Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"FACES" BACK
|
GLM result group-wise
 |
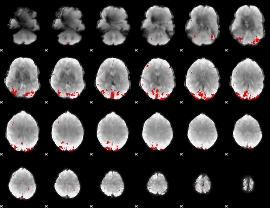
GLM binary result thresholded by 1
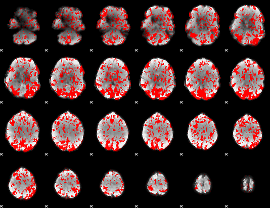 |
GLM binary result thresholded by 1.5
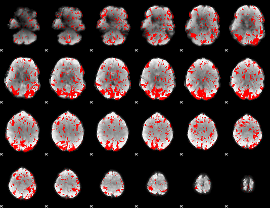 |
GLM binary result thresholded by 2
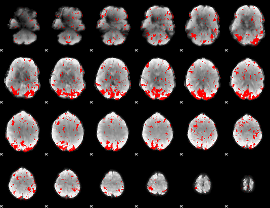 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
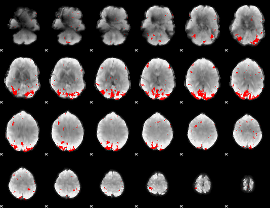 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
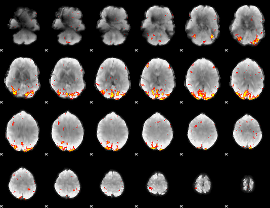
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
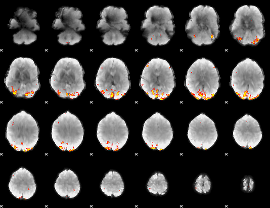
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:118; Jaccard:0.1306
 |
Component:026; Jaccard:0.19024
 |
Component:026; Jaccard:0.2737
 |
Component:026; Jaccard:0.34245
 |
Component:026; Jaccard:0.37304
 |
Component:026; Jaccard:0.36023
 |
Component:026; Jaccard:0.3353
 |
Correlation:0.21169
  |
Correlation:0.35705
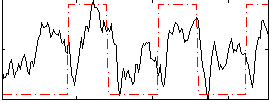 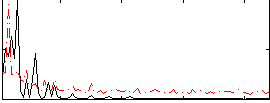 |
Correlation:0.35705
  |
Correlation:0.35705
  |
Correlation:0.35705
  |
Correlation:0.35705
  |
Correlation:0.35705
  |
Component:026; Jaccard:0.1246
 |
Component:118; Jaccard:0.1708
 |
Component:118; Jaccard:0.20245
 |
Component:118; Jaccard:0.21155
 |
Component:096; Jaccard:0.19718
 |
Component:096; Jaccard:0.18658
 |
Component:096; Jaccard:0.16963
 |
Correlation:0.35705
  |
Correlation:0.21169
  |
Correlation:0.21169
  |
Correlation:0.21169
  |
Correlation:0.44556
  |
Correlation:0.44556
  |
Correlation:0.44556
  |
Component:258; Jaccard:0.12165
 |
Component:258; Jaccard:0.15204
 |
Component:258; Jaccard:0.17275
 |
Component:096; Jaccard:0.19347
 |
Component:118; Jaccard:0.18946
 |
Component:350; Jaccard:0.16957
 |
Component:350; Jaccard:0.15324
 |
Correlation:-0.036691
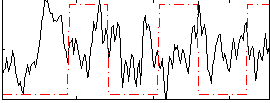 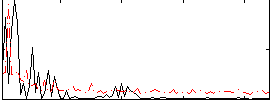 |
Correlation:-0.036691
  |
Correlation:-0.036691
  |
Correlation:0.44556
  |
Correlation:0.21169
  |
Correlation:-0.22631
  |
Correlation:-0.22631
  |
Component:096; Jaccard:0.095227
 |
Component:096; Jaccard:0.13095
 |
Component:096; Jaccard:0.16961
 |
Component:258; Jaccard:0.16979
 |
Component:350; Jaccard:0.17521
 |
Component:118; Jaccard:0.164
 |
Component:118; Jaccard:0.13829
 |
Correlation:0.44556
  |
Correlation:0.44556
  |
Correlation:0.44556
  |
Correlation:-0.036691
  |
Correlation:-0.22631
  |
Correlation:0.21169
  |
Correlation:0.21169
  |
Component:087; Jaccard:0.088151
 |
Component:350; Jaccard:0.11308
 |
Component:350; Jaccard:0.14403
 |
Component:350; Jaccard:0.16689
 |
Component:258; Jaccard:0.15583
 |
Component:258; Jaccard:0.13793
 |
Component:258; Jaccard:0.12248
 |
Correlation:-0.10788
  |
Correlation:-0.22631
  |
Correlation:-0.22631
  |
Correlation:-0.22631
  |
Correlation:-0.036691
  |
Correlation:-0.036691
  |
Correlation:-0.036691
  |
Component:272; Jaccard:0.08707
 |
Component:053; Jaccard:0.10328
 |
Component:053; Jaccard:0.11939
 |
Component:053; Jaccard:0.12461
 |
Component:010; Jaccard:0.11952
 |
Component:010; Jaccard:0.10952
 |
Component:010; Jaccard:0.095786
 |
Correlation:0.39004
  |
Correlation:0.031282
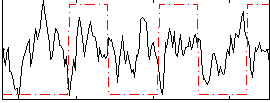  |
Correlation:0.031282
  |
Correlation:0.031282
  |
Correlation:0.67259
  |
Correlation:0.67259
  |
Correlation:0.67259
  |
Component:183; Jaccard:0.086998
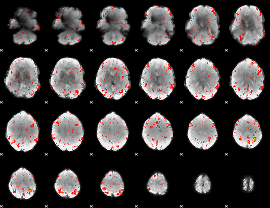 |
Component:087; Jaccard:0.10275
 |
Component:010; Jaccard:0.11787
 |
Component:010; Jaccard:0.12307
 |
Component:053; Jaccard:0.11902
 |
Component:053; Jaccard:0.10483
 |
Component:053; Jaccard:0.089717
 |
Correlation:0.38952
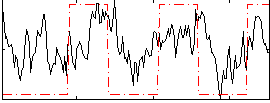  |
Correlation:-0.10788
  |
Correlation:0.67259
  |
Correlation:0.67259
  |
Correlation:0.031282
  |
Correlation:0.031282
  |
Correlation:0.031282
  |
Component:053; Jaccard:0.083625
 |
Component:010; Jaccard:0.10185
 |
Component:087; Jaccard:0.11647
 |
Component:087; Jaccard:0.12039
 |
Component:087; Jaccard:0.11416
 |
Component:087; Jaccard:0.09964
 |
Component:087; Jaccard:0.08395
 |
Correlation:0.031282
  |
Correlation:0.67259
  |
Correlation:-0.10788
  |
Correlation:-0.10788
  |
Correlation:-0.10788
  |
Correlation:-0.10788
  |
Correlation:-0.10788
  |
Component:350; Jaccard:0.083462
 |
Component:272; Jaccard:0.099597
 |
Component:272; Jaccard:0.10595
 |
Component:136; Jaccard:0.11252
 |
Component:136; Jaccard:0.10669
 |
Component:136; Jaccard:0.094843
 |
Component:136; Jaccard:0.081112
 |
Correlation:-0.22631
  |
Correlation:0.39004
  |
Correlation:0.39004
  |
Correlation:0.3122
 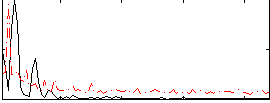 |
Correlation:0.3122
  |
Correlation:0.3122
  |
Correlation:0.3122
  |
Component:028; Jaccard:0.083456
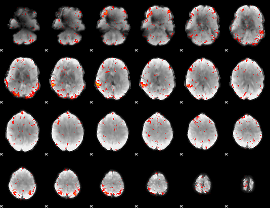 |
Component:183; Jaccard:0.094965
 |
Component:136; Jaccard:0.10551
 |
Component:272; Jaccard:0.10203
 |
Component:272; Jaccard:0.094236
 |
Component:272; Jaccard:0.083827
 |
Component:272; Jaccard:0.070829
 |
Correlation:0.30599
  |
Correlation:0.38952
  |
Correlation:0.3122
  |
Correlation:0.39004
  |
Correlation:0.39004
  |
Correlation:0.39004
  |
Correlation:0.39004
  |
Cope: 02:"SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
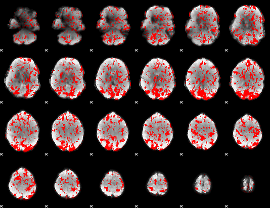 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
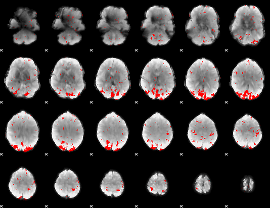 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
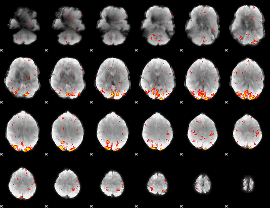 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:258; Jaccard:0.13955
 |
Component:350; Jaccard:0.18418
 |
Component:350; Jaccard:0.24943
 |
Component:350; Jaccard:0.29501
 |
Component:350; Jaccard:0.28885
 |
Component:350; Jaccard:0.25813
 |
Component:350; Jaccard:0.22111
 |
Correlation:0.15288
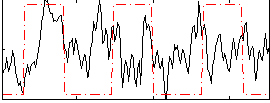 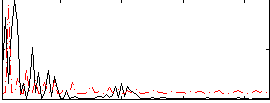 |
Correlation:0.41166
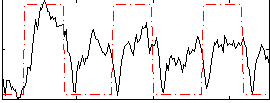 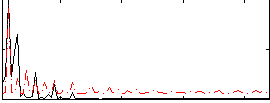 |
Correlation:0.41166
  |
Correlation:0.41166
  |
Correlation:0.41166
  |
Correlation:0.41166
  |
Correlation:0.41166
  |
Component:026; Jaccard:0.12893
 |
Component:026; Jaccard:0.17552
 |
Component:026; Jaccard:0.22363
 |
Component:026; Jaccard:0.25033
 |
Component:026; Jaccard:0.24519
 |
Component:026; Jaccard:0.21593
 |
Component:026; Jaccard:0.18297
 |
Correlation:-0.22831
  |
Correlation:-0.22831
  |
Correlation:-0.22831
  |
Correlation:-0.22831
  |
Correlation:-0.22831
  |
Correlation:-0.22831
  |
Correlation:-0.22831
  |
Component:350; Jaccard:0.12241
 |
Component:258; Jaccard:0.17138
 |
Component:258; Jaccard:0.18942
 |
Component:258; Jaccard:0.18128
 |
Component:258; Jaccard:0.1571
 |
Component:258; Jaccard:0.12798
 |
Component:258; Jaccard:0.10621
 |
Correlation:0.41166
  |
Correlation:0.15288
  |
Correlation:0.15288
  |
Correlation:0.15288
  |
Correlation:0.15288
  |
Correlation:0.15288
  |
Correlation:0.15288
  |
Component:118; Jaccard:0.12238
 |
Component:118; Jaccard:0.14774
 |
Component:118; Jaccard:0.16128
 |
Component:118; Jaccard:0.15564
 |
Component:118; Jaccard:0.13227
 |
Component:118; Jaccard:0.10626
 |
Component:053; Jaccard:0.084064
 |
Correlation:-0.068455
  |
Correlation:-0.068455
  |
Correlation:-0.068455
  |
Correlation:-0.068455
  |
Correlation:-0.068455
  |
Correlation:-0.068455
  |
Correlation:0.097323
  |
Component:053; Jaccard:0.098698
 |
Component:053; Jaccard:0.11945
 |
Component:053; Jaccard:0.13414
 |
Component:053; Jaccard:0.13622
 |
Component:053; Jaccard:0.12222
 |
Component:053; Jaccard:0.10076
 |
Component:118; Jaccard:0.082784
 |
Correlation:0.097323
  |
Correlation:0.097323
  |
Correlation:0.097323
  |
Correlation:0.097323
  |
Correlation:0.097323
  |
Correlation:0.097323
  |
Correlation:-0.068455
  |
Component:087; Jaccard:0.094395
 |
Component:087; Jaccard:0.10601
 |
Component:087; Jaccard:0.10964
 |
Component:087; Jaccard:0.10757
 |
Component:087; Jaccard:0.095463
 |
Component:096; Jaccard:0.082456
 |
Component:096; Jaccard:0.071767
 |
Correlation:0.30397
  |
Correlation:0.30397
  |
Correlation:0.30397
  |
Correlation:0.30397
  |
Correlation:0.30397
  |
Correlation:-0.26474
  |
Correlation:-0.26474
  |
Component:349; Jaccard:0.086954
 |
Component:349; Jaccard:0.086983
 |
Component:096; Jaccard:0.084428
 |
Component:096; Jaccard:0.092337
 |
Component:096; Jaccard:0.09106
 |
Component:087; Jaccard:0.080234
 |
Component:087; Jaccard:0.066988
 |
Correlation:0.11122
  |
Correlation:0.11122
  |
Correlation:-0.26474
  |
Correlation:-0.26474
  |
Correlation:-0.26474
  |
Correlation:0.30397
  |
Correlation:0.30397
  |
Component:298; Jaccard:0.075058
 |
Component:252; Jaccard:0.073525
 |
Component:136; Jaccard:0.07593
 |
Component:136; Jaccard:0.0789
 |
Component:136; Jaccard:0.074493
 |
Component:136; Jaccard:0.065763
 |
Component:067; Jaccard:0.054989
 |
Correlation:-0.062299
  |
Correlation:0.18309
  |
Correlation:0.088327
  |
Correlation:0.088327
  |
Correlation:0.088327
  |
Correlation:0.088327
  |
Correlation:0.15336
  |
Component:252; Jaccard:0.074395
 |
Component:298; Jaccard:0.071246
 |
Component:349; Jaccard:0.074126
 |
Component:067; Jaccard:0.060236
 |
Component:067; Jaccard:0.061758
 |
Component:067; Jaccard:0.059545
 |
Component:136; Jaccard:0.054651
 |
Correlation:0.18309
  |
Correlation:-0.062299
  |
Correlation:0.11122
  |
Correlation:0.15336
  |
Correlation:0.15336
  |
Correlation:0.15336
  |
Correlation:0.088327
  |
Component:198; Jaccard:0.071879
 |
Component:096; Jaccard:0.070214
 |
Component:210; Jaccard:0.061321
 |
Component:349; Jaccard:0.054487
 |
Component:004; Jaccard:0.040834
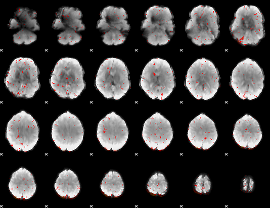 |
Component:004; Jaccard:0.037126
 |
Component:004; Jaccard:0.032485
 |
Correlation:0.20077
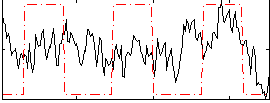  |
Correlation:-0.26474
  |
Correlation:0.039328
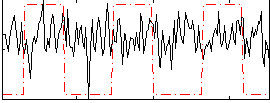  |
Correlation:0.11122
  |
Correlation:0.050314
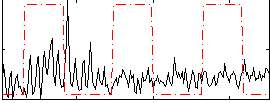  |
Correlation:0.050314
  |
Correlation:0.050314
  |
Cope: 03:"FACES-SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
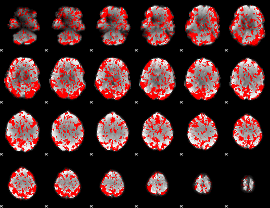 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
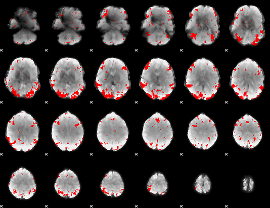 |
GLM binary result thresholded by 3.5
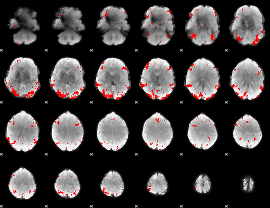 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
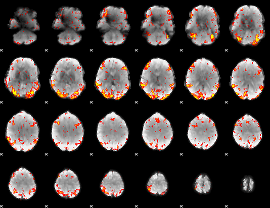 |
GLM Z-score result thresholded by 3
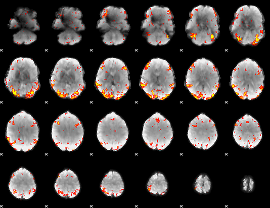 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
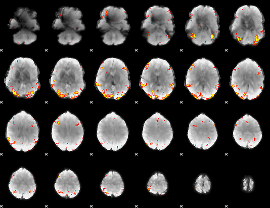 |
Component:374; Jaccard:0.12732
 |
Component:374; Jaccard:0.17242
 |
Component:374; Jaccard:0.2178
 |
Component:374; Jaccard:0.24625
 |
Component:374; Jaccard:0.25079
 |
Component:374; Jaccard:0.23642
 |
Component:010; Jaccard:0.22839
 |
Correlation:0.47676
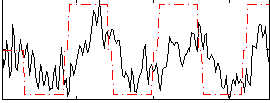  |
Correlation:0.47676
  |
Correlation:0.47676
  |
Correlation:0.47676
  |
Correlation:0.47676
  |
Correlation:0.47676
  |
Correlation:0.72261
  |
Component:028; Jaccard:0.119
 |
Component:028; Jaccard:0.15527
 |
Component:028; Jaccard:0.18997
 |
Component:010; Jaccard:0.21382
 |
Component:010; Jaccard:0.23232
 |
Component:010; Jaccard:0.23407
 |
Component:374; Jaccard:0.21457
 |
Correlation:0.35124
  |
Correlation:0.35124
  |
Correlation:0.35124
  |
Correlation:0.72261
  |
Correlation:0.72261
  |
Correlation:0.72261
  |
Correlation:0.47676
  |
Component:183; Jaccard:0.11591
 |
Component:183; Jaccard:0.14414
 |
Component:010; Jaccard:0.17682
 |
Component:028; Jaccard:0.21146
 |
Component:028; Jaccard:0.21485
 |
Component:096; Jaccard:0.20201
 |
Component:096; Jaccard:0.19572
 |
Correlation:0.43049
  |
Correlation:0.43049
  |
Correlation:0.72261
  |
Correlation:0.35124
  |
Correlation:0.35124
  |
Correlation:0.38527
  |
Correlation:0.38527
  |
Component:272; Jaccard:0.10493
 |
Component:010; Jaccard:0.13596
 |
Component:183; Jaccard:0.16754
 |
Component:183; Jaccard:0.18051
 |
Component:096; Jaccard:0.196
 |
Component:028; Jaccard:0.20144
 |
Component:028; Jaccard:0.17811
 |
Correlation:0.46145
  |
Correlation:0.72261
  |
Correlation:0.43049
  |
Correlation:0.43049
  |
Correlation:0.38527
  |
Correlation:0.35124
  |
Correlation:0.35124
  |
Component:283; Jaccard:0.1043
 |
Component:272; Jaccard:0.13291
 |
Component:272; Jaccard:0.15742
 |
Component:096; Jaccard:0.17606
 |
Component:272; Jaccard:0.18087
 |
Component:272; Jaccard:0.17761
 |
Component:272; Jaccard:0.16851
 |
Correlation:0.19281
  |
Correlation:0.46145
  |
Correlation:0.46145
  |
Correlation:0.38527
  |
Correlation:0.46145
  |
Correlation:0.46145
  |
Correlation:0.46145
  |
Component:358; Jaccard:0.10309
 |
Component:358; Jaccard:0.12802
 |
Component:358; Jaccard:0.14752
 |
Component:272; Jaccard:0.1752
 |
Component:183; Jaccard:0.1754
 |
Component:026; Jaccard:0.15957
 |
Component:026; Jaccard:0.15343
 |
Correlation:0.31489
  |
Correlation:0.31489
  |
Correlation:0.31489
  |
Correlation:0.46145
  |
Correlation:0.43049
  |
Correlation:0.31751
  |
Correlation:0.31751
  |
Component:010; Jaccard:0.099078
 |
Component:125; Jaccard:0.11923
 |
Component:096; Jaccard:0.14479
 |
Component:358; Jaccard:0.15299
 |
Component:026; Jaccard:0.15768
 |
Component:183; Jaccard:0.15862
 |
Component:183; Jaccard:0.13264
 |
Correlation:0.72261
  |
Correlation:0.47469
  |
Correlation:0.38527
  |
Correlation:0.31489
  |
Correlation:0.31751
  |
Correlation:0.43049
  |
Correlation:0.43049
  |
Component:125; Jaccard:0.096831
 |
Component:283; Jaccard:0.11257
 |
Component:125; Jaccard:0.13862
 |
Component:125; Jaccard:0.14892
 |
Component:125; Jaccard:0.14687
 |
Component:125; Jaccard:0.13555
 |
Component:125; Jaccard:0.11904
 |
Correlation:0.47469
  |
Correlation:0.19281
  |
Correlation:0.47469
  |
Correlation:0.47469
  |
Correlation:0.47469
  |
Correlation:0.47469
  |
Correlation:0.47469
  |
Component:051; Jaccard:0.094016
 |
Component:096; Jaccard:0.11183
 |
Component:026; Jaccard:0.12712
 |
Component:026; Jaccard:0.1453
 |
Component:358; Jaccard:0.14597
 |
Component:358; Jaccard:0.1295
 |
Component:358; Jaccard:0.10975
 |
Correlation:0.0063101
  |
Correlation:0.38527
  |
Correlation:0.31751
  |
Correlation:0.31751
  |
Correlation:0.31489
  |
Correlation:0.31489
  |
Correlation:0.31489
  |
Component:282; Jaccard:0.091532
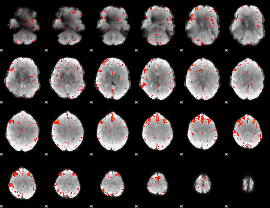 |
Component:051; Jaccard:0.10687
 |
Component:051; Jaccard:0.11286
 |
Component:051; Jaccard:0.11133
 |
Component:136; Jaccard:0.10051
 |
Component:136; Jaccard:0.091633
 |
Component:136; Jaccard:0.079245
 |
Correlation:0.07436
  |
Correlation:0.0063101
  |
Correlation:0.0063101
  |
Correlation:0.0063101
  |
Correlation:0.12143
  |
Correlation:0.12143
  |
Correlation:0.12143
  |
Cope: 04:"neg_FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
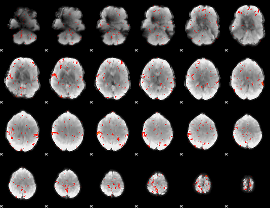 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:143; Jaccard:0.12437
 |
Component:143; Jaccard:0.1398
 |
Component:143; Jaccard:0.11981
 |
Component:143; Jaccard:0.079253
 |
Component:143; Jaccard:0.037959
 |
Component:143; Jaccard:0.015952
 |
Component:143; Jaccard:0.006499
 |
Correlation:0.33038
  |
Correlation:0.33038
  |
Correlation:0.33038
  |
Correlation:0.33038
  |
Correlation:0.33038
  |
Correlation:0.33038
  |
Correlation:0.33038
  |
Component:161; Jaccard:0.090219
 |
Component:161; Jaccard:0.083674
 |
Component:161; Jaccard:0.057118
 |
Component:161; Jaccard:0.0313
 |
Component:161; Jaccard:0.014914
 |
Component:159; Jaccard:0.007294
 |
Component:178; Jaccard:0.004931
 |
Correlation:0.066578
  |
Correlation:0.066578
  |
Correlation:0.066578
  |
Correlation:0.066578
  |
Correlation:0.066578
  |
Correlation:0.020125
  |
Correlation:0.0076814
  |
Component:359; Jaccard:0.085103
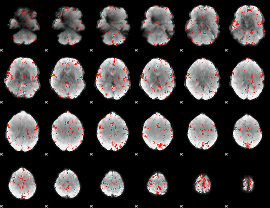 |
Component:359; Jaccard:0.075545
 |
Component:359; Jaccard:0.053119
 |
Component:085; Jaccard:0.030165
 |
Component:159; Jaccard:0.013555
 |
Component:178; Jaccard:0.006814
 |
Component:161; Jaccard:0.003869
 |
Correlation:-0.059591
  |
Correlation:-0.059591
  |
Correlation:-0.059591
  |
Correlation:0.095473
  |
Correlation:0.020125
  |
Correlation:0.0076814
  |
Correlation:0.066578
  |
Component:085; Jaccard:0.072678
 |
Component:085; Jaccard:0.067011
 |
Component:085; Jaccard:0.049378
 |
Component:359; Jaccard:0.028476
 |
Component:085; Jaccard:0.012647
 |
Component:161; Jaccard:0.006763
 |
Component:159; Jaccard:0.003733
 |
Correlation:0.095473
  |
Correlation:0.095473
  |
Correlation:0.095473
  |
Correlation:-0.059591
  |
Correlation:0.095473
  |
Correlation:0.066578
  |
Correlation:0.020125
  |
Component:268; Jaccard:0.072662
 |
Component:268; Jaccard:0.060919
 |
Component:171; Jaccard:0.044809
 |
Component:287; Jaccard:0.026816
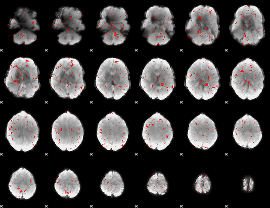 |
Component:287; Jaccard:0.012564
 |
Component:287; Jaccard:0.00667
 |
Component:337; Jaccard:0.003307
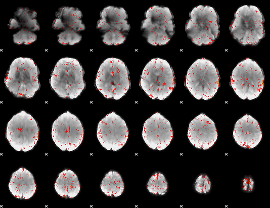 |
Correlation:0.13215
  |
Correlation:0.13215
  |
Correlation:0.07001
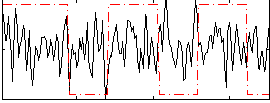 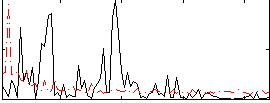 |
Correlation:0.21362
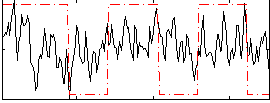 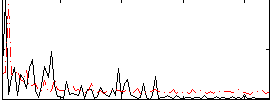 |
Correlation:0.21362
  |
Correlation:0.21362
  |
Correlation:0.2168
  |
Component:001; Jaccard:0.067818
 |
Component:001; Jaccard:0.060739
 |
Component:248; Jaccard:0.044243
 |
Component:248; Jaccard:0.026806
 |
Component:359; Jaccard:0.012531
 |
Component:216; Jaccard:0.00661
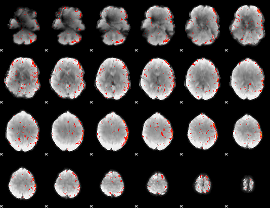 |
Component:287; Jaccard:0.002799
 |
Correlation:0.33284
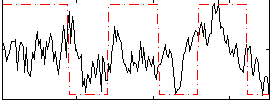  |
Correlation:0.33284
  |
Correlation:0.16378
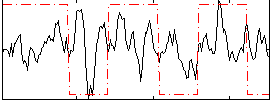  |
Correlation:0.16378
  |
Correlation:-0.059591
  |
Correlation:-0.038393
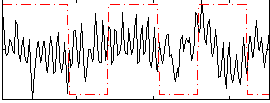  |
Correlation:0.21362
  |
Component:166; Jaccard:0.06644
 |
Component:234; Jaccard:0.060703
 |
Component:287; Jaccard:0.042852
 |
Component:171; Jaccard:0.026665
 |
Component:234; Jaccard:0.012094
 |
Component:337; Jaccard:0.005432
 |
Component:354; Jaccard:0.002503
 |
Correlation:-0.0029996
  |
Correlation:0.10419
  |
Correlation:0.21362
  |
Correlation:0.07001
  |
Correlation:0.10419
  |
Correlation:0.2168
  |
Correlation:0.074864
  |
Component:234; Jaccard:0.065625
 |
Component:296; Jaccard:0.055834
 |
Component:234; Jaccard:0.04236
 |
Component:177; Jaccard:0.025238
 |
Component:216; Jaccard:0.011529
 |
Component:085; Jaccard:0.005253
 |
Component:216; Jaccard:0.002502
 |
Correlation:0.10419
  |
Correlation:0.040199
  |
Correlation:0.10419
  |
Correlation:0.027358
  |
Correlation:-0.038393
  |
Correlation:0.095473
  |
Correlation:-0.038393
  |
Component:168; Jaccard:0.064948
 |
Component:171; Jaccard:0.055589
 |
Component:177; Jaccard:0.04204
 |
Component:234; Jaccard:0.02516
 |
Component:337; Jaccard:0.011324
 |
Component:286; Jaccard:0.005024
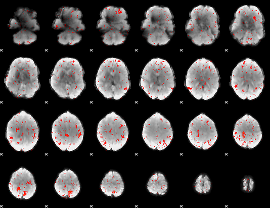 |
Component:268; Jaccard:0.0022
 |
Correlation:0.12049
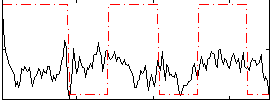  |
Correlation:0.07001
  |
Correlation:0.027358
  |
Correlation:0.10419
  |
Correlation:0.2168
  |
Correlation:0.12414
  |
Correlation:0.13215
  |
Component:337; Jaccard:0.062278
 |
Component:337; Jaccard:0.055441
 |
Component:001; Jaccard:0.041453
 |
Component:214; Jaccard:0.023493
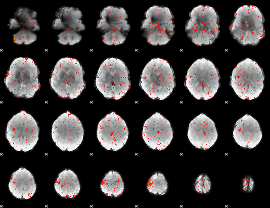 |
Component:178; Jaccard:0.011319
 |
Component:234; Jaccard:0.004615
 |
Component:286; Jaccard:0.002195
 |
Correlation:0.2168
  |
Correlation:0.2168
  |
Correlation:0.33284
  |
Correlation:-0.041429
  |
Correlation:0.0076814
  |
Correlation:0.10419
  |
Correlation:0.12414
  |
Cope: 05:"neg_SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:125; Jaccard:0.123
 |
Component:125; Jaccard:0.14922
 |
Component:125; Jaccard:0.15832
 |
Component:125; Jaccard:0.13499
 |
Component:125; Jaccard:0.087648
 |
Component:125; Jaccard:0.054337
 |
Component:125; Jaccard:0.033372
 |
Correlation:0.47008
  |
Correlation:0.47008
  |
Correlation:0.47008
  |
Correlation:0.47008
  |
Correlation:0.47008
  |
Correlation:0.47008
  |
Correlation:0.47008
  |
Component:161; Jaccard:0.1057
 |
Component:161; Jaccard:0.11088
 |
Component:358; Jaccard:0.11206
 |
Component:143; Jaccard:0.088447
 |
Component:143; Jaccard:0.055474
 |
Component:143; Jaccard:0.030497
 |
Component:358; Jaccard:0.01513
 |
Correlation:0.10353
  |
Correlation:0.10353
  |
Correlation:0.32884
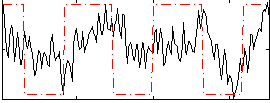  |
Correlation:-0.17092
  |
Correlation:-0.17092
  |
Correlation:-0.17092
  |
Correlation:0.32884
  |
Component:166; Jaccard:0.10491
 |
Component:358; Jaccard:0.1108
 |
Component:143; Jaccard:0.10749
 |
Component:358; Jaccard:0.086785
 |
Component:358; Jaccard:0.052489
 |
Component:358; Jaccard:0.029859
 |
Component:301; Jaccard:0.014642
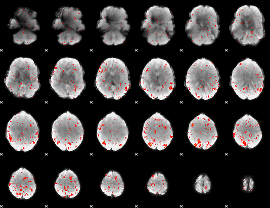 |
Correlation:0.05376
  |
Correlation:0.32884
  |
Correlation:-0.17092
  |
Correlation:0.32884
  |
Correlation:0.32884
  |
Correlation:0.32884
  |
Correlation:0.14333
  |
Component:212; Jaccard:0.097739
 |
Component:166; Jaccard:0.10682
 |
Component:085; Jaccard:0.098785
 |
Component:085; Jaccard:0.074785
 |
Component:360; Jaccard:0.044986
 |
Component:301; Jaccard:0.025983
 |
Component:374; Jaccard:0.013819
 |
Correlation:0.055967
  |
Correlation:0.05376
  |
Correlation:-0.089844
  |
Correlation:-0.089844
  |
Correlation:0.072657
  |
Correlation:0.14333
  |
Correlation:0.45262
  |
Component:358; Jaccard:0.096795
 |
Component:143; Jaccard:0.10606
 |
Component:161; Jaccard:0.096937
 |
Component:360; Jaccard:0.072031
 |
Component:301; Jaccard:0.042579
 |
Component:360; Jaccard:0.024113
 |
Component:360; Jaccard:0.013195
 |
Correlation:0.32884
  |
Correlation:-0.17092
  |
Correlation:0.10353
  |
Correlation:0.072657
  |
Correlation:0.14333
  |
Correlation:0.072657
  |
Correlation:0.072657
  |
Component:143; Jaccard:0.096231
 |
Component:085; Jaccard:0.10158
 |
Component:166; Jaccard:0.093617
 |
Component:161; Jaccard:0.069181
 |
Component:085; Jaccard:0.041649
 |
Component:374; Jaccard:0.023464
 |
Component:143; Jaccard:0.013129
 |
Correlation:-0.17092
  |
Correlation:-0.089844
  |
Correlation:0.05376
  |
Correlation:0.10353
  |
Correlation:-0.089844
  |
Correlation:0.45262
  |
Correlation:-0.17092
  |
Component:085; Jaccard:0.089131
 |
Component:212; Jaccard:0.097138
 |
Component:360; Jaccard:0.091271
 |
Component:301; Jaccard:0.065425
 |
Component:216; Jaccard:0.0402
 |
Component:085; Jaccard:0.022169
 |
Component:272; Jaccard:0.012149
 |
Correlation:-0.089844
  |
Correlation:0.055967
  |
Correlation:0.072657
  |
Correlation:0.14333
  |
Correlation:0.14081
  |
Correlation:-0.089844
  |
Correlation:0.46071
  |
Component:360; Jaccard:0.084919
 |
Component:360; Jaccard:0.093923
 |
Component:301; Jaccard:0.08325
 |
Component:216; Jaccard:0.063781
 |
Component:161; Jaccard:0.038607
 |
Component:161; Jaccard:0.020114
 |
Component:085; Jaccard:0.010927
 |
Correlation:0.072657
  |
Correlation:0.072657
  |
Correlation:0.14333
  |
Correlation:0.14081
  |
Correlation:0.10353
  |
Correlation:0.10353
  |
Correlation:-0.089844
  |
Component:216; Jaccard:0.079165
 |
Component:301; Jaccard:0.085213
 |
Component:212; Jaccard:0.081057
 |
Component:166; Jaccard:0.063283
 |
Component:374; Jaccard:0.038368
 |
Component:216; Jaccard:0.020039
 |
Component:117; Jaccard:0.010776
 |
Correlation:0.14081
  |
Correlation:0.14333
  |
Correlation:0.055967
  |
Correlation:0.05376
  |
Correlation:0.45262
  |
Correlation:0.14081
  |
Correlation:0.063005
  |
Component:359; Jaccard:0.07822
 |
Component:216; Jaccard:0.08426
 |
Component:216; Jaccard:0.080828
 |
Component:374; Jaccard:0.060484
 |
Component:166; Jaccard:0.033095
 |
Component:272; Jaccard:0.018818
 |
Component:321; Jaccard:0.010488
 |
Correlation:0.063219
  |
Correlation:0.14081
  |
Correlation:0.14081
  |
Correlation:0.45262
  |
Correlation:0.05376
  |
Correlation:0.46071
  |
Correlation:0.10293
  |
Cope: 06:"SHAPES-FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:069; Jaccard:0.10321
 |
Component:069; Jaccard:0.10895
 |
Component:069; Jaccard:0.098852
 |
Component:069; Jaccard:0.071312
 |
Component:069; Jaccard:0.038607
 |
Component:069; Jaccard:0.016945
 |
Component:350; Jaccard:0.008493
 |
Correlation:0.49172
  |
Correlation:0.49172
  |
Correlation:0.49172
  |
Correlation:0.49172
  |
Correlation:0.49172
  |
Correlation:0.49172
  |
Correlation:0.34604
  |
Component:353; Jaccard:0.10186
 |
Component:353; Jaccard:0.099925
 |
Component:353; Jaccard:0.079961
 |
Component:174; Jaccard:0.054934
 |
Component:350; Jaccard:0.031399
 |
Component:350; Jaccard:0.016022
 |
Component:069; Jaccard:0.00752
 |
Correlation:0.19895
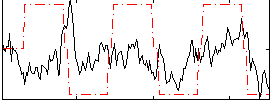  |
Correlation:0.19895
  |
Correlation:0.19895
  |
Correlation:0.18403
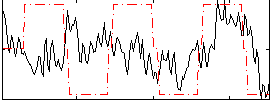  |
Correlation:0.34604
  |
Correlation:0.34604
  |
Correlation:0.49172
  |
Component:001; Jaccard:0.087589
 |
Component:174; Jaccard:0.088231
 |
Component:174; Jaccard:0.077702
 |
Component:353; Jaccard:0.049539
 |
Component:174; Jaccard:0.029682
 |
Component:174; Jaccard:0.015331
 |
Component:174; Jaccard:0.006483
 |
Correlation:0.32095
  |
Correlation:0.18403
  |
Correlation:0.18403
  |
Correlation:0.19895
  |
Correlation:0.18403
  |
Correlation:0.18403
  |
Correlation:0.18403
  |
Component:174; Jaccard:0.084527
 |
Component:001; Jaccard:0.086061
 |
Component:252; Jaccard:0.071474
 |
Component:252; Jaccard:0.047485
 |
Component:353; Jaccard:0.023981
 |
Component:353; Jaccard:0.0108
 |
Component:353; Jaccard:0.004602
 |
Correlation:0.18403
  |
Correlation:0.32095
  |
Correlation:0.25718
  |
Correlation:0.25718
  |
Correlation:0.19895
  |
Correlation:0.19895
  |
Correlation:0.19895
  |
Component:252; Jaccard:0.083189
 |
Component:252; Jaccard:0.0846
 |
Component:001; Jaccard:0.067474
 |
Component:350; Jaccard:0.045179
 |
Component:252; Jaccard:0.022522
 |
Component:252; Jaccard:0.009965
 |
Component:252; Jaccard:0.003793
 |
Correlation:0.25718
  |
Correlation:0.25718
  |
Correlation:0.32095
  |
Correlation:0.34604
  |
Correlation:0.25718
  |
Correlation:0.25718
  |
Correlation:0.25718
  |
Component:198; Jaccard:0.079695
 |
Component:198; Jaccard:0.079068
 |
Component:198; Jaccard:0.065585
 |
Component:001; Jaccard:0.040011
 |
Component:302; Jaccard:0.021419
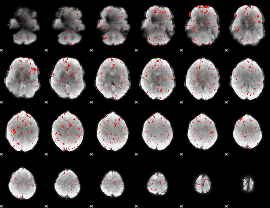 |
Component:001; Jaccard:0.009576
 |
Component:001; Jaccard:0.003447
 |
Correlation:0.24331
  |
Correlation:0.24331
  |
Correlation:0.24331
  |
Correlation:0.32095
  |
Correlation:0.3447
  |
Correlation:0.32095
  |
Correlation:0.32095
  |
Component:355; Jaccard:0.07421
 |
Component:355; Jaccard:0.073248
 |
Component:355; Jaccard:0.058437
 |
Component:198; Jaccard:0.039809
 |
Component:001; Jaccard:0.020272
 |
Component:302; Jaccard:0.008479
 |
Component:121; Jaccard:0.003344
 |
Correlation:0.18356
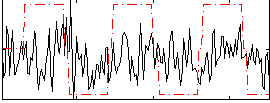  |
Correlation:0.18356
  |
Correlation:0.18356
  |
Correlation:0.24331
  |
Correlation:0.32095
  |
Correlation:0.3447
  |
Correlation:0.13363
  |
Component:182; Jaccard:0.069639
 |
Component:302; Jaccard:0.067036
 |
Component:302; Jaccard:0.055135
 |
Component:355; Jaccard:0.038631
 |
Component:198; Jaccard:0.020178
 |
Component:198; Jaccard:0.008475
 |
Component:198; Jaccard:0.003278
 |
Correlation:0.16986
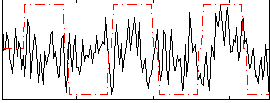  |
Correlation:0.3447
  |
Correlation:0.3447
  |
Correlation:0.18356
  |
Correlation:0.24331
  |
Correlation:0.24331
  |
Correlation:0.24331
  |
Component:121; Jaccard:0.067796
 |
Component:182; Jaccard:0.064817
 |
Component:350; Jaccard:0.05382
 |
Component:302; Jaccard:0.037417
 |
Component:355; Jaccard:0.019603
 |
Component:121; Jaccard:0.008322
 |
Component:188; Jaccard:0.003218
 |
Correlation:0.13363
  |
Correlation:0.16986
  |
Correlation:0.34604
  |
Correlation:0.3447
  |
Correlation:0.18356
  |
Correlation:0.13363
  |
Correlation:0.31697
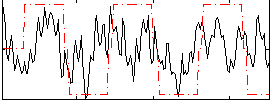  |
Component:017; Jaccard:0.067708
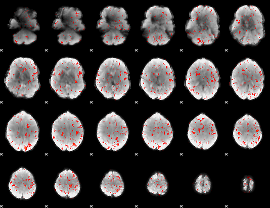 |
Component:121; Jaccard:0.06286
 |
Component:121; Jaccard:0.051958
 |
Component:121; Jaccard:0.035028
 |
Component:121; Jaccard:0.016968
 |
Component:355; Jaccard:0.008176
 |
Component:265; Jaccard:0.003113
 |
Correlation:0.16671
  |
Correlation:0.13363
  |
Correlation:0.13363
  |
Correlation:0.13363
  |
Correlation:0.13363
  |
Correlation:0.18356
  |
Correlation:0.23972
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































