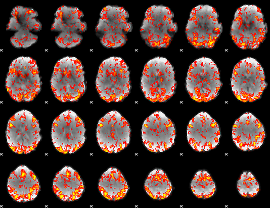 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
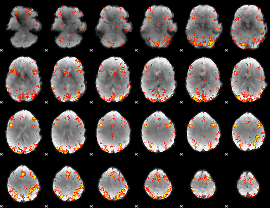 |
GLM Z-score result thresholded by 4
 |
Component:274; Jaccard:0.11243
 |
Component:274; Jaccard:0.13426
 |
Component:274; Jaccard:0.15709
 |
Component:316; Jaccard:0.18003
 |
Component:316; Jaccard:0.21263
 |
Component:316; Jaccard:0.2399
 |
Component:316; Jaccard:0.25469
 |
Correlation:-0.0047703
  |
Correlation:-0.0047703
  |
Correlation:-0.0047703
  |
Correlation:0.15914
  |
Correlation:0.15914
  |
Correlation:0.15914
  |
Correlation:0.15914
  |
Component:297; Jaccard:0.1059
 |
Component:251; Jaccard:0.12356
 |
Component:316; Jaccard:0.14819
 |
Component:274; Jaccard:0.1759
 |
Component:274; Jaccard:0.18663
 |
Component:379; Jaccard:0.21265
 |
Component:379; Jaccard:0.22964
 |
Correlation:-0.069335
  |
Correlation:0.18646
  |
Correlation:0.15914
  |
Correlation:-0.0047703
  |
Correlation:-0.0047703
  |
Correlation:0.21327
  |
Correlation:0.21327
  |
Component:370; Jaccard:0.10396
 |
Component:297; Jaccard:0.12193
 |
Component:251; Jaccard:0.14519
 |
Component:251; Jaccard:0.16556
 |
Component:379; Jaccard:0.18509
 |
Component:209; Jaccard:0.18942
 |
Component:209; Jaccard:0.19207
 |
Correlation:-0.078349
  |
Correlation:-0.069335
  |
Correlation:0.18646
  |
Correlation:0.18646
  |
Correlation:0.21327
  |
Correlation:0.1444
  |
Correlation:0.1444
  |
Component:251; Jaccard:0.10324
 |
Component:370; Jaccard:0.12079
 |
Component:297; Jaccard:0.1384
 |
Component:209; Jaccard:0.15969
 |
Component:251; Jaccard:0.17888
 |
Component:274; Jaccard:0.18749
 |
Component:191; Jaccard:0.1874
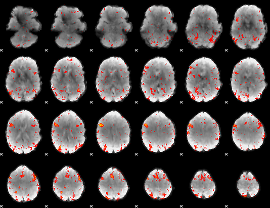 |
Correlation:0.18646
  |
Correlation:-0.078349
  |
Correlation:-0.069335
  |
Correlation:0.1444
  |
Correlation:0.18646
  |
Correlation:-0.0047703
  |
Correlation:0.12842
  |
Component:327; Jaccard:0.10263
 |
Component:316; Jaccard:0.11899
 |
Component:370; Jaccard:0.13802
 |
Component:191; Jaccard:0.15686
 |
Component:209; Jaccard:0.17743
 |
Component:191; Jaccard:0.18689
 |
Component:274; Jaccard:0.17748
 |
Correlation:-0.11771
  |
Correlation:0.15914
  |
Correlation:-0.078349
  |
Correlation:0.12842
  |
Correlation:0.1444
  |
Correlation:0.12842
  |
Correlation:-0.0047703
  |
Component:232; Jaccard:0.098357
 |
Component:327; Jaccard:0.11792
 |
Component:209; Jaccard:0.13757
 |
Component:379; Jaccard:0.1551
 |
Component:191; Jaccard:0.17622
 |
Component:251; Jaccard:0.18145
 |
Component:251; Jaccard:0.17179
 |
Correlation:-0.090768
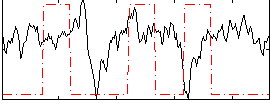 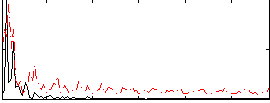 |
Correlation:-0.11771
  |
Correlation:0.1444
  |
Correlation:0.21327
  |
Correlation:0.12842
  |
Correlation:0.18646
  |
Correlation:0.18646
  |
Component:071; Jaccard:0.097874
 |
Component:209; Jaccard:0.1159
 |
Component:191; Jaccard:0.13565
 |
Component:370; Jaccard:0.15181
 |
Component:194; Jaccard:0.16502
 |
Component:194; Jaccard:0.17477
 |
Component:194; Jaccard:0.17156
 |
Correlation:-0.041774
 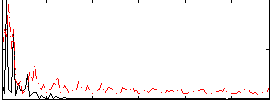 |
Correlation:0.1444
  |
Correlation:0.12842
  |
Correlation:-0.078349
  |
Correlation:0.17804
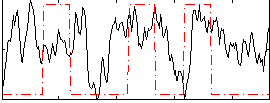  |
Correlation:0.17804
  |
Correlation:0.17804
  |
Component:209; Jaccard:0.097118
 |
Component:232; Jaccard:0.11432
 |
Component:327; Jaccard:0.13455
 |
Component:297; Jaccard:0.15105
 |
Component:297; Jaccard:0.15957
 |
Component:297; Jaccard:0.15927
 |
Component:231; Jaccard:0.15828
 |
Correlation:0.1444
  |
Correlation:-0.090768
  |
Correlation:-0.11771
  |
Correlation:-0.069335
  |
Correlation:-0.069335
  |
Correlation:-0.069335
  |
Correlation:0.078656
  |
Component:349; Jaccard:0.096299
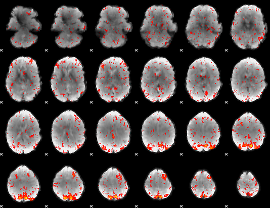 |
Component:071; Jaccard:0.11398
 |
Component:071; Jaccard:0.13104
 |
Component:327; Jaccard:0.14819
 |
Component:327; Jaccard:0.15624
 |
Component:327; Jaccard:0.15514
 |
Component:222; Jaccard:0.15153
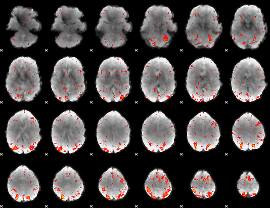 |
Correlation:-0.062452
  |
Correlation:-0.041774
  |
Correlation:-0.041774
  |
Correlation:-0.11771
  |
Correlation:-0.11771
  |
Correlation:-0.11771
  |
Correlation:-0.071057
  |
Component:316; Jaccard:0.095922
 |
Component:191; Jaccard:0.11378
 |
Component:232; Jaccard:0.13061
 |
Component:194; Jaccard:0.14735
 |
Component:370; Jaccard:0.15552
 |
Component:232; Jaccard:0.15493
 |
Component:297; Jaccard:0.15045
 |
Correlation:0.15914
  |
Correlation:0.12842
  |
Correlation:-0.090768
  |
Correlation:0.17804
  |
Correlation:-0.078349
  |
Correlation:-0.090768
  |
Correlation:-0.069335
  |
Cope: 02:"REL" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
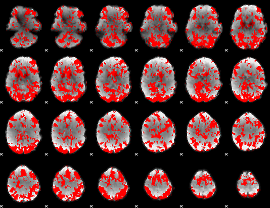 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
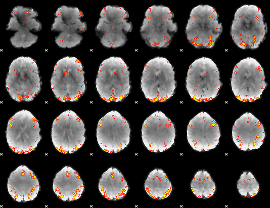 |
Component:071; Jaccard:0.12787
 |
Component:071; Jaccard:0.14742
 |
Component:379; Jaccard:0.17167
 |
Component:379; Jaccard:0.20547
 |
Component:379; Jaccard:0.23876
 |
Component:379; Jaccard:0.26493
 |
Component:379; Jaccard:0.27432
 |
Correlation:-0.072035
  |
Correlation:-0.072035
  |
Correlation:0.17607
  |
Correlation:0.17607
  |
Correlation:0.17607
  |
Correlation:0.17607
  |
Correlation:0.17607
  |
Component:370; Jaccard:0.12078
 |
Component:316; Jaccard:0.14185
 |
Component:071; Jaccard:0.16609
 |
Component:316; Jaccard:0.18625
 |
Component:316; Jaccard:0.20487
 |
Component:231; Jaccard:0.21563
 |
Component:231; Jaccard:0.22555
 |
Correlation:0.21347
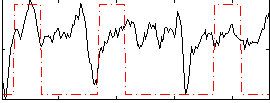  |
Correlation:0.15518
  |
Correlation:-0.072035
  |
Correlation:0.15518
  |
Correlation:0.15518
  |
Correlation:0.2725
  |
Correlation:0.2725
  |
Component:232; Jaccard:0.12016
 |
Component:379; Jaccard:0.14021
 |
Component:316; Jaccard:0.16576
 |
Component:071; Jaccard:0.18074
 |
Component:231; Jaccard:0.19752
 |
Component:316; Jaccard:0.21351
 |
Component:316; Jaccard:0.21091
 |
Correlation:0.035166
  |
Correlation:0.17607
  |
Correlation:0.15518
  |
Correlation:-0.072035
  |
Correlation:0.2725
  |
Correlation:0.15518
  |
Correlation:0.15518
  |
Component:316; Jaccard:0.12007
 |
Component:222; Jaccard:0.13578
 |
Component:222; Jaccard:0.15394
 |
Component:231; Jaccard:0.17303
 |
Component:071; Jaccard:0.18775
 |
Component:071; Jaccard:0.18477
 |
Component:222; Jaccard:0.17777
 |
Correlation:0.15518
  |
Correlation:0.2028
  |
Correlation:0.2028
  |
Correlation:0.2725
  |
Correlation:-0.072035
  |
Correlation:-0.072035
  |
Correlation:0.2028
  |
Component:274; Jaccard:0.11988
 |
Component:232; Jaccard:0.13509
 |
Component:232; Jaccard:0.14954
 |
Component:222; Jaccard:0.17002
 |
Component:222; Jaccard:0.18202
 |
Component:222; Jaccard:0.18272
 |
Component:209; Jaccard:0.17121
 |
Correlation:0.0091015
  |
Correlation:0.035166
  |
Correlation:0.035166
  |
Correlation:0.2028
  |
Correlation:0.2028
  |
Correlation:0.2028
  |
Correlation:0.048935
  |
Component:222; Jaccard:0.11654
 |
Component:274; Jaccard:0.13337
 |
Component:231; Jaccard:0.14752
 |
Component:209; Jaccard:0.15833
 |
Component:209; Jaccard:0.17029
 |
Component:209; Jaccard:0.17378
 |
Component:071; Jaccard:0.16578
 |
Correlation:0.2028
  |
Correlation:0.0091015
  |
Correlation:0.2725
  |
Correlation:0.048935
  |
Correlation:0.048935
  |
Correlation:0.048935
  |
Correlation:-0.072035
  |
Component:379; Jaccard:0.11256
 |
Component:370; Jaccard:0.13335
 |
Component:274; Jaccard:0.14498
 |
Component:232; Jaccard:0.15761
 |
Component:232; Jaccard:0.15799
 |
Component:362; Jaccard:0.15622
 |
Component:362; Jaccard:0.15504
 |
Correlation:0.17607
  |
Correlation:0.21347
  |
Correlation:0.0091015
  |
Correlation:0.035166
  |
Correlation:0.035166
  |
Correlation:0.46361
  |
Correlation:0.46361
  |
Component:297; Jaccard:0.11255
 |
Component:209; Jaccard:0.12653
 |
Component:370; Jaccard:0.14492
 |
Component:274; Jaccard:0.15344
 |
Component:274; Jaccard:0.15574
 |
Component:232; Jaccard:0.14857
 |
Component:232; Jaccard:0.13554
 |
Correlation:0.14822
  |
Correlation:0.048935
  |
Correlation:0.21347
  |
Correlation:0.0091015
  |
Correlation:0.0091015
  |
Correlation:0.035166
  |
Correlation:0.035166
  |
Component:209; Jaccard:0.11093
 |
Component:231; Jaccard:0.12401
 |
Component:209; Jaccard:0.14376
 |
Component:370; Jaccard:0.15127
 |
Component:370; Jaccard:0.15214
 |
Component:274; Jaccard:0.14805
 |
Component:274; Jaccard:0.13199
 |
Correlation:0.048935
  |
Correlation:0.2725
  |
Correlation:0.048935
  |
Correlation:0.21347
  |
Correlation:0.21347
  |
Correlation:0.0091015
  |
Correlation:0.0091015
  |
Component:327; Jaccard:0.10487
 |
Component:297; Jaccard:0.12301
 |
Component:297; Jaccard:0.13187
 |
Component:297; Jaccard:0.13868
 |
Component:362; Jaccard:0.14941
 |
Component:370; Jaccard:0.14381
 |
Component:367; Jaccard:0.13024
 |
Correlation:0.22289
  |
Correlation:0.14822
  |
Correlation:0.14822
  |
Correlation:0.14822
  |
Correlation:0.46361
  |
Correlation:0.21347
  |
Correlation:-0.047276
  |
Cope: 03:"MATCH-REL" BACK
|
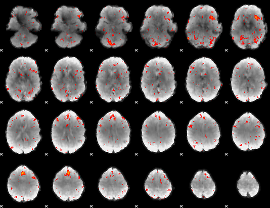
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
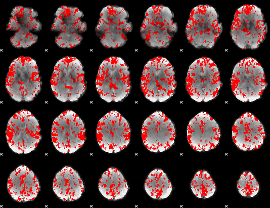 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:377; Jaccard:0.11241
 |
Component:377; Jaccard:0.14087
 |
Component:377; Jaccard:0.1735
 |
Component:377; Jaccard:0.2065
 |
Component:377; Jaccard:0.22415
 |
Component:377; Jaccard:0.20871
 |
Component:377; Jaccard:0.16366
 |
Correlation:0.0069774
 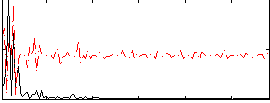 |
Correlation:0.0069774
  |
Correlation:0.0069774
  |
Correlation:0.0069774
  |
Correlation:0.0069774
  |
Correlation:0.0069774
  |
Correlation:0.0069774
  |
Component:047; Jaccard:0.1008
 |
Component:047; Jaccard:0.11955
 |
Component:047; Jaccard:0.13988
 |
Component:047; Jaccard:0.15588
 |
Component:375; Jaccard:0.16963
 |
Component:375; Jaccard:0.16947
 |
Component:375; Jaccard:0.14239
 |
Correlation:-0.068776
  |
Correlation:-0.068776
  |
Correlation:-0.068776
  |
Correlation:-0.068776
  |
Correlation:0.14601
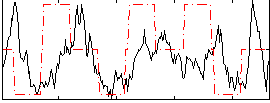  |
Correlation:0.14601
  |
Correlation:0.14601
  |
Component:304; Jaccard:0.091459
 |
Component:375; Jaccard:0.10641
 |
Component:375; Jaccard:0.12987
 |
Component:375; Jaccard:0.15324
 |
Component:047; Jaccard:0.15828
 |
Component:047; Jaccard:0.14409
 |
Component:047; Jaccard:0.11146
 |
Correlation:-0.09061
  |
Correlation:0.14601
  |
Correlation:0.14601
  |
Correlation:0.14601
  |
Correlation:-0.068776
  |
Correlation:-0.068776
  |
Correlation:-0.068776
  |
Component:354; Jaccard:0.089806
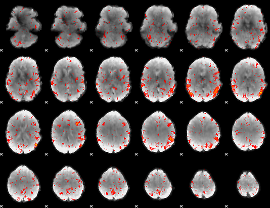 |
Component:354; Jaccard:0.10617
 |
Component:354; Jaccard:0.12099
 |
Component:354; Jaccard:0.13093
 |
Component:354; Jaccard:0.12878
 |
Component:354; Jaccard:0.1094
 |
Component:354; Jaccard:0.084396
 |
Correlation:0.16943
  |
Correlation:0.16943
  |
Correlation:0.16943
  |
Correlation:0.16943
  |
Correlation:0.16943
  |
Correlation:0.16943
  |
Correlation:0.16943
  |
Component:228; Jaccard:0.087029
 |
Component:304; Jaccard:0.099542
 |
Component:228; Jaccard:0.10613
 |
Component:228; Jaccard:0.10962
 |
Component:228; Jaccard:0.10588
 |
Component:050; Jaccard:0.090928
 |
Component:050; Jaccard:0.074198
 |
Correlation:-0.082165
  |
Correlation:-0.09061
  |
Correlation:-0.082165
  |
Correlation:-0.082165
  |
Correlation:-0.082165
  |
Correlation:0.16111
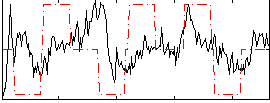  |
Correlation:0.16111
  |
Component:349; Jaccard:0.086716
 |
Component:228; Jaccard:0.097924
 |
Component:304; Jaccard:0.10318
 |
Component:283; Jaccard:0.10701
 |
Component:283; Jaccard:0.10441
 |
Component:283; Jaccard:0.08943
 |
Component:308; Jaccard:0.07135
 |
Correlation:-0.10517
  |
Correlation:-0.082165
  |
Correlation:-0.09061
  |
Correlation:-0.1141
  |
Correlation:-0.1141
  |
Correlation:-0.1141
  |
Correlation:0.13609
  |
Component:186; Jaccard:0.086073
 |
Component:186; Jaccard:0.094829
 |
Component:283; Jaccard:0.10141
 |
Component:088; Jaccard:0.10199
 |
Component:088; Jaccard:0.098048
 |
Component:228; Jaccard:0.086541
 |
Component:283; Jaccard:0.067833
 |
Correlation:-0.15396
  |
Correlation:-0.15396
  |
Correlation:-0.1141
  |
Correlation:0.1803
  |
Correlation:0.1803
  |
Correlation:-0.082165
  |
Correlation:-0.1141
  |
Component:375; Jaccard:0.085385
 |
Component:283; Jaccard:0.09197
 |
Component:186; Jaccard:0.10102
 |
Component:304; Jaccard:0.10094
 |
Component:050; Jaccard:0.096909
 |
Component:369; Jaccard:0.082918
 |
Component:369; Jaccard:0.066047
 |
Correlation:0.14601
  |
Correlation:-0.1141
  |
Correlation:-0.15396
  |
Correlation:-0.09061
  |
Correlation:0.16111
  |
Correlation:-0.051045
  |
Correlation:-0.051045
  |
Component:318; Jaccard:0.08507
 |
Component:318; Jaccard:0.089853
 |
Component:088; Jaccard:0.097838
 |
Component:186; Jaccard:0.10012
 |
Component:255; Jaccard:0.093557
 |
Component:088; Jaccard:0.082435
 |
Component:228; Jaccard:0.063664
 |
Correlation:-0.087525
  |
Correlation:-0.087525
  |
Correlation:0.1803
  |
Correlation:-0.15396
  |
Correlation:0.052455
  |
Correlation:0.1803
  |
Correlation:-0.082165
  |
Component:230; Jaccard:0.081828
 |
Component:349; Jaccard:0.088878
 |
Component:230; Jaccard:0.094336
 |
Component:255; Jaccard:0.097376
 |
Component:346; Jaccard:0.093061
 |
Component:308; Jaccard:0.08164
 |
Component:346; Jaccard:0.061579
 |
Correlation:0.006389
  |
Correlation:-0.10517
  |
Correlation:0.006389
  |
Correlation:0.052455
  |
Correlation:0.043373
  |
Correlation:0.13609
  |
Correlation:0.043373
  |
Cope: 04:"REL-MATCH" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
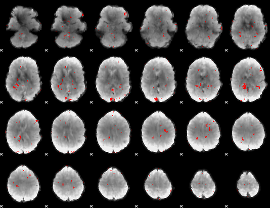 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
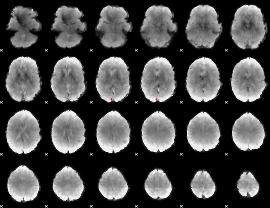 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
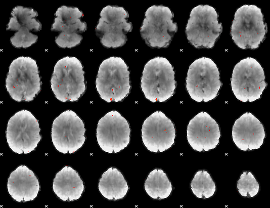 |
GLM Z-score result thresholded by 4
 |
Component:362; Jaccard:0.088451
 |
Component:123; Jaccard:0.090959
 |
Component:123; Jaccard:0.09617
 |
Component:123; Jaccard:0.071874
 |
Component:123; Jaccard:0.030335
 |
Component:362; Jaccard:0.011565
 |
Component:362; Jaccard:0.005961
 |
Correlation:0.29857
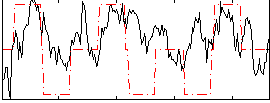  |
Correlation:-0.00025461
  |
Correlation:-0.00025461
  |
Correlation:-0.00025461
  |
Correlation:-0.00025461
  |
Correlation:0.29857
  |
Correlation:0.29857
  |
Component:256; Jaccard:0.076952
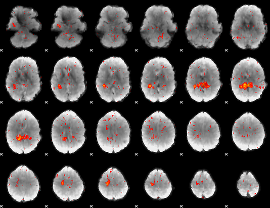 |
Component:256; Jaccard:0.083074
 |
Component:256; Jaccard:0.078675
 |
Component:256; Jaccard:0.0512
 |
Component:362; Jaccard:0.023167
 |
Component:234; Jaccard:0.009928
 |
Component:286; Jaccard:0.004399
 |
Correlation:0.072381
  |
Correlation:0.072381
  |
Correlation:0.072381
  |
Correlation:0.072381
  |
Correlation:0.29857
  |
Correlation:0.33085
  |
Correlation:0.25957
  |
Component:123; Jaccard:0.076405
 |
Component:362; Jaccard:0.082353
 |
Component:118; Jaccard:0.063483
 |
Component:118; Jaccard:0.042965
 |
Component:256; Jaccard:0.022731
 |
Component:124; Jaccard:0.009206
 |
Component:231; Jaccard:0.004358
 |
Correlation:-0.00025461
  |
Correlation:0.29857
  |
Correlation:-0.048488
  |
Correlation:-0.048488
  |
Correlation:0.072381
  |
Correlation:0.19706
  |
Correlation:0.11489
  |
Component:352; Jaccard:0.073642
 |
Component:352; Jaccard:0.073112
 |
Component:384; Jaccard:0.062596
 |
Component:234; Jaccard:0.040913
 |
Component:234; Jaccard:0.02257
 |
Component:123; Jaccard:0.008741
 |
Component:124; Jaccard:0.003934
 |
Correlation:0.10015
  |
Correlation:0.10015
  |
Correlation:0.092617
  |
Correlation:0.33085
  |
Correlation:0.33085
  |
Correlation:-0.00025461
  |
Correlation:0.19706
  |
Component:384; Jaccard:0.070732
 |
Component:118; Jaccard:0.072727
 |
Component:352; Jaccard:0.062087
 |
Component:362; Jaccard:0.040614
 |
Component:027; Jaccard:0.021063
 |
Component:027; Jaccard:0.008113
 |
Component:379; Jaccard:0.003303
 |
Correlation:0.092617
  |
Correlation:-0.048488
  |
Correlation:0.10015
  |
Correlation:0.29857
  |
Correlation:0.031941
  |
Correlation:0.031941
  |
Correlation:-0.022045
  |
Component:118; Jaccard:0.069242
 |
Component:384; Jaccard:0.072213
 |
Component:362; Jaccard:0.061585
 |
Component:384; Jaccard:0.039983
 |
Component:118; Jaccard:0.020688
 |
Component:118; Jaccard:0.008014
 |
Component:234; Jaccard:0.003222
 |
Correlation:-0.048488
  |
Correlation:0.092617
  |
Correlation:0.29857
  |
Correlation:0.092617
  |
Correlation:-0.048488
  |
Correlation:-0.048488
  |
Correlation:0.33085
  |
Component:231; Jaccard:0.066819
 |
Component:234; Jaccard:0.065078
 |
Component:234; Jaccard:0.058597
 |
Component:027; Jaccard:0.037441
 |
Component:124; Jaccard:0.017893
 |
Component:076; Jaccard:0.007902
 |
Component:343; Jaccard:0.002806
 |
Correlation:0.11489
  |
Correlation:0.33085
  |
Correlation:0.33085
  |
Correlation:0.031941
  |
Correlation:0.19706
  |
Correlation:0.087512
  |
Correlation:0.019462
  |
Component:156; Jaccard:0.062301
 |
Component:156; Jaccard:0.062892
 |
Component:156; Jaccard:0.053493
 |
Component:352; Jaccard:0.037229
 |
Component:384; Jaccard:0.017736
 |
Component:231; Jaccard:0.007901
 |
Component:075; Jaccard:0.002531
 |
Correlation:0.044442
  |
Correlation:0.044442
  |
Correlation:0.044442
  |
Correlation:0.10015
  |
Correlation:0.092617
  |
Correlation:0.11489
  |
Correlation:0.051582
  |
Component:234; Jaccard:0.06173
 |
Component:231; Jaccard:0.058471
 |
Component:027; Jaccard:0.046222
 |
Component:076; Jaccard:0.034596
 |
Component:076; Jaccard:0.01762
 |
Component:286; Jaccard:0.007817
 |
Component:197; Jaccard:0.002276
 |
Correlation:0.33085
  |
Correlation:0.11489
  |
Correlation:0.031941
  |
Correlation:0.087512
  |
Correlation:0.087512
  |
Correlation:0.25957
  |
Correlation:0.16487
  |
Component:241; Jaccard:0.057556
 |
Component:241; Jaccard:0.055349
 |
Component:241; Jaccard:0.046201
 |
Component:156; Jaccard:0.034207
 |
Component:352; Jaccard:0.016865
 |
Component:293; Jaccard:0.007063
 |
Component:123; Jaccard:0.00224
 |
Correlation:0.090918
  |
Correlation:0.090918
  |
Correlation:0.090918
  |
Correlation:0.044442
  |
Correlation:0.10015
  |
Correlation:0.28877
  |
Correlation:-0.00025461
  |
Cope: 05:"neg_MATCH" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:255; Jaccard:0.094803
 |
Component:082; Jaccard:0.093537
 |
Component:082; Jaccard:0.10566
 |
Component:082; Jaccard:0.11016
 |
Component:082; Jaccard:0.09575
 |
Component:082; Jaccard:0.075135
 |
Component:082; Jaccard:0.042501
 |
Correlation:0.11293
  |
Correlation:0.1002
  |
Correlation:0.1002
  |
Correlation:0.1002
  |
Correlation:0.1002
  |
Correlation:0.1002
  |
Correlation:0.1002
  |
Component:017; Jaccard:0.087278
 |
Component:255; Jaccard:0.093167
 |
Component:006; Jaccard:0.085592
 |
Component:006; Jaccard:0.087991
 |
Component:006; Jaccard:0.075457
 |
Component:006; Jaccard:0.052022
 |
Component:161; Jaccard:0.035642
 |
Correlation:0.25805
  |
Correlation:0.11293
  |
Correlation:0.11446
  |
Correlation:0.11446
  |
Correlation:0.11446
  |
Correlation:0.11446
  |
Correlation:0.10505
  |
Component:272; Jaccard:0.084427
 |
Component:017; Jaccard:0.089784
 |
Component:255; Jaccard:0.083816
 |
Component:356; Jaccard:0.074239
 |
Component:161; Jaccard:0.063806
 |
Component:161; Jaccard:0.051439
 |
Component:237; Jaccard:0.028746
 |
Correlation:0.085998
  |
Correlation:0.25805
  |
Correlation:0.11293
  |
Correlation:0.23559
  |
Correlation:0.10505
  |
Correlation:0.10505
  |
Correlation:0.12371
  |
Component:107; Jaccard:0.08412
 |
Component:272; Jaccard:0.083011
 |
Component:017; Jaccard:0.083327
 |
Component:237; Jaccard:0.073229
 |
Component:237; Jaccard:0.062973
 |
Component:237; Jaccard:0.045077
 |
Component:006; Jaccard:0.02604
 |
Correlation:0.26601
  |
Correlation:0.085998
  |
Correlation:0.25805
  |
Correlation:0.12371
  |
Correlation:0.12371
  |
Correlation:0.12371
  |
Correlation:0.11446
  |
Component:321; Jaccard:0.079785
 |
Component:107; Jaccard:0.080169
 |
Component:356; Jaccard:0.077624
 |
Component:359; Jaccard:0.073125
 |
Component:356; Jaccard:0.059445
 |
Component:356; Jaccard:0.037384
 |
Component:356; Jaccard:0.019147
 |
Correlation:0.095116
  |
Correlation:0.26601
  |
Correlation:0.23559
  |
Correlation:0.12185
  |
Correlation:0.23559
  |
Correlation:0.23559
  |
Correlation:0.23559
  |
Component:082; Jaccard:0.077635
 |
Component:321; Jaccard:0.079043
 |
Component:359; Jaccard:0.076503
 |
Component:161; Jaccard:0.06987
 |
Component:359; Jaccard:0.056452
 |
Component:100; Jaccard:0.033513
 |
Component:100; Jaccard:0.019093
 |
Correlation:0.1002
  |
Correlation:0.095116
  |
Correlation:0.12185
  |
Correlation:0.10505
  |
Correlation:0.12185
  |
Correlation:-0.052405
  |
Correlation:-0.052405
  |
Component:047; Jaccard:0.074865
 |
Component:006; Jaccard:0.07889
 |
Component:237; Jaccard:0.075849
 |
Component:017; Jaccard:0.068233
 |
Component:341; Jaccard:0.049988
 |
Component:359; Jaccard:0.031292
 |
Component:115; Jaccard:0.016677
 |
Correlation:0.17819
  |
Correlation:0.11446
  |
Correlation:0.12371
  |
Correlation:0.25805
  |
Correlation:0.21593
  |
Correlation:0.12185
  |
Correlation:0.053173
  |
Component:356; Jaccard:0.072815
 |
Component:356; Jaccard:0.076845
 |
Component:272; Jaccard:0.075035
 |
Component:341; Jaccard:0.065352
 |
Component:017; Jaccard:0.047649
 |
Component:105; Jaccard:0.028494
 |
Component:293; Jaccard:0.015714
 |
Correlation:0.23559
  |
Correlation:0.23559
  |
Correlation:0.085998
  |
Correlation:0.21593
  |
Correlation:0.25805
  |
Correlation:0.00097453
  |
Correlation:0.29116
  |
Component:266; Jaccard:0.072695
 |
Component:237; Jaccard:0.074846
 |
Component:107; Jaccard:0.072327
 |
Component:255; Jaccard:0.065157
 |
Component:058; Jaccard:0.047078
 |
Component:341; Jaccard:0.028314
 |
Component:344; Jaccard:0.015454
 |
Correlation:0.15711
  |
Correlation:0.12371
  |
Correlation:0.26601
  |
Correlation:0.11293
  |
Correlation:0.0083203
  |
Correlation:0.21593
  |
Correlation:-0.022224
  |
Component:341; Jaccard:0.068857
 |
Component:359; Jaccard:0.072062
 |
Component:321; Jaccard:0.072213
 |
Component:058; Jaccard:0.060087
 |
Component:100; Jaccard:0.044231
 |
Component:058; Jaccard:0.028309
 |
Component:238; Jaccard:0.015377
 |
Correlation:0.21593
  |
Correlation:0.12185
  |
Correlation:0.095116
  |
Correlation:0.0083203
  |
Correlation:-0.052405
  |
Correlation:0.0083203
  |
Correlation:0.14548
  |
Cope: 06:"neg_REL" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:047; Jaccard:0.12688
 |
Component:047; Jaccard:0.14513
 |
Component:047; Jaccard:0.16348
 |
Component:047; Jaccard:0.178
 |
Component:047; Jaccard:0.18145
 |
Component:047; Jaccard:0.16866
 |
Component:047; Jaccard:0.14293
 |
Correlation:0.062157
  |
Correlation:0.062157
  |
Correlation:0.062157
  |
Correlation:0.062157
  |
Correlation:0.062157
  |
Correlation:0.062157
  |
Correlation:0.062157
  |
Component:255; Jaccard:0.10921
 |
Component:255; Jaccard:0.12777
 |
Component:255; Jaccard:0.14795
 |
Component:255; Jaccard:0.16353
 |
Component:255; Jaccard:0.17105
 |
Component:255; Jaccard:0.16618
 |
Component:255; Jaccard:0.1414
 |
Correlation:0.20143
  |
Correlation:0.20143
  |
Correlation:0.20143
  |
Correlation:0.20143
  |
Correlation:0.20143
  |
Correlation:0.20143
  |
Correlation:0.20143
  |
Component:375; Jaccard:0.10037
 |
Component:375; Jaccard:0.11671
 |
Component:375; Jaccard:0.13367
 |
Component:375; Jaccard:0.14926
 |
Component:375; Jaccard:0.15641
 |
Component:375; Jaccard:0.15157
 |
Component:375; Jaccard:0.13265
 |
Correlation:0.32423
  |
Correlation:0.32423
  |
Correlation:0.32423
  |
Correlation:0.32423
  |
Correlation:0.32423
  |
Correlation:0.32423
  |
Correlation:0.32423
  |
Component:228; Jaccard:0.096935
 |
Component:228; Jaccard:0.10496
 |
Component:017; Jaccard:0.11515
 |
Component:017; Jaccard:0.12681
 |
Component:017; Jaccard:0.13526
 |
Component:050; Jaccard:0.1383
 |
Component:050; Jaccard:0.13203
 |
Correlation:0.047618
  |
Correlation:0.047618
  |
Correlation:0.026693
  |
Correlation:0.026693
  |
Correlation:0.026693
  |
Correlation:0.18646
  |
Correlation:0.18646
  |
Component:266; Jaccard:0.095211
 |
Component:266; Jaccard:0.10478
 |
Component:272; Jaccard:0.11477
 |
Component:272; Jaccard:0.12627
 |
Component:272; Jaccard:0.13327
 |
Component:017; Jaccard:0.13532
 |
Component:017; Jaccard:0.12786
 |
Correlation:0.19943
  |
Correlation:0.19943
  |
Correlation:0.074988
  |
Correlation:0.074988
  |
Correlation:0.074988
  |
Correlation:0.026693
  |
Correlation:0.026693
  |
Component:377; Jaccard:0.090222
 |
Component:272; Jaccard:0.10248
 |
Component:266; Jaccard:0.11255
 |
Component:050; Jaccard:0.11836
 |
Component:050; Jaccard:0.13143
 |
Component:272; Jaccard:0.13414
 |
Component:272; Jaccard:0.12075
 |
Correlation:0.10487
  |
Correlation:0.074988
  |
Correlation:0.19943
  |
Correlation:0.18646
  |
Correlation:0.18646
  |
Correlation:0.074988
  |
Correlation:0.074988
  |
Component:272; Jaccard:0.090024
 |
Component:017; Jaccard:0.10084
 |
Component:228; Jaccard:0.11212
 |
Component:266; Jaccard:0.11831
 |
Component:266; Jaccard:0.12118
 |
Component:266; Jaccard:0.11122
 |
Component:266; Jaccard:0.096982
 |
Correlation:0.074988
  |
Correlation:0.026693
  |
Correlation:0.047618
  |
Correlation:0.19943
  |
Correlation:0.19943
  |
Correlation:0.19943
  |
Correlation:0.19943
  |
Component:107; Jaccard:0.08826
 |
Component:377; Jaccard:0.097212
 |
Component:050; Jaccard:0.10578
 |
Component:228; Jaccard:0.117
 |
Component:228; Jaccard:0.11489
 |
Component:228; Jaccard:0.10362
 |
Component:228; Jaccard:0.082787
 |
Correlation:0.059471
  |
Correlation:0.10487
  |
Correlation:0.18646
  |
Correlation:0.047618
  |
Correlation:0.047618
  |
Correlation:0.047618
  |
Correlation:0.047618
  |
Component:176; Jaccard:0.087853
 |
Component:107; Jaccard:0.096889
 |
Component:107; Jaccard:0.10456
 |
Component:107; Jaccard:0.10779
 |
Component:107; Jaccard:0.1041
 |
Component:107; Jaccard:0.095993
 |
Component:346; Jaccard:0.080379
 |
Correlation:0.12362
  |
Correlation:0.059471
  |
Correlation:0.059471
  |
Correlation:0.059471
  |
Correlation:0.059471
  |
Correlation:0.059471
  |
Correlation:0.17218
  |
Component:017; Jaccard:0.087002
 |
Component:176; Jaccard:0.095264
 |
Component:377; Jaccard:0.10265
 |
Component:377; Jaccard:0.10469
 |
Component:369; Jaccard:0.10088
 |
Component:346; Jaccard:0.090681
 |
Component:107; Jaccard:0.080046
 |
Correlation:0.026693
  |
Correlation:0.12362
  |
Correlation:0.10487
  |
Correlation:0.10487
  |
Correlation:0.02149
  |
Correlation:0.17218
  |
Correlation:0.059471
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































