Task name: GAMBLING, TR=0.72, totally 253 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"PUNISH" BACK
|
GLM result group-wise
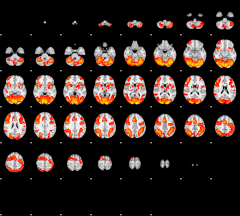 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
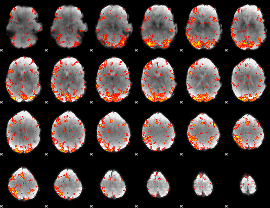 |
GLM Z-score result thresholded by 3
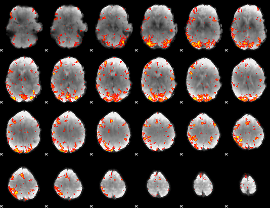 |
GLM Z-score result thresholded by 3.5
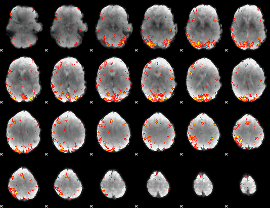 |
GLM Z-score result thresholded by 4
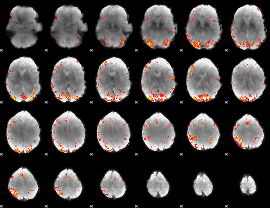 |
Component:291; Jaccard:0.14115
 |
Component:291; Jaccard:0.17519
 |
Component:291; Jaccard:0.21227
 |
Component:291; Jaccard:0.24757
 |
Component:291; Jaccard:0.27059
 |
Component:291; Jaccard:0.26888
 |
Component:291; Jaccard:0.24374
 |
Correlation:0.26615
  |
Correlation:0.26615
  |
Correlation:0.26615
  |
Correlation:0.26615
  |
Correlation:0.26615
  |
Correlation:0.26615
  |
Correlation:0.26615
  |
Component:193; Jaccard:0.13086
 |
Component:137; Jaccard:0.1563
 |
Component:204; Jaccard:0.18613
 |
Component:204; Jaccard:0.21224
 |
Component:137; Jaccard:0.22332
 |
Component:137; Jaccard:0.22106
 |
Component:137; Jaccard:0.19927
 |
Correlation:0.22178
  |
Correlation:0.30495
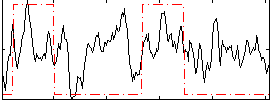 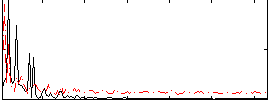 |
Correlation:0.092425
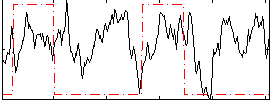 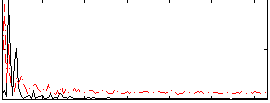 |
Correlation:0.092425
  |
Correlation:0.30495
  |
Correlation:0.30495
  |
Correlation:0.30495
  |
Component:137; Jaccard:0.12893
 |
Component:204; Jaccard:0.15609
 |
Component:137; Jaccard:0.18508
 |
Component:137; Jaccard:0.20971
 |
Component:204; Jaccard:0.22135
 |
Component:204; Jaccard:0.21562
 |
Component:399; Jaccard:0.19256
 |
Correlation:0.30495
  |
Correlation:0.092425
  |
Correlation:0.30495
  |
Correlation:0.30495
  |
Correlation:0.092425
  |
Correlation:0.092425
  |
Correlation:0.18911
  |
Component:204; Jaccard:0.12603
 |
Component:193; Jaccard:0.15226
 |
Component:193; Jaccard:0.17036
 |
Component:193; Jaccard:0.17978
 |
Component:399; Jaccard:0.18996
 |
Component:399; Jaccard:0.19815
 |
Component:204; Jaccard:0.18939
 |
Correlation:0.092425
  |
Correlation:0.22178
  |
Correlation:0.22178
  |
Correlation:0.22178
  |
Correlation:0.18911
  |
Correlation:0.18911
  |
Correlation:0.092425
  |
Component:031; Jaccard:0.12177
 |
Component:031; Jaccard:0.13637
 |
Component:031; Jaccard:0.14792
 |
Component:399; Jaccard:0.17058
 |
Component:193; Jaccard:0.1747
 |
Component:289; Jaccard:0.1735
 |
Component:289; Jaccard:0.16083
 |
Correlation:0.15646
  |
Correlation:0.15646
  |
Correlation:0.15646
  |
Correlation:0.18911
  |
Correlation:0.22178
  |
Correlation:0.25656
  |
Correlation:0.25656
  |
Component:293; Jaccard:0.11327
 |
Component:348; Jaccard:0.12125
 |
Component:399; Jaccard:0.14503
 |
Component:289; Jaccard:0.16221
 |
Component:289; Jaccard:0.1734
 |
Component:364; Jaccard:0.16029
 |
Component:364; Jaccard:0.15088
 |
Correlation:0.1648
  |
Correlation:0.055663
  |
Correlation:0.18911
  |
Correlation:0.25656
  |
Correlation:0.25656
  |
Correlation:0.43402
  |
Correlation:0.43402
  |
Component:348; Jaccard:0.11128
 |
Component:289; Jaccard:0.1212
 |
Component:289; Jaccard:0.14411
 |
Component:031; Jaccard:0.14927
 |
Component:364; Jaccard:0.15861
 |
Component:193; Jaccard:0.15696
 |
Component:193; Jaccard:0.12884
 |
Correlation:0.055663
  |
Correlation:0.25656
  |
Correlation:0.25656
  |
Correlation:0.15646
  |
Correlation:0.43402
  |
Correlation:0.22178
  |
Correlation:0.22178
  |
Component:239; Jaccard:0.10993
 |
Component:399; Jaccard:0.12001
 |
Component:364; Jaccard:0.12791
 |
Component:364; Jaccard:0.14528
 |
Component:031; Jaccard:0.13936
 |
Component:354; Jaccard:0.13275
 |
Component:354; Jaccard:0.11977
 |
Correlation:0.20999
  |
Correlation:0.18911
  |
Correlation:0.43402
  |
Correlation:0.43402
  |
Correlation:0.15646
  |
Correlation:0.35527
  |
Correlation:0.35527
  |
Component:354; Jaccard:0.10279
 |
Component:239; Jaccard:0.11895
 |
Component:354; Jaccard:0.12758
 |
Component:354; Jaccard:0.13599
 |
Component:354; Jaccard:0.13936
 |
Component:350; Jaccard:0.12404
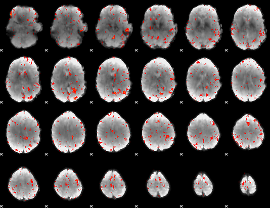 |
Component:350; Jaccard:0.1122
 |
Correlation:0.35527
  |
Correlation:0.20999
  |
Correlation:0.35527
  |
Correlation:0.35527
  |
Correlation:0.35527
  |
Correlation:0.36101
  |
Correlation:0.36101
  |
Component:135; Jaccard:0.10218
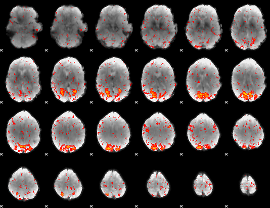 |
Component:293; Jaccard:0.11838
 |
Component:348; Jaccard:0.12667
 |
Component:350; Jaccard:0.12705
 |
Component:350; Jaccard:0.12972
 |
Component:031; Jaccard:0.12345
 |
Component:259; Jaccard:0.10775
 |
Correlation:0.1789
  |
Correlation:0.1648
  |
Correlation:0.055663
  |
Correlation:0.36101
  |
Correlation:0.36101
  |
Correlation:0.15646
  |
Correlation:0.33926
  |
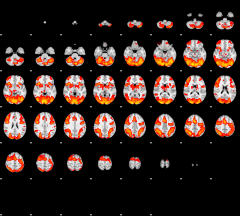
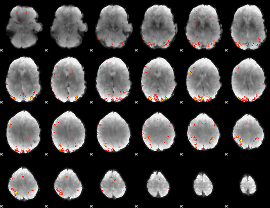
Cope: 02:"REWARD" BACK
|
GLM result group-wise
 |
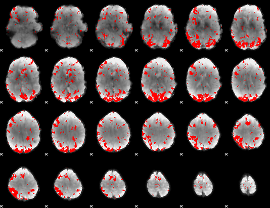
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
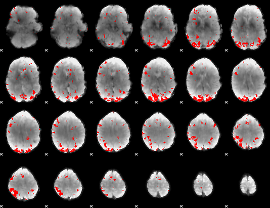
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:291; Jaccard:0.17807
 |
Component:291; Jaccard:0.20993
 |
Component:291; Jaccard:0.23039
 |
Component:291; Jaccard:0.23451
 |
Component:399; Jaccard:0.21817
 |
Component:399; Jaccard:0.2014
 |
Component:399; Jaccard:0.17162
 |
Correlation:0.068705
  |
Correlation:0.068705
  |
Correlation:0.068705
  |
Correlation:0.068705
  |
Correlation:0.21271
  |
Correlation:0.21271
  |
Correlation:0.21271
  |
Component:137; Jaccard:0.16962
 |
Component:137; Jaccard:0.19828
 |
Component:137; Jaccard:0.21781
 |
Component:137; Jaccard:0.22095
 |
Component:291; Jaccard:0.2147
 |
Component:291; Jaccard:0.18068
 |
Component:291; Jaccard:0.14132
 |
Correlation:0.0096251
  |
Correlation:0.0096251
  |
Correlation:0.0096251
  |
Correlation:0.0096251
  |
Correlation:0.068705
  |
Correlation:0.068705
  |
Correlation:0.068705
  |
Component:204; Jaccard:0.15146
 |
Component:204; Jaccard:0.17427
 |
Component:399; Jaccard:0.20239
 |
Component:399; Jaccard:0.21916
 |
Component:137; Jaccard:0.20093
 |
Component:137; Jaccard:0.17295
 |
Component:137; Jaccard:0.13657
 |
Correlation:-0.060198
  |
Correlation:-0.060198
  |
Correlation:0.21271
  |
Correlation:0.21271
  |
Correlation:0.0096251
  |
Correlation:0.0096251
  |
Correlation:0.0096251
  |
Component:399; Jaccard:0.14166
 |
Component:399; Jaccard:0.174
 |
Component:204; Jaccard:0.18623
 |
Component:289; Jaccard:0.18184
 |
Component:289; Jaccard:0.17406
 |
Component:289; Jaccard:0.15179
 |
Component:289; Jaccard:0.1243
 |
Correlation:0.21271
  |
Correlation:0.21271
  |
Correlation:-0.060198
  |
Correlation:0.029212
  |
Correlation:0.029212
  |
Correlation:0.029212
  |
Correlation:0.029212
  |
Component:289; Jaccard:0.13389
 |
Component:289; Jaccard:0.15651
 |
Component:289; Jaccard:0.17482
 |
Component:204; Jaccard:0.18123
 |
Component:204; Jaccard:0.1618
 |
Component:204; Jaccard:0.13065
 |
Component:204; Jaccard:0.097355
 |
Correlation:0.029212
  |
Correlation:0.029212
  |
Correlation:0.029212
  |
Correlation:-0.060198
  |
Correlation:-0.060198
  |
Correlation:-0.060198
  |
Correlation:-0.060198
  |
Component:253; Jaccard:0.12082
 |
Component:253; Jaccard:0.13283
 |
Component:253; Jaccard:0.13854
 |
Component:253; Jaccard:0.13516
 |
Component:253; Jaccard:0.12292
 |
Component:364; Jaccard:0.10643
 |
Component:364; Jaccard:0.088477
 |
Correlation:0.20615
  |
Correlation:0.20615
  |
Correlation:0.20615
  |
Correlation:0.20615
  |
Correlation:0.20615
  |
Correlation:0.011792
  |
Correlation:0.011792
  |
Component:135; Jaccard:0.11605
 |
Component:154; Jaccard:0.12793
 |
Component:154; Jaccard:0.13341
 |
Component:364; Jaccard:0.13077
 |
Component:364; Jaccard:0.11938
 |
Component:253; Jaccard:0.10582
 |
Component:253; Jaccard:0.085096
 |
Correlation:-0.038142
  |
Correlation:0.071665
  |
Correlation:0.071665
  |
Correlation:0.011792
  |
Correlation:0.011792
  |
Correlation:0.20615
  |
Correlation:0.20615
  |
Component:193; Jaccard:0.11527
 |
Component:135; Jaccard:0.12544
 |
Component:364; Jaccard:0.12739
 |
Component:154; Jaccard:0.13061
 |
Component:154; Jaccard:0.11902
 |
Component:154; Jaccard:0.1054
 |
Component:154; Jaccard:0.083109
 |
Correlation:-0.30776
  |
Correlation:-0.038142
  |
Correlation:0.011792
  |
Correlation:0.071665
  |
Correlation:0.071665
  |
Correlation:0.071665
  |
Correlation:0.071665
  |
Component:154; Jaccard:0.11446
 |
Component:193; Jaccard:0.11757
 |
Component:135; Jaccard:0.12709
 |
Component:135; Jaccard:0.11939
 |
Component:135; Jaccard:0.099655
 |
Component:033; Jaccard:0.082914
 |
Component:033; Jaccard:0.067186
 |
Correlation:0.071665
  |
Correlation:-0.30776
  |
Correlation:-0.038142
  |
Correlation:-0.038142
  |
Correlation:-0.038142
  |
Correlation:-0.044348
  |
Correlation:-0.044348
  |
Component:031; Jaccard:0.112
 |
Component:364; Jaccard:0.11494
 |
Component:033; Jaccard:0.11467
 |
Component:033; Jaccard:0.10999
 |
Component:033; Jaccard:0.098928
 |
Component:350; Jaccard:0.077022
 |
Component:350; Jaccard:0.062302
 |
Correlation:-0.13604
  |
Correlation:0.011792
  |
Correlation:-0.044348
  |
Correlation:-0.044348
  |
Correlation:-0.044348
  |
Correlation:0.056344
  |
Correlation:0.056344
  |
Cope: 03:"PUNISH-REWARD" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
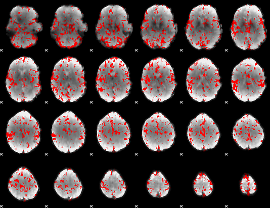 |
GLM binary result thresholded by 2.5
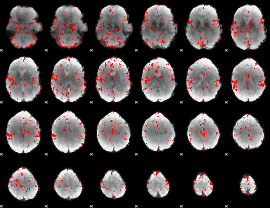 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
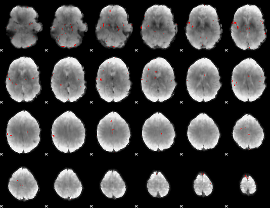 |
GLM Z-score result thresholded by 1
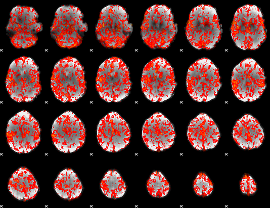 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
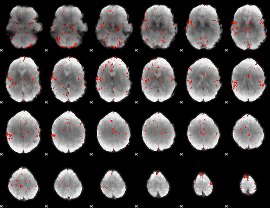 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
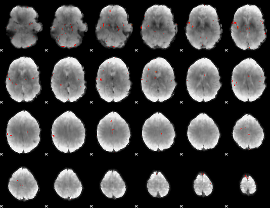 |
Component:293; Jaccard:0.11879
 |
Component:293; Jaccard:0.13631
 |
Component:076; Jaccard:0.16064
 |
Component:076; Jaccard:0.16789
 |
Component:076; Jaccard:0.13674
 |
Component:076; Jaccard:0.086793
 |
Component:076; Jaccard:0.041332
 |
Correlation:0.27132
  |
Correlation:0.27132
  |
Correlation:0.28621
  |
Correlation:0.28621
  |
Correlation:0.28621
  |
Correlation:0.28621
  |
Correlation:0.28621
  |
Component:286; Jaccard:0.11416
 |
Component:076; Jaccard:0.13612
 |
Component:286; Jaccard:0.14657
 |
Component:131; Jaccard:0.15726
 |
Component:131; Jaccard:0.13359
 |
Component:131; Jaccard:0.080234
 |
Component:131; Jaccard:0.036002
 |
Correlation:0.25627
  |
Correlation:0.28621
  |
Correlation:0.25627
  |
Correlation:0.31016
  |
Correlation:0.31016
  |
Correlation:0.31016
  |
Correlation:0.31016
  |
Component:076; Jaccard:0.10958
 |
Component:286; Jaccard:0.13434
 |
Component:131; Jaccard:0.14637
 |
Component:286; Jaccard:0.13944
 |
Component:286; Jaccard:0.10414
 |
Component:286; Jaccard:0.061993
 |
Component:286; Jaccard:0.027572
 |
Correlation:0.28621
  |
Correlation:0.25627
  |
Correlation:0.31016
  |
Correlation:0.25627
  |
Correlation:0.25627
  |
Correlation:0.25627
  |
Correlation:0.25627
  |
Component:055; Jaccard:0.10635
 |
Component:131; Jaccard:0.12273
 |
Component:293; Jaccard:0.14561
 |
Component:293; Jaccard:0.13364
 |
Component:293; Jaccard:0.098565
 |
Component:293; Jaccard:0.053079
 |
Component:019; Jaccard:0.026707
 |
Correlation:0.2766
  |
Correlation:0.31016
  |
Correlation:0.27132
  |
Correlation:0.27132
  |
Correlation:0.27132
  |
Correlation:0.27132
  |
Correlation:0.35308
  |
Component:221; Jaccard:0.10406
 |
Component:055; Jaccard:0.12085
 |
Component:055; Jaccard:0.12547
 |
Component:055; Jaccard:0.11346
 |
Component:055; Jaccard:0.077957
 |
Component:019; Jaccard:0.047075
 |
Component:293; Jaccard:0.02248
 |
Correlation:0.24286
  |
Correlation:0.2766
  |
Correlation:0.2766
  |
Correlation:0.2766
  |
Correlation:0.2766
  |
Correlation:0.35308
  |
Correlation:0.27132
  |
Component:087; Jaccard:0.097551
 |
Component:221; Jaccard:0.11504
 |
Component:221; Jaccard:0.11781
 |
Component:221; Jaccard:0.10386
 |
Component:120; Jaccard:0.073378
 |
Component:055; Jaccard:0.040753
 |
Component:256; Jaccard:0.019629
 |
Correlation:0.2382
  |
Correlation:0.24286
  |
Correlation:0.24286
  |
Correlation:0.24286
  |
Correlation:0.31065
  |
Correlation:0.2766
  |
Correlation:0.19062
  |
Component:131; Jaccard:0.09697
 |
Component:087; Jaccard:0.1101
 |
Component:087; Jaccard:0.11517
 |
Component:087; Jaccard:0.10302
 |
Component:087; Jaccard:0.07304
 |
Component:256; Jaccard:0.040662
 |
Component:371; Jaccard:0.019357
 |
Correlation:0.31016
  |
Correlation:0.2382
  |
Correlation:0.2382
  |
Correlation:0.2382
  |
Correlation:0.2382
  |
Correlation:0.19062
  |
Correlation:0.15897
  |
Component:218; Jaccard:0.096635
 |
Component:218; Jaccard:0.10431
 |
Component:120; Jaccard:0.1059
 |
Component:120; Jaccard:0.098378
 |
Component:221; Jaccard:0.069122
 |
Component:087; Jaccard:0.040621
 |
Component:120; Jaccard:0.018273
 |
Correlation:0.2762
  |
Correlation:0.2762
  |
Correlation:0.31065
  |
Correlation:0.31065
  |
Correlation:0.24286
  |
Correlation:0.2382
  |
Correlation:0.31065
  |
Component:239; Jaccard:0.096241
 |
Component:239; Jaccard:0.10132
 |
Component:218; Jaccard:0.10449
 |
Component:218; Jaccard:0.089062
 |
Component:256; Jaccard:0.068048
 |
Component:371; Jaccard:0.040491
 |
Component:066; Jaccard:0.017929
 |
Correlation:0.25266
  |
Correlation:0.25266
  |
Correlation:0.2762
  |
Correlation:0.2762
  |
Correlation:0.19062
  |
Correlation:0.15897
  |
Correlation:0.25963
  |
Component:193; Jaccard:0.088505
 |
Component:120; Jaccard:0.099134
 |
Component:239; Jaccard:0.0963
 |
Component:256; Jaccard:0.087961
 |
Component:019; Jaccard:0.06734
 |
Component:120; Jaccard:0.039811
 |
Component:079; Jaccard:0.017206
 |
Correlation:0.31142
  |
Correlation:0.31065
  |
Correlation:0.25266
  |
Correlation:0.19062
  |
Correlation:0.35308
  |
Correlation:0.31065
  |
Correlation:0.070348
  |
Cope: 04:"neg_PUNISH" BACK
|
GLM result group-wise
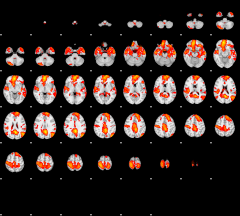 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
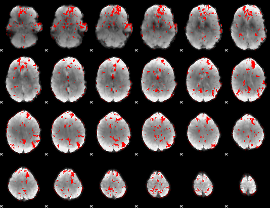 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
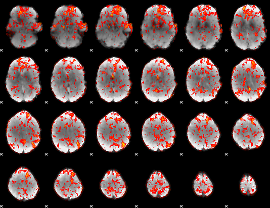 |
GLM Z-score result thresholded by 1.5
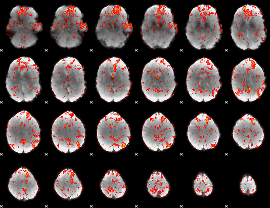 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
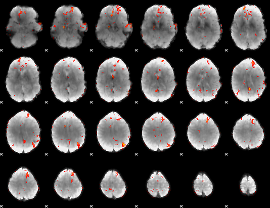 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
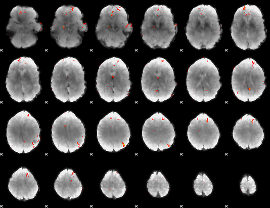 |
Component:243; Jaccard:0.12576
 |
Component:243; Jaccard:0.13861
 |
Component:243; Jaccard:0.13839
 |
Component:243; Jaccard:0.12629
 |
Component:243; Jaccard:0.10136
 |
Component:243; Jaccard:0.073295
 |
Component:252; Jaccard:0.047615
 |
Correlation:0.13308
  |
Correlation:0.13308
  |
Correlation:0.13308
  |
Correlation:0.13308
  |
Correlation:0.13308
  |
Correlation:0.13308
  |
Correlation:0.017292
  |
Component:370; Jaccard:0.1169
 |
Component:370; Jaccard:0.12449
 |
Component:370; Jaccard:0.12361
 |
Component:069; Jaccard:0.11767
 |
Component:069; Jaccard:0.099681
 |
Component:370; Jaccard:0.072774
 |
Component:370; Jaccard:0.047085
 |
Correlation:0.084817
  |
Correlation:0.084817
  |
Correlation:0.084817
  |
Correlation:0.41672
  |
Correlation:0.41672
  |
Correlation:0.084817
  |
Correlation:0.084817
  |
Component:252; Jaccard:0.10353
 |
Component:252; Jaccard:0.11326
 |
Component:069; Jaccard:0.11811
 |
Component:370; Jaccard:0.11147
 |
Component:370; Jaccard:0.094406
 |
Component:252; Jaccard:0.069803
 |
Component:243; Jaccard:0.04691
 |
Correlation:0.017292
  |
Correlation:0.017292
  |
Correlation:0.41672
  |
Correlation:0.084817
  |
Correlation:0.084817
  |
Correlation:0.017292
  |
Correlation:0.13308
  |
Component:194; Jaccard:0.09902
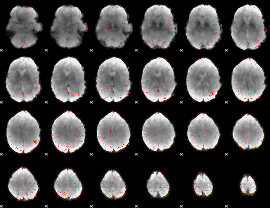 |
Component:194; Jaccard:0.11258
 |
Component:252; Jaccard:0.1162
 |
Component:252; Jaccard:0.11014
 |
Component:252; Jaccard:0.09266
 |
Component:069; Jaccard:0.067009
 |
Component:069; Jaccard:0.041715
 |
Correlation:0.037561
  |
Correlation:0.037561
  |
Correlation:0.017292
  |
Correlation:0.017292
  |
Correlation:0.017292
  |
Correlation:0.41672
  |
Correlation:0.41672
  |
Component:251; Jaccard:0.096456
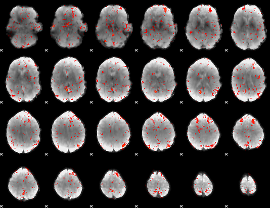 |
Component:069; Jaccard:0.10431
 |
Component:194; Jaccard:0.1119
 |
Component:251; Jaccard:0.095808
 |
Component:251; Jaccard:0.079633
 |
Component:251; Jaccard:0.056498
 |
Component:251; Jaccard:0.035607
 |
Correlation:0.088239
  |
Correlation:0.41672
  |
Correlation:0.037561
  |
Correlation:0.088239
  |
Correlation:0.088239
  |
Correlation:0.088239
  |
Correlation:0.088239
  |
Component:025; Jaccard:0.095036
 |
Component:251; Jaccard:0.10254
 |
Component:251; Jaccard:0.10233
 |
Component:194; Jaccard:0.094678
 |
Component:025; Jaccard:0.070479
 |
Component:116; Jaccard:0.053932
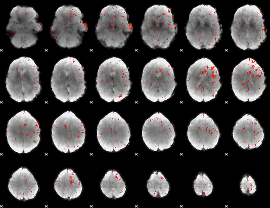 |
Component:116; Jaccard:0.034687
 |
Correlation:0.15352
  |
Correlation:0.088239
  |
Correlation:0.088239
  |
Correlation:0.037561
  |
Correlation:0.15352
  |
Correlation:0.16759
  |
Correlation:0.16759
  |
Component:018; Jaccard:0.087913
 |
Component:025; Jaccard:0.098961
 |
Component:018; Jaccard:0.097186
 |
Component:018; Jaccard:0.087785
 |
Component:116; Jaccard:0.066629
 |
Component:025; Jaccard:0.050446
 |
Component:025; Jaccard:0.032763
 |
Correlation:0.1267
  |
Correlation:0.15352
  |
Correlation:0.1267
  |
Correlation:0.1267
  |
Correlation:0.16759
  |
Correlation:0.15352
  |
Correlation:0.15352
  |
Component:069; Jaccard:0.087428
 |
Component:018; Jaccard:0.095392
 |
Component:025; Jaccard:0.096553
 |
Component:025; Jaccard:0.086388
 |
Component:194; Jaccard:0.06633
 |
Component:164; Jaccard:0.046607
 |
Component:164; Jaccard:0.029409
 |
Correlation:0.41672
  |
Correlation:0.1267
  |
Correlation:0.15352
  |
Correlation:0.15352
  |
Correlation:0.037561
  |
Correlation:-0.08092
  |
Correlation:-0.08092
  |
Component:221; Jaccard:0.084298
 |
Component:281; Jaccard:0.087623
 |
Component:281; Jaccard:0.089608
 |
Component:118; Jaccard:0.080591
 |
Component:118; Jaccard:0.065833
 |
Component:118; Jaccard:0.044073
 |
Component:041; Jaccard:0.02908
 |
Correlation:-0.006097
  |
Correlation:0.054653
  |
Correlation:0.054653
  |
Correlation:0.092307
  |
Correlation:0.092307
  |
Correlation:0.092307
  |
Correlation:0.046573
  |
Component:281; Jaccard:0.076291
 |
Component:118; Jaccard:0.079972
 |
Component:118; Jaccard:0.084412
 |
Component:281; Jaccard:0.078161
 |
Component:018; Jaccard:0.065215
 |
Component:038; Jaccard:0.041897
 |
Component:118; Jaccard:0.027302
 |
Correlation:0.054653
  |
Correlation:0.092307
  |
Correlation:0.092307
  |
Correlation:0.054653
  |
Correlation:0.1267
  |
Correlation:0.37734
  |
Correlation:0.092307
  |
Cope: 05:"neg_REWARD" BACK
|
GLM result group-wise
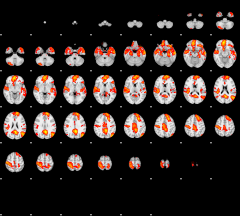 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
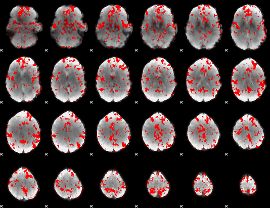 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
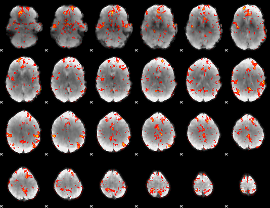 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:221; Jaccard:0.14026
 |
Component:221; Jaccard:0.1646
 |
Component:221; Jaccard:0.1855
 |
Component:243; Jaccard:0.19705
 |
Component:243; Jaccard:0.19844
 |
Component:243; Jaccard:0.17401
 |
Component:243; Jaccard:0.13065
 |
Correlation:0.40687
  |
Correlation:0.40687
  |
Correlation:0.40687
  |
Correlation:0.33502
  |
Correlation:0.33502
  |
Correlation:0.33502
  |
Correlation:0.33502
  |
Component:243; Jaccard:0.11766
 |
Component:243; Jaccard:0.14603
 |
Component:243; Jaccard:0.17616
 |
Component:221; Jaccard:0.18834
 |
Component:221; Jaccard:0.16981
 |
Component:252; Jaccard:0.14504
 |
Component:252; Jaccard:0.11088
 |
Correlation:0.33502
  |
Correlation:0.33502
  |
Correlation:0.33502
  |
Correlation:0.40687
  |
Correlation:0.40687
  |
Correlation:0.13506
  |
Correlation:0.13506
  |
Component:256; Jaccard:0.10562
 |
Component:252; Jaccard:0.12336
 |
Component:252; Jaccard:0.1466
 |
Component:252; Jaccard:0.16301
 |
Component:252; Jaccard:0.16356
 |
Component:221; Jaccard:0.13502
 |
Component:221; Jaccard:0.094897
 |
Correlation:0.45974
  |
Correlation:0.13506
  |
Correlation:0.13506
  |
Correlation:0.13506
  |
Correlation:0.13506
  |
Correlation:0.40687
  |
Correlation:0.40687
  |
Component:252; Jaccard:0.10169
 |
Component:256; Jaccard:0.12334
 |
Component:256; Jaccard:0.13922
 |
Component:256; Jaccard:0.14608
 |
Component:256; Jaccard:0.13782
 |
Component:256; Jaccard:0.11614
 |
Component:256; Jaccard:0.081276
 |
Correlation:0.13506
  |
Correlation:0.45974
  |
Correlation:0.45974
  |
Correlation:0.45974
  |
Correlation:0.45974
  |
Correlation:0.45974
  |
Correlation:0.45974
  |
Component:076; Jaccard:0.098116
 |
Component:076; Jaccard:0.10543
 |
Component:370; Jaccard:0.1075
 |
Component:370; Jaccard:0.10901
 |
Component:370; Jaccard:0.10099
 |
Component:025; Jaccard:0.083119
 |
Component:370; Jaccard:0.058877
 |
Correlation:0.30924
  |
Correlation:0.30924
  |
Correlation:0.14503
  |
Correlation:0.14503
  |
Correlation:0.14503
  |
Correlation:0.32987
  |
Correlation:0.14503
  |
Component:295; Jaccard:0.091765
 |
Component:370; Jaccard:0.099703
 |
Component:076; Jaccard:0.10632
 |
Component:025; Jaccard:0.10563
 |
Component:025; Jaccard:0.098123
 |
Component:370; Jaccard:0.081867
 |
Component:025; Jaccard:0.057475
 |
Correlation:0.44274
  |
Correlation:0.14503
  |
Correlation:0.30924
  |
Correlation:0.32987
  |
Correlation:0.32987
  |
Correlation:0.14503
  |
Correlation:0.32987
  |
Component:370; Jaccard:0.088722
 |
Component:025; Jaccard:0.09684
 |
Component:025; Jaccard:0.10446
 |
Component:076; Jaccard:0.098889
 |
Component:268; Jaccard:0.087336
 |
Component:268; Jaccard:0.070571
 |
Component:268; Jaccard:0.052054
 |
Correlation:0.14503
  |
Correlation:0.32987
  |
Correlation:0.32987
  |
Correlation:0.30924
  |
Correlation:0.24202
  |
Correlation:0.24202
  |
Correlation:0.24202
  |
Component:025; Jaccard:0.086242
 |
Component:295; Jaccard:0.096477
 |
Component:268; Jaccard:0.097131
 |
Component:268; Jaccard:0.094858
 |
Component:147; Jaccard:0.083846
 |
Component:147; Jaccard:0.067127
 |
Component:147; Jaccard:0.048744
 |
Correlation:0.32987
  |
Correlation:0.44274
  |
Correlation:0.24202
  |
Correlation:0.24202
  |
Correlation:0.010617
  |
Correlation:0.010617
  |
Correlation:0.010617
  |
Component:268; Jaccard:0.085435
 |
Component:268; Jaccard:0.092124
 |
Component:295; Jaccard:0.096792
 |
Component:295; Jaccard:0.091344
 |
Component:076; Jaccard:0.081971
 |
Component:295; Jaccard:0.064516
 |
Component:164; Jaccard:0.047504
 |
Correlation:0.24202
  |
Correlation:0.24202
  |
Correlation:0.44274
  |
Correlation:0.44274
  |
Correlation:0.30924
  |
Correlation:0.44274
  |
Correlation:0.061384
  |
Component:269; Jaccard:0.084048
 |
Component:066; Jaccard:0.086814
 |
Component:066; Jaccard:0.091635
 |
Component:147; Jaccard:0.089076
 |
Component:295; Jaccard:0.080716
 |
Component:066; Jaccard:0.063118
 |
Component:251; Jaccard:0.046685
 |
Correlation:0.23045
  |
Correlation:0.40117
  |
Correlation:0.40117
  |
Correlation:0.010617
  |
Correlation:0.44274
  |
Correlation:0.40117
  |
Correlation:-0.036667
  |
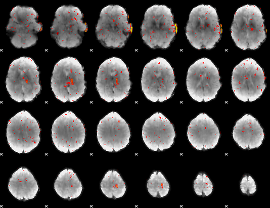
Cope: 06:"REWARD-PUNISH" BACK
|
GLM result group-wise
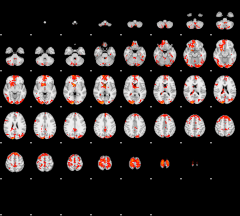 |
GLM binary result thresholded by 1
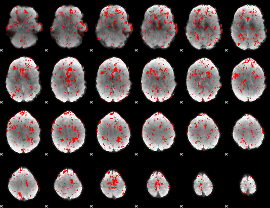 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:118; Jaccard:0.085935
 |
Component:118; Jaccard:0.090802
 |
Component:118; Jaccard:0.082374
 |
Component:375; Jaccard:0.073558
 |
Component:375; Jaccard:0.061394
 |
Component:375; Jaccard:0.040569
 |
Component:375; Jaccard:0.020088
 |
Correlation:0.03135
  |
Correlation:0.03135
  |
Correlation:0.03135
  |
Correlation:0.12781
  |
Correlation:0.12781
  |
Correlation:0.12781
  |
Correlation:0.12781
  |
Component:186; Jaccard:0.082351
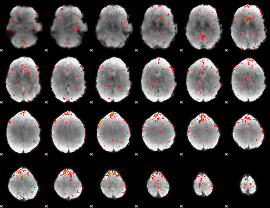 |
Component:356; Jaccard:0.08136
 |
Component:375; Jaccard:0.075576
 |
Component:118; Jaccard:0.056602
 |
Component:356; Jaccard:0.031334
 |
Component:045; Jaccard:0.01988
 |
Component:045; Jaccard:0.009807
 |
Correlation:0.0086253
  |
Correlation:0.049976
  |
Correlation:0.12781
  |
Correlation:0.03135
  |
Correlation:0.049976
  |
Correlation:0.041955
  |
Correlation:0.041955
  |
Component:005; Jaccard:0.078161
 |
Component:005; Jaccard:0.075075
 |
Component:356; Jaccard:0.073415
 |
Component:356; Jaccard:0.054727
 |
Component:045; Jaccard:0.031282
 |
Component:356; Jaccard:0.017225
 |
Component:356; Jaccard:0.007061
 |
Correlation:0.10847
  |
Correlation:0.10847
  |
Correlation:0.049976
  |
Correlation:0.049976
  |
Correlation:0.041955
  |
Correlation:0.049976
  |
Correlation:0.049976
  |
Component:356; Jaccard:0.077028
 |
Component:302; Jaccard:0.072783
 |
Component:302; Jaccard:0.065494
 |
Component:276; Jaccard:0.049064
 |
Component:118; Jaccard:0.029505
 |
Component:097; Jaccard:0.015102
 |
Component:116; Jaccard:0.006766
 |
Correlation:0.049976
  |
Correlation:0.18411
  |
Correlation:0.18411
  |
Correlation:0.046171
  |
Correlation:0.03135
  |
Correlation:0.2264
  |
Correlation:0.18803
  |
Component:049; Jaccard:0.074414
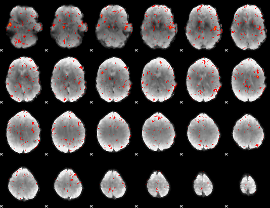 |
Component:186; Jaccard:0.072327
 |
Component:276; Jaccard:0.064654
 |
Component:302; Jaccard:0.047903
 |
Component:276; Jaccard:0.027933
 |
Component:049; Jaccard:0.014746
 |
Component:097; Jaccard:0.006751
 |
Correlation:0.086093
  |
Correlation:0.0086253
  |
Correlation:0.046171
  |
Correlation:0.18411
  |
Correlation:0.046171
  |
Correlation:0.086093
  |
Correlation:0.2264
  |
Component:302; Jaccard:0.068498
 |
Component:276; Jaccard:0.070962
 |
Component:049; Jaccard:0.0607
 |
Component:049; Jaccard:0.043208
 |
Component:302; Jaccard:0.026925
 |
Component:048; Jaccard:0.014245
 |
Component:049; Jaccard:0.006068
 |
Correlation:0.18411
  |
Correlation:0.046171
  |
Correlation:0.086093
  |
Correlation:0.086093
  |
Correlation:0.18411
  |
Correlation:0.17549
  |
Correlation:0.086093
  |
Component:276; Jaccard:0.065735
 |
Component:049; Jaccard:0.068908
 |
Component:005; Jaccard:0.058451
 |
Component:045; Jaccard:0.042546
 |
Component:049; Jaccard:0.026361
 |
Component:116; Jaccard:0.014149
 |
Component:163; Jaccard:0.006037
 |
Correlation:0.046171
  |
Correlation:0.086093
  |
Correlation:0.10847
  |
Correlation:0.041955
  |
Correlation:0.086093
  |
Correlation:0.18803
  |
Correlation:-0.012483
  |
Component:240; Jaccard:0.064722
 |
Component:375; Jaccard:0.067704
 |
Component:186; Jaccard:0.052981
 |
Component:116; Jaccard:0.040132
 |
Component:116; Jaccard:0.026095
 |
Component:276; Jaccard:0.012441
 |
Component:077; Jaccard:0.005713
 |
Correlation:0.068163
  |
Correlation:0.12781
  |
Correlation:0.0086253
  |
Correlation:0.18803
  |
Correlation:0.18803
  |
Correlation:0.046171
  |
Correlation:0.14334
  |
Component:024; Jaccard:0.062144
 |
Component:024; Jaccard:0.060572
 |
Component:045; Jaccard:0.052879
 |
Component:005; Jaccard:0.036824
 |
Component:048; Jaccard:0.025981
 |
Component:118; Jaccard:0.012017
 |
Component:276; Jaccard:0.005412
 |
Correlation:0.16251
  |
Correlation:0.16251
  |
Correlation:0.041955
  |
Correlation:0.10847
  |
Correlation:0.17549
  |
Correlation:0.03135
  |
Correlation:0.046171
  |
Component:217; Jaccard:0.061777
 |
Component:240; Jaccard:0.057577
 |
Component:116; Jaccard:0.049514
 |
Component:048; Jaccard:0.035773
 |
Component:097; Jaccard:0.021005
 |
Component:077; Jaccard:0.011743
 |
Component:048; Jaccard:0.005399
 |
Correlation:0.16665
  |
Correlation:0.068163
  |
Correlation:0.18803
  |
Correlation:0.17549
  |
Correlation:0.2264
  |
Correlation:0.14334
  |
Correlation:0.17549
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































