Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
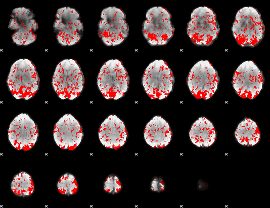 |
GLM binary result thresholded by 1.5
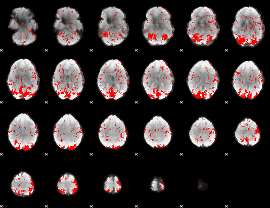 |
GLM binary result thresholded by 2
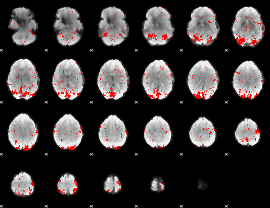 |
GLM binary result thresholded by 2.5
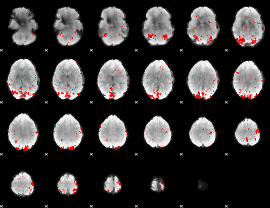 |
GLM binary result thresholded by 3
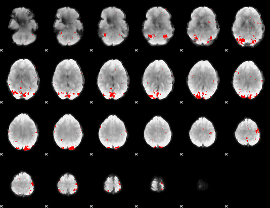 |
GLM binary result thresholded by 3.5
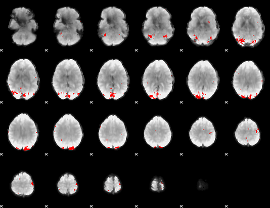 |
GLM binary result thresholded by 4
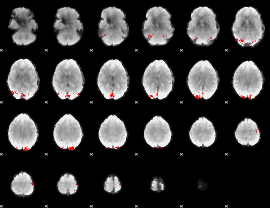 |
GLM Z-score result thresholded by 1
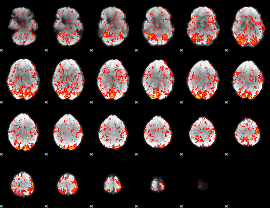 |
GLM Z-score result thresholded by 1.5
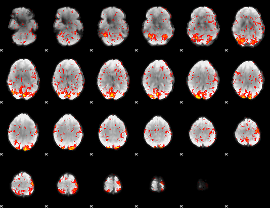 |
GLM Z-score result thresholded by 2
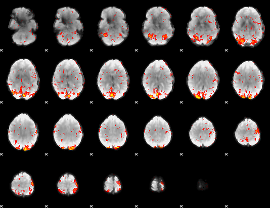 |
GLM Z-score result thresholded by 2.5
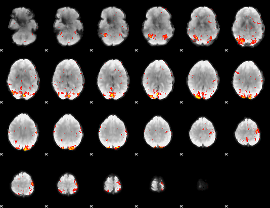 |
GLM Z-score result thresholded by 3
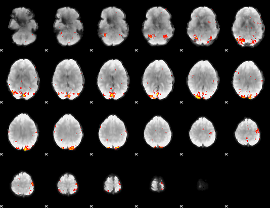 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:257; Jaccard:0.15531
 |
Component:257; Jaccard:0.20354
 |
Component:257; Jaccard:0.23881
 |
Component:257; Jaccard:0.24252
 |
Component:257; Jaccard:0.21971
 |
Component:257; Jaccard:0.17863
 |
Component:257; Jaccard:0.1402
 |
Correlation:0.24905
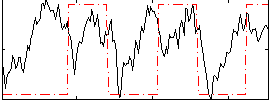 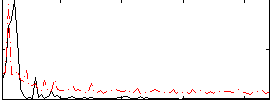 |
Correlation:0.24905
  |
Correlation:0.24905
  |
Correlation:0.24905
  |
Correlation:0.24905
  |
Correlation:0.24905
  |
Correlation:0.24905
  |
Component:282; Jaccard:0.15094
 |
Component:282; Jaccard:0.19887
 |
Component:282; Jaccard:0.23407
 |
Component:282; Jaccard:0.23671
 |
Component:282; Jaccard:0.21435
 |
Component:282; Jaccard:0.16785
 |
Component:282; Jaccard:0.13012
 |
Correlation:-0.2021
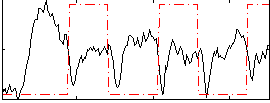 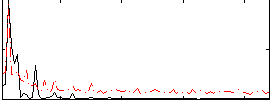 |
Correlation:-0.2021
  |
Correlation:-0.2021
  |
Correlation:-0.2021
  |
Correlation:-0.2021
  |
Correlation:-0.2021
  |
Correlation:-0.2021
  |
Component:084; Jaccard:0.12812
 |
Component:084; Jaccard:0.15468
 |
Component:084; Jaccard:0.17281
 |
Component:084; Jaccard:0.17376
 |
Component:084; Jaccard:0.15384
 |
Component:084; Jaccard:0.12342
 |
Component:084; Jaccard:0.091194
 |
Correlation:0.61921
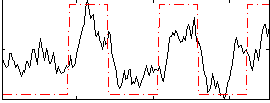 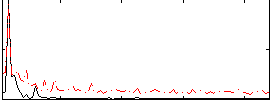 |
Correlation:0.61921
  |
Correlation:0.61921
  |
Correlation:0.61921
  |
Correlation:0.61921
  |
Correlation:0.61921
  |
Correlation:0.61921
  |
Component:389; Jaccard:0.12363
 |
Component:359; Jaccard:0.1348
 |
Component:359; Jaccard:0.14271
 |
Component:359; Jaccard:0.13559
 |
Component:359; Jaccard:0.11603
 |
Component:359; Jaccard:0.085741
 |
Component:359; Jaccard:0.059096
 |
Correlation:0.0048902
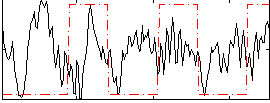 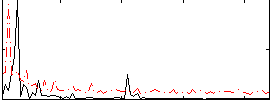 |
Correlation:0.10009
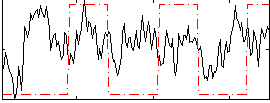 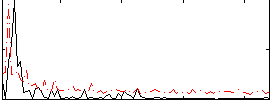 |
Correlation:0.10009
  |
Correlation:0.10009
  |
Correlation:0.10009
  |
Correlation:0.10009
  |
Correlation:0.10009
  |
Component:359; Jaccard:0.11354
 |
Component:389; Jaccard:0.12784
 |
Component:028; Jaccard:0.1292
 |
Component:028; Jaccard:0.11756
 |
Component:028; Jaccard:0.095092
 |
Component:028; Jaccard:0.073038
 |
Component:028; Jaccard:0.052106
 |
Correlation:0.10009
  |
Correlation:0.0048902
  |
Correlation:0.1469
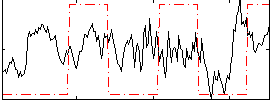 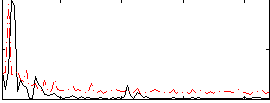 |
Correlation:0.1469
  |
Correlation:0.1469
  |
Correlation:0.1469
  |
Correlation:0.1469
  |
Component:028; Jaccard:0.11071
 |
Component:028; Jaccard:0.12529
 |
Component:389; Jaccard:0.12234
 |
Component:389; Jaccard:0.10076
 |
Component:389; Jaccard:0.07476
 |
Component:389; Jaccard:0.048374
 |
Component:238; Jaccard:0.032958
 |
Correlation:0.1469
  |
Correlation:0.1469
  |
Correlation:0.0048902
  |
Correlation:0.0048902
  |
Correlation:0.0048902
  |
Correlation:0.0048902
  |
Correlation:0.032635
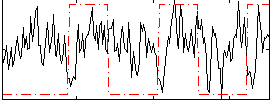 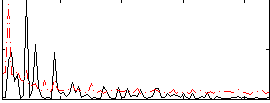 |
Component:100; Jaccard:0.08895
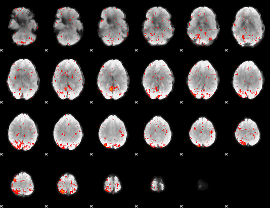 |
Component:208; Jaccard:0.093484
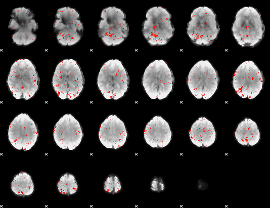 |
Component:208; Jaccard:0.095248
 |
Component:208; Jaccard:0.083551
 |
Component:265; Jaccard:0.063117
 |
Component:265; Jaccard:0.044509
 |
Component:265; Jaccard:0.031946
 |
Correlation:-0.092421
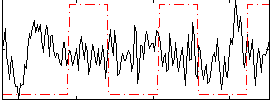 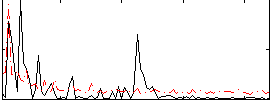 |
Correlation:0.046919
  |
Correlation:0.046919
  |
Correlation:0.046919
  |
Correlation:-0.21989
  |
Correlation:-0.21989
  |
Correlation:-0.21989
  |
Component:265; Jaccard:0.085098
 |
Component:100; Jaccard:0.093426
 |
Component:265; Jaccard:0.090915
 |
Component:265; Jaccard:0.079569
 |
Component:208; Jaccard:0.059121
 |
Component:238; Jaccard:0.043071
 |
Component:389; Jaccard:0.029891
 |
Correlation:-0.21989
  |
Correlation:-0.092421
  |
Correlation:-0.21989
  |
Correlation:-0.21989
  |
Correlation:0.046919
  |
Correlation:0.032635
  |
Correlation:0.0048902
  |
Component:202; Jaccard:0.081295
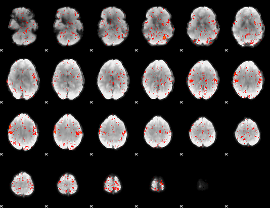 |
Component:265; Jaccard:0.091226
 |
Component:100; Jaccard:0.084839
 |
Component:238; Jaccard:0.06767
 |
Component:238; Jaccard:0.054946
 |
Component:208; Jaccard:0.040819
 |
Component:034; Jaccard:0.028577
 |
Correlation:-0.00067838
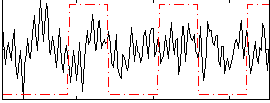 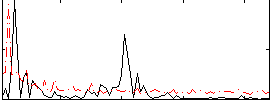 |
Correlation:-0.21989
  |
Correlation:-0.092421
  |
Correlation:0.032635
  |
Correlation:0.032635
  |
Correlation:0.046919
  |
Correlation:-0.34603
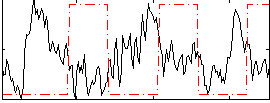 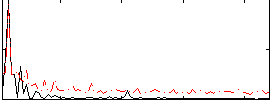 |
Component:208; Jaccard:0.081078
 |
Component:238; Jaccard:0.081611
 |
Component:034; Jaccard:0.077058
 |
Component:100; Jaccard:0.067017
 |
Component:034; Jaccard:0.051536
 |
Component:034; Jaccard:0.038388
 |
Component:145; Jaccard:0.028458
 |
Correlation:0.046919
  |
Correlation:0.032635
  |
Correlation:-0.34603
  |
Correlation:-0.092421
  |
Correlation:-0.34603
  |
Correlation:-0.34603
  |
Correlation:0.60398
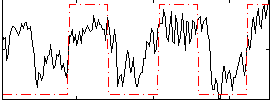 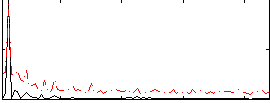 |
Cope: 02:"SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
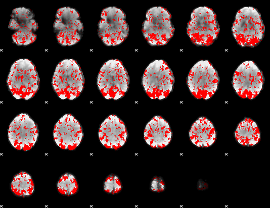 |
GLM binary result thresholded by 1.5
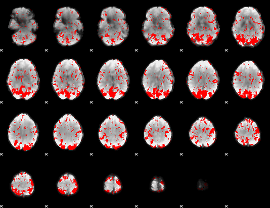 |
GLM binary result thresholded by 2
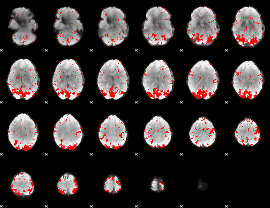 |
GLM binary result thresholded by 2.5
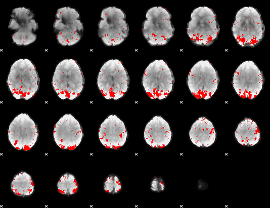 |
GLM binary result thresholded by 3
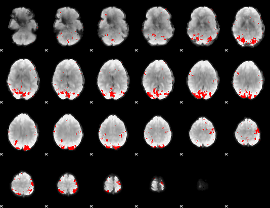 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
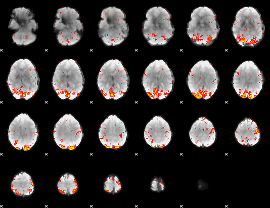 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
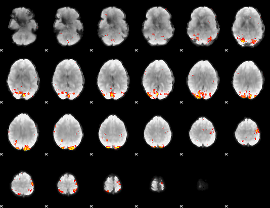 |
GLM Z-score result thresholded by 4
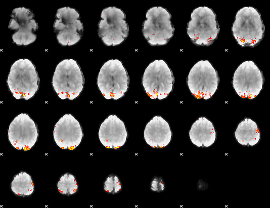 |
Component:282; Jaccard:0.14906
 |
Component:282; Jaccard:0.21265
 |
Component:282; Jaccard:0.28409
 |
Component:282; Jaccard:0.33815
 |
Component:282; Jaccard:0.3538
 |
Component:282; Jaccard:0.33497
 |
Component:282; Jaccard:0.28742
 |
Correlation:0.41062
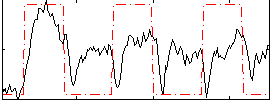  |
Correlation:0.41062
  |
Correlation:0.41062
  |
Correlation:0.41062
  |
Correlation:0.41062
  |
Correlation:0.41062
  |
Correlation:0.41062
  |
Component:257; Jaccard:0.1326
 |
Component:257; Jaccard:0.17469
 |
Component:257; Jaccard:0.21297
 |
Component:257; Jaccard:0.2342
 |
Component:257; Jaccard:0.23024
 |
Component:257; Jaccard:0.20562
 |
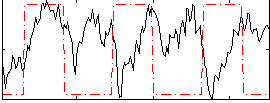
Component:257; Jaccard:0.1776
 |
Correlation:-0.1541
 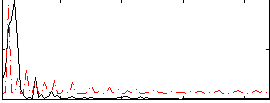 |
Correlation:-0.1541
  |
Correlation:-0.1541
  |
Correlation:-0.1541
  |
Correlation:-0.1541
  |
Correlation:-0.1541
  |
Correlation:-0.1541
  |
Component:389; Jaccard:0.13006
 |
Component:034; Jaccard:0.14911
 |
Component:359; Jaccard:0.1688
 |
Component:359; Jaccard:0.17663
 |
Component:359; Jaccard:0.16883
 |
Component:359; Jaccard:0.14537
 |
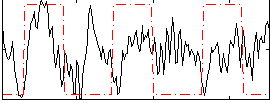
Component:359; Jaccard:0.11758
 |
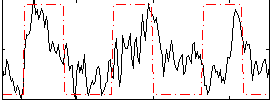
Correlation:0.132
  |
Correlation:0.43276
  |
Correlation:0.14769
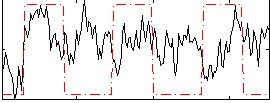 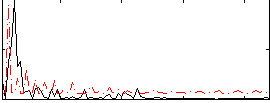 |
Correlation:0.14769
  |
Correlation:0.14769
  |
Correlation:0.14769
  |
Correlation:0.14769
  |
Component:034; Jaccard:0.12602
 |
Component:359; Jaccard:0.14504
 |
Component:034; Jaccard:0.16652
 |
Component:034; Jaccard:0.16739
 |
Component:034; Jaccard:0.151
 |
Component:034; Jaccard:0.12354
 |
Component:034; Jaccard:0.096316
 |
Correlation:0.43276
  |
Correlation:0.14769
  |
Correlation:0.43276
  |
Correlation:0.43276
  |
Correlation:0.43276
  |
Correlation:0.43276
  |
Correlation:0.43276
  |
Component:265; Jaccard:0.12113
 |
Component:389; Jaccard:0.14499
 |
Component:265; Jaccard:0.15646
 |
Component:265; Jaccard:0.15849
 |
Component:265; Jaccard:0.14532
 |
Component:265; Jaccard:0.1167
 |
Component:265; Jaccard:0.092154
 |
Correlation:0.23682
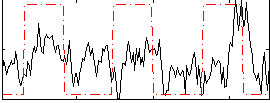  |
Correlation:0.132
  |
Correlation:0.23682
  |
Correlation:0.23682
  |
Correlation:0.23682
  |
Correlation:0.23682
  |
Correlation:0.23682
  |
Component:368; Jaccard:0.11844
 |
Component:265; Jaccard:0.14288
 |
Component:368; Jaccard:0.15242
 |
Component:368; Jaccard:0.14478
 |
Component:028; Jaccard:0.1313
 |
Component:028; Jaccard:0.10966
 |
Component:028; Jaccard:0.086665
 |
Correlation:0.27609
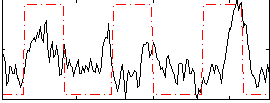  |
Correlation:0.23682
  |
Correlation:0.27609
  |
Correlation:0.27609
  |
Correlation:-0.037643
  |
Correlation:-0.037643
  |
Correlation:-0.037643
  |
Component:359; Jaccard:0.11573
 |
Component:368; Jaccard:0.1412
 |
Component:389; Jaccard:0.14639
 |
Component:028; Jaccard:0.1418
 |
Component:368; Jaccard:0.1241
 |
Component:368; Jaccard:0.097475
 |
Component:011; Jaccard:0.076235
 |
Correlation:0.14769
  |
Correlation:0.27609
  |
Correlation:0.132
  |
Correlation:-0.037643
  |
Correlation:0.27609
  |
Correlation:0.27609
  |
Correlation:0.21216
  |
Component:183; Jaccard:0.11237
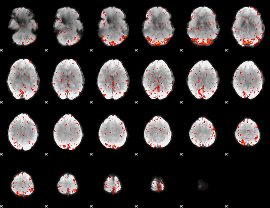 |
Component:028; Jaccard:0.12581
 |
Component:028; Jaccard:0.14003
 |
Component:389; Jaccard:0.13396
 |
Component:011; Jaccard:0.11571
 |
Component:011; Jaccard:0.095051
 |
Component:368; Jaccard:0.071568
 |
Correlation:0.051818
  |
Correlation:-0.037643
  |
Correlation:-0.037643
  |
Correlation:0.132
  |
Correlation:0.21216
  |
Correlation:0.21216
  |
Correlation:0.27609
  |
Component:028; Jaccard:0.10721
 |
Component:011; Jaccard:0.11864
 |
Component:011; Jaccard:0.12906
 |
Component:011; Jaccard:0.12526
 |
Component:389; Jaccard:0.11098
 |
Component:120; Jaccard:0.08866
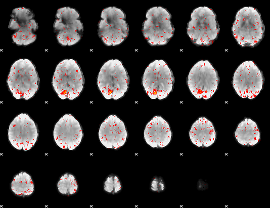 |
Component:120; Jaccard:0.066623
 |
Correlation:-0.037643
  |
Correlation:0.21216
  |
Correlation:0.21216
  |
Correlation:0.21216
  |
Correlation:0.132
  |
Correlation:0.22261
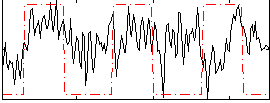 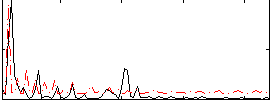 |
Correlation:0.22261
  |
Component:100; Jaccard:0.10412
 |
Component:100; Jaccard:0.11807
 |
Component:100; Jaccard:0.12534
 |
Component:120; Jaccard:0.11695
 |
Component:120; Jaccard:0.10508
 |
Component:389; Jaccard:0.087964
 |
Component:389; Jaccard:0.065516
 |
Correlation:0.2042
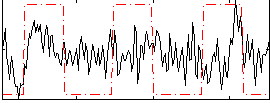  |
Correlation:0.2042
  |
Correlation:0.2042
  |
Correlation:0.22261
  |
Correlation:0.22261
  |
Correlation:0.132
  |
Correlation:0.132
  |
Cope: 03:"FACES-SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
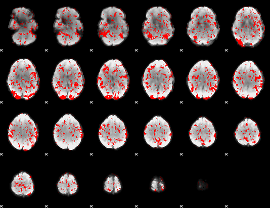 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
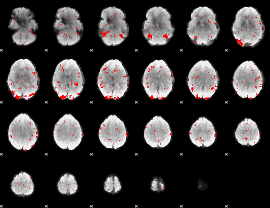 |
GLM binary result thresholded by 2.5
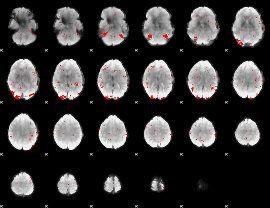 |
GLM binary result thresholded by 3
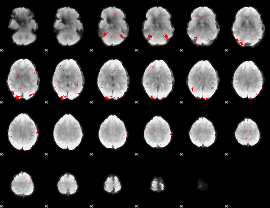 |
GLM binary result thresholded by 3.5
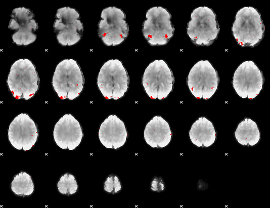 |
GLM binary result thresholded by 4
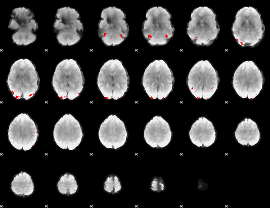 |
GLM Z-score result thresholded by 1
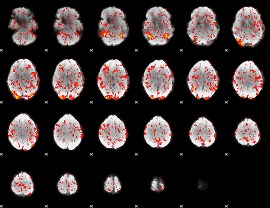 |
GLM Z-score result thresholded by 1.5
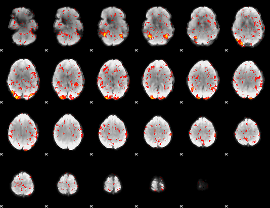 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
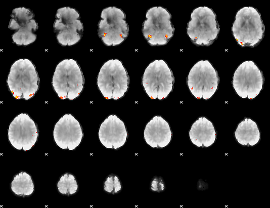 |
Component:084; Jaccard:0.17588
 |
Component:084; Jaccard:0.21754
 |
Component:084; Jaccard:0.23165
 |
Component:084; Jaccard:0.20411
 |
Component:084; Jaccard:0.15768
 |
Component:084; Jaccard:0.11127
 |
Component:084; Jaccard:0.082452
 |
Correlation:0.63957
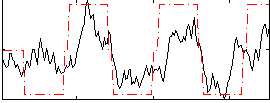 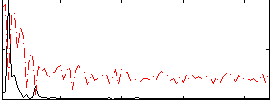 |
Correlation:0.63957
  |
Correlation:0.63957
  |
Correlation:0.63957
  |
Correlation:0.63957
  |
Correlation:0.63957
  |
Correlation:0.63957
  |
Component:145; Jaccard:0.12398
 |
Component:145; Jaccard:0.14755
 |
Component:145; Jaccard:0.15717
 |
Component:145; Jaccard:0.15006
 |
Component:145; Jaccard:0.12283
 |
Component:145; Jaccard:0.095974
 |
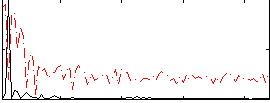
Component:145; Jaccard:0.07445
 |
Correlation:0.68087
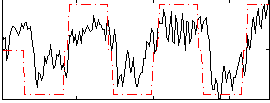  |
Correlation:0.68087
  |
Correlation:0.68087
  |
Correlation:0.68087
  |
Correlation:0.68087
  |
Correlation:0.68087
  |
Correlation:0.68087
  |
Component:010; Jaccard:0.10034
 |
Component:010; Jaccard:0.10865
 |
Component:010; Jaccard:0.10607
 |
Component:010; Jaccard:0.092654
 |
Component:010; Jaccard:0.070653
 |
Component:010; Jaccard:0.051473
 |
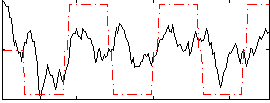
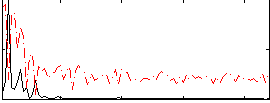
Component:010; Jaccard:0.0408
 |
Correlation:0.3928
  |
Correlation:0.3928
  |
Correlation:0.3928
  |
Correlation:0.3928
  |
Correlation:0.3928
  |
Correlation:0.3928
  |
Correlation:0.3928
  |
Component:238; Jaccard:0.07819
 |
Component:238; Jaccard:0.07423
 |
Component:238; Jaccard:0.061682
 |
Component:238; Jaccard:0.046798
 |
Component:238; Jaccard:0.032972
 |
Component:227; Jaccard:0.023063
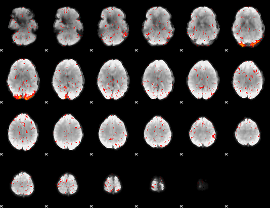 |
Component:227; Jaccard:0.017919
 |
Correlation:0.023375
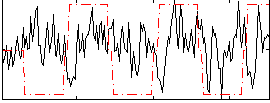 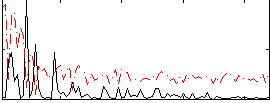 |
Correlation:0.023375
  |
Correlation:0.023375
  |
Correlation:0.023375
  |
Correlation:0.023375
  |
Correlation:0.042768
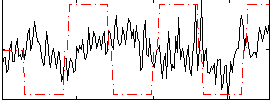 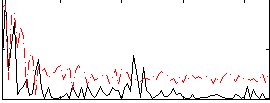 |
Correlation:0.042768
  |
Component:312; Jaccard:0.069582
 |
Component:162; Jaccard:0.066686
 |
Component:162; Jaccard:0.057411
 |
Component:122; Jaccard:0.043941
 |
Component:257; Jaccard:0.03175
 |
Component:238; Jaccard:0.022848
 |
Component:238; Jaccard:0.016531
 |
Correlation:0.26252
  |
Correlation:-0.07456
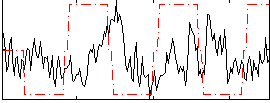 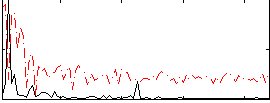 |
Correlation:-0.07456
  |
Correlation:0.29466
 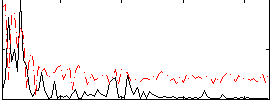 |
Correlation:0.21867
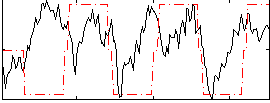  |
Correlation:0.023375
  |
Correlation:0.023375
  |
Component:162; Jaccard:0.06917
 |
Component:314; Jaccard:0.06482
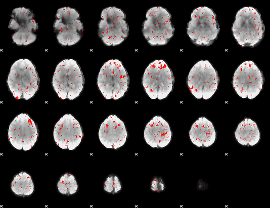 |
Component:122; Jaccard:0.05645
 |
Component:257; Jaccard:0.040925
 |
Component:227; Jaccard:0.03076
 |
Component:257; Jaccard:0.022802
 |
Component:122; Jaccard:0.014501
 |
Correlation:-0.07456
  |
Correlation:0.1378
  |
Correlation:0.29466
  |
Correlation:0.21867
  |
Correlation:0.042768
  |
Correlation:0.21867
  |
Correlation:0.29466
  |
Component:314; Jaccard:0.067746
 |
Component:312; Jaccard:0.064548
 |
Component:314; Jaccard:0.053698
 |
Component:162; Jaccard:0.039876
 |
Component:122; Jaccard:0.030444
 |
Component:122; Jaccard:0.020733
 |
Component:153; Jaccard:0.014137
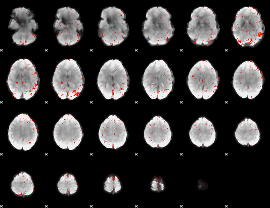 |
Correlation:0.1378
  |
Correlation:0.26252
  |
Correlation:0.1378
  |
Correlation:-0.07456
  |
Correlation:0.29466
  |
Correlation:0.29466
  |
Correlation:0.22761
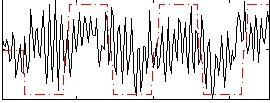 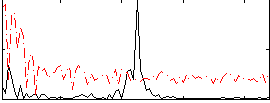 |
Component:122; Jaccard:0.064873
 |
Component:122; Jaccard:0.063376
 |
Component:312; Jaccard:0.049988
 |
Component:314; Jaccard:0.039829
 |
Component:314; Jaccard:0.025338
 |
Component:153; Jaccard:0.017541
 |
Component:257; Jaccard:0.014068
 |
Correlation:0.29466
  |
Correlation:0.29466
  |
Correlation:0.26252
  |
Correlation:0.1378
  |
Correlation:0.1378
  |
Correlation:0.22761
  |
Correlation:0.21867
  |
Component:185; Jaccard:0.061378
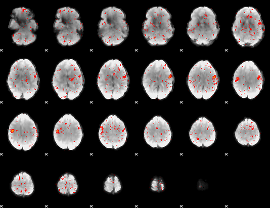 |
Component:185; Jaccard:0.057864
 |
Component:318; Jaccard:0.046383
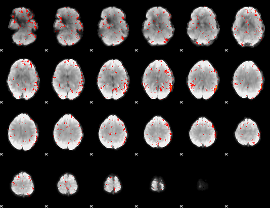 |
Component:227; Jaccard:0.037859
 |
Component:153; Jaccard:0.024523
 |
Component:314; Jaccard:0.016637
 |
Component:314; Jaccard:0.012559
 |
Correlation:-0.0087455
 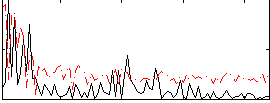 |
Correlation:-0.0087455
  |
Correlation:0.15197
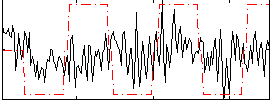  |
Correlation:0.042768
  |
Correlation:0.22761
  |
Correlation:0.1378
  |
Correlation:0.1378
  |
Component:318; Jaccard:0.060482
 |
Component:318; Jaccard:0.057597
 |
Component:287; Jaccard:0.045332
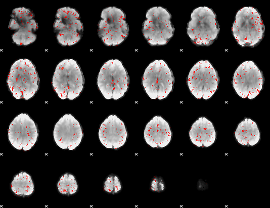 |
Component:318; Jaccard:0.034381
 |
Component:162; Jaccard:0.024245
 |
Component:318; Jaccard:0.013796
 |
Component:324; Jaccard:0.01015
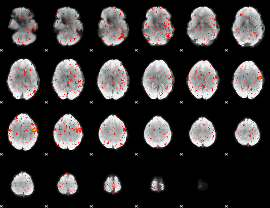 |
Correlation:0.15197
  |
Correlation:0.15197
  |
Correlation:0.1832
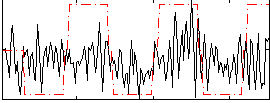 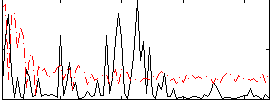 |
Correlation:0.15197
  |
Correlation:-0.07456
  |
Correlation:0.15197
  |
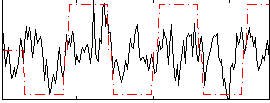
Correlation:0.29564
  |
Cope: 04:"neg_FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
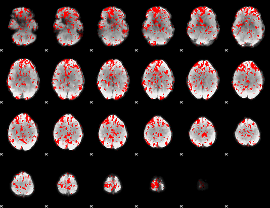 |
GLM binary result thresholded by 2
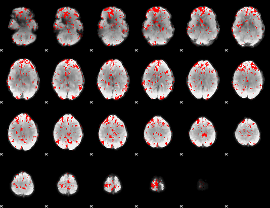 |
GLM binary result thresholded by 2.5
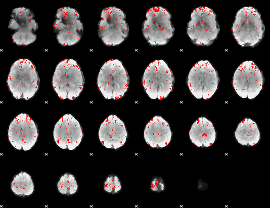 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
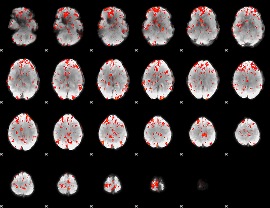 |
GLM Z-score result thresholded by 2.5
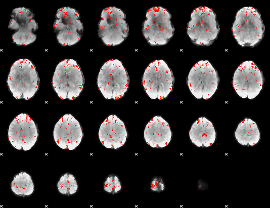 |
GLM Z-score result thresholded by 3
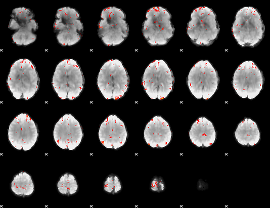 |
GLM Z-score result thresholded by 3.5
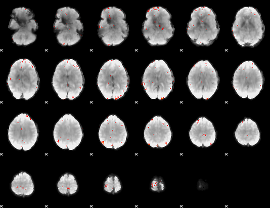 |
GLM Z-score result thresholded by 4
 |
Component:259; Jaccard:0.13259
 |
Component:259; Jaccard:0.1686
 |
Component:259; Jaccard:0.19703
 |
Component:259; Jaccard:0.18452
 |
Component:259; Jaccard:0.12699
 |
Component:259; Jaccard:0.067708
 |
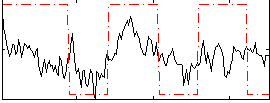
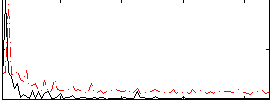
Component:259; Jaccard:0.026059
 |
Correlation:0.41164
  |
Correlation:0.41164
  |
Correlation:0.41164
  |
Correlation:0.41164
  |
Correlation:0.41164
  |
Correlation:0.41164
  |
Correlation:0.41164
  |
Component:319; Jaccard:0.095382
 |
Component:319; Jaccard:0.10489
 |
Component:230; Jaccard:0.10895
 |
Component:306; Jaccard:0.10094
 |
Component:306; Jaccard:0.076802
 |
Component:306; Jaccard:0.046278
 |
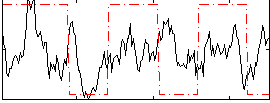
Component:306; Jaccard:0.021767
 |
Correlation:0.15764
  |
Correlation:0.15764
  |
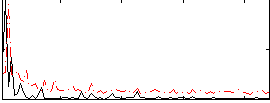
Correlation:0.24697
  |
Correlation:0.089813
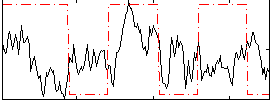 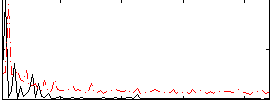 |
Correlation:0.089813
  |
Correlation:0.089813
  |
Correlation:0.089813
  |
Component:230; Jaccard:0.089872
 |
Component:230; Jaccard:0.10339
 |
Component:306; Jaccard:0.10654
 |
Component:230; Jaccard:0.097427
 |
Component:214; Jaccard:0.067418
 |
Component:121; Jaccard:0.043329
 |
Component:121; Jaccard:0.019386
 |
Correlation:0.24697
  |
Correlation:0.24697
  |
Correlation:0.089813
  |
Correlation:0.24697
  |
Correlation:0.10845
 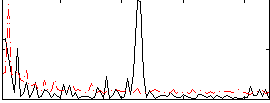 |
Correlation:-0.033268
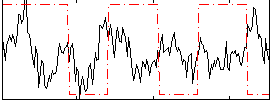  |
Correlation:-0.033268
  |
Component:306; Jaccard:0.086583
 |
Component:306; Jaccard:0.10263
 |
Component:319; Jaccard:0.10309
 |
Component:214; Jaccard:0.089526
 |
Component:121; Jaccard:0.067306
 |
Component:214; Jaccard:0.037818
 |
Component:010; Jaccard:0.0166
 |
Correlation:0.089813
  |
Correlation:0.089813
  |
Correlation:0.15764
  |
Correlation:0.10845
  |
Correlation:-0.033268
  |
Correlation:0.10845
  |
Correlation:-0.2959
  |
Component:223; Jaccard:0.080607
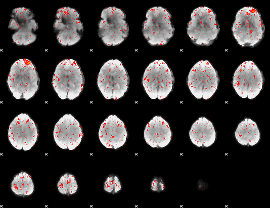 |
Component:214; Jaccard:0.087147
 |
Component:214; Jaccard:0.096909
 |
Component:121; Jaccard:0.087569
 |
Component:230; Jaccard:0.066013
 |
Component:010; Jaccard:0.035221
 |
Component:214; Jaccard:0.013366
 |
Correlation:0.055409
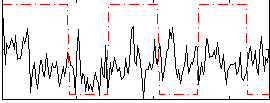 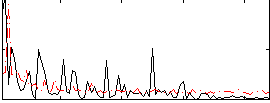 |
Correlation:0.10845
  |
Correlation:0.10845
  |
Correlation:-0.033268
  |
Correlation:0.24697
  |
Correlation:-0.2959
  |
Correlation:0.10845
  |
Component:264; Jaccard:0.0793
 |
Component:223; Jaccard:0.086665
 |
Component:121; Jaccard:0.088336
 |
Component:319; Jaccard:0.085865
 |
Component:010; Jaccard:0.060974
 |
Component:230; Jaccard:0.033547
 |
Component:230; Jaccard:0.012021
 |
Correlation:0.24121
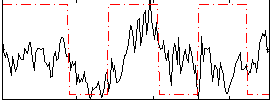 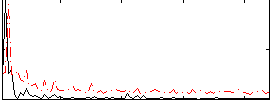 |
Correlation:0.055409
  |
Correlation:-0.033268
  |
Correlation:0.15764
  |
Correlation:-0.2959
  |
Correlation:0.24697
  |
Correlation:0.24697
  |
Component:332; Jaccard:0.077592
 |
Component:332; Jaccard:0.086553
 |
Component:010; Jaccard:0.087332
 |
Component:010; Jaccard:0.082073
 |
Component:319; Jaccard:0.054082
 |
Component:216; Jaccard:0.02929
 |
Component:332; Jaccard:0.011477
 |
Correlation:0.27022
  |
Correlation:0.27022
  |
Correlation:-0.2959
  |
Correlation:-0.2959
  |
Correlation:0.15764
  |
Correlation:0.19446
  |
Correlation:0.27022
  |
Component:119; Jaccard:0.077069
 |
Component:010; Jaccard:0.085313
 |
Component:332; Jaccard:0.08638
 |
Component:393; Jaccard:0.075427
 |
Component:393; Jaccard:0.053871
 |
Component:332; Jaccard:0.028088
 |
Component:252; Jaccard:0.011338
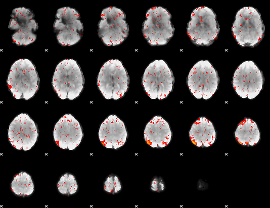 |
Correlation:-0.010849
  |
Correlation:-0.2959
  |
Correlation:0.27022
  |
Correlation:0.041636
  |
Correlation:0.041636
  |
Correlation:0.27022
  |
Correlation:0.2557
  |
Component:010; Jaccard:0.075929
 |
Component:264; Jaccard:0.082887
 |
Component:223; Jaccard:0.084573
 |
Component:216; Jaccard:0.073706
 |
Component:216; Jaccard:0.051279
 |
Component:393; Jaccard:0.027298
 |
Component:393; Jaccard:0.010982
 |
Correlation:-0.2959
  |
Correlation:0.24121
  |
Correlation:0.055409
  |
Correlation:0.19446
  |
Correlation:0.19446
  |
Correlation:0.041636
  |
Correlation:0.041636
  |
Component:214; Jaccard:0.075481
 |
Component:121; Jaccard:0.082258
 |
Component:393; Jaccard:0.08431
 |
Component:332; Jaccard:0.073211
 |
Component:332; Jaccard:0.049783
 |
Component:319; Jaccard:0.026169
 |
Component:216; Jaccard:0.010269
 |
Correlation:0.10845
  |
Correlation:-0.033268
  |
Correlation:0.041636
  |
Correlation:0.27022
  |
Correlation:0.27022
  |
Correlation:0.15764
  |
Correlation:0.19446
  |
Cope: 05:"neg_SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
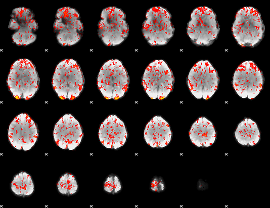 |
GLM Z-score result thresholded by 2
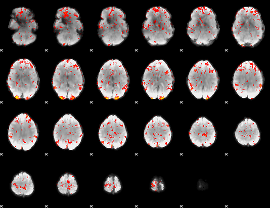 |
GLM Z-score result thresholded by 2.5
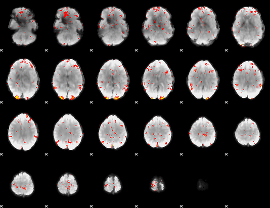 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
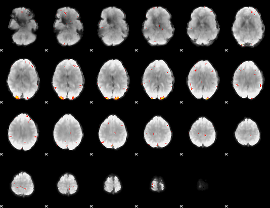 |
GLM Z-score result thresholded by 4
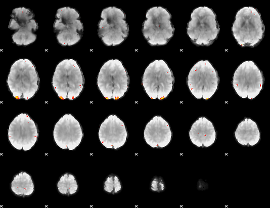 |
Component:010; Jaccard:0.12718
 |
Component:010; Jaccard:0.16219
 |
Component:010; Jaccard:0.19032
 |
Component:010; Jaccard:0.18197
 |
Component:010; Jaccard:0.14902
 |
Component:010; Jaccard:0.11169
 |
Component:010; Jaccard:0.07889
 |
Correlation:0.42827
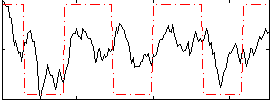 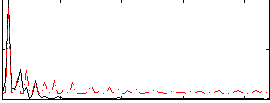 |
Correlation:0.42827
  |
Correlation:0.42827
  |
Correlation:0.42827
  |
Correlation:0.42827
  |
Correlation:0.42827
  |
Correlation:0.42827
  |
Component:259; Jaccard:0.10132
 |
Component:259; Jaccard:0.1106
 |
Component:121; Jaccard:0.11811
 |
Component:121; Jaccard:0.11013
 |
Component:121; Jaccard:0.078914
 |
Component:145; Jaccard:0.052616
 |
Component:145; Jaccard:0.043144
 |
Correlation:-0.26978
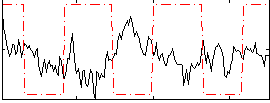  |
Correlation:-0.26978
  |
Correlation:0.24958
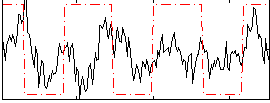 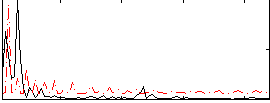 |
Correlation:0.24958
  |
Correlation:0.24958
  |
Correlation:0.65128
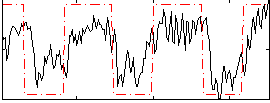 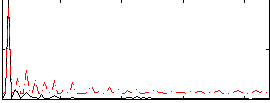 |
Correlation:0.65128
  |
Component:121; Jaccard:0.091949
 |
Component:121; Jaccard:0.1098
 |
Component:259; Jaccard:0.10488
 |
Component:259; Jaccard:0.08257
 |
Component:145; Jaccard:0.058014
 |
Component:121; Jaccard:0.046459
 |
Component:121; Jaccard:0.023969
 |
Correlation:0.24958
  |
Correlation:0.24958
  |
Correlation:-0.26978
  |
Correlation:-0.26978
  |
Correlation:0.65128
  |
Correlation:0.24958
  |
Correlation:0.24958
  |
Component:230; Jaccard:0.08781
 |
Component:214; Jaccard:0.097395
 |
Component:214; Jaccard:0.099637
 |
Component:214; Jaccard:0.081789
 |
Component:214; Jaccard:0.055556
 |
Component:214; Jaccard:0.028622
 |
Component:314; Jaccard:0.017099
 |
Correlation:-0.019682
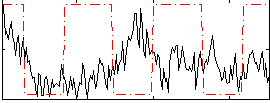 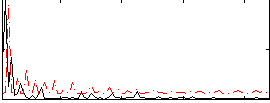 |
Correlation:0.14622
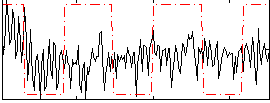 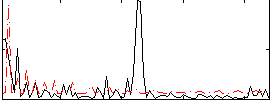 |
Correlation:0.14622
  |
Correlation:0.14622
  |
Correlation:0.14622
  |
Correlation:0.14622
  |
Correlation:0.13421
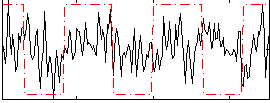 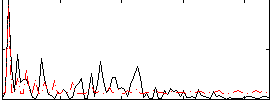 |
Component:214; Jaccard:0.083417
 |
Component:230; Jaccard:0.095956
 |
Component:230; Jaccard:0.093233
 |
Component:306; Jaccard:0.073554
 |
Component:259; Jaccard:0.048493
 |
Component:306; Jaccard:0.024933
 |
Component:227; Jaccard:0.013333
 |
Correlation:0.14622
  |
Correlation:-0.019682
  |
Correlation:-0.019682
  |
Correlation:0.032832
  |
Correlation:-0.26978
  |
Correlation:0.032832
  |
Correlation:0.029555
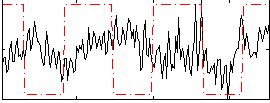 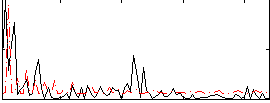 |
Component:086; Jaccard:0.083113
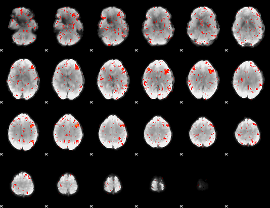 |
Component:306; Jaccard:0.088233
 |
Component:306; Jaccard:0.085108
 |
Component:230; Jaccard:0.073286
 |
Component:306; Jaccard:0.047078
 |
Component:259; Jaccard:0.024084
 |
Component:306; Jaccard:0.012696
 |
Correlation:0.31897
 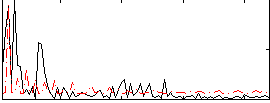 |
Correlation:0.032832
  |
Correlation:0.032832
  |
Correlation:-0.019682
  |
Correlation:0.032832
  |
Correlation:-0.26978
  |
Correlation:0.032832
  |
Component:223; Jaccard:0.082322
 |
Component:086; Jaccard:0.087733
 |
Component:086; Jaccard:0.08412
 |
Component:223; Jaccard:0.066012
 |
Component:194; Jaccard:0.042049
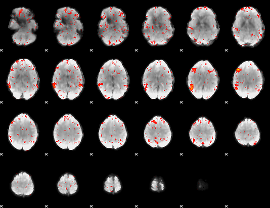 |
Component:318; Jaccard:0.023954
 |
Component:223; Jaccard:0.012172
 |
Correlation:0.08502
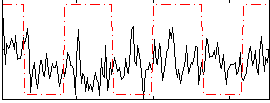  |
Correlation:0.31897
  |
Correlation:0.31897
  |
Correlation:0.08502
  |
Correlation:0.45521
  |
Correlation:0.2146
  |
Correlation:0.08502
  |
Component:306; Jaccard:0.080629
 |
Component:223; Jaccard:0.084521
 |
Component:223; Jaccard:0.080231
 |
Component:086; Jaccard:0.064908
 |
Component:223; Jaccard:0.041861
 |
Component:223; Jaccard:0.023361
 |
Component:194; Jaccard:0.012138
 |
Correlation:0.032832
  |
Correlation:0.08502
  |
Correlation:0.08502
  |
Correlation:0.31897
  |
Correlation:0.08502
  |
Correlation:0.08502
  |
Correlation:0.45521
  |
Component:194; Jaccard:0.077327
 |
Component:194; Jaccard:0.080839
 |
Component:194; Jaccard:0.079606
 |
Component:194; Jaccard:0.064289
 |
Component:230; Jaccard:0.041263
 |
Component:194; Jaccard:0.02324
 |
Component:318; Jaccard:0.012107
 |
Correlation:0.45521
  |
Correlation:0.45521
  |
Correlation:0.45521
  |
Correlation:0.45521
  |
Correlation:-0.019682
  |
Correlation:0.45521
  |
Correlation:0.2146
  |
Component:119; Jaccard:0.072355
 |
Component:242; Jaccard:0.072697
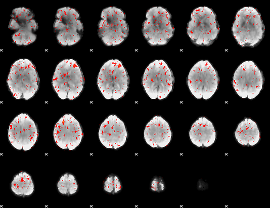 |
Component:242; Jaccard:0.065826
 |
Component:145; Jaccard:0.0642
 |
Component:318; Jaccard:0.039306
 |
Component:314; Jaccard:0.02264
 |
Component:230; Jaccard:0.012047
 |
Correlation:0.084292
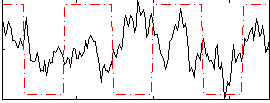 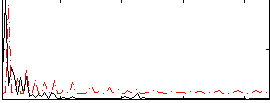 |
Correlation:-0.045332
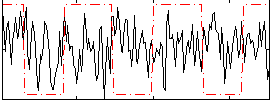 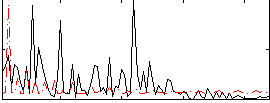 |
Correlation:-0.045332
  |
Correlation:0.65128
  |
Correlation:0.2146
  |
Correlation:0.13421
  |
Correlation:-0.019682
  |
Cope: 06:"SHAPES-FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
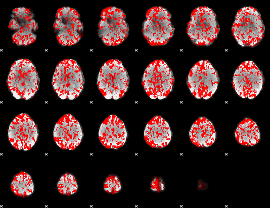 |
GLM binary result thresholded by 1.5
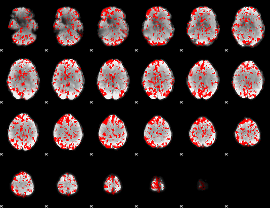 |
GLM binary result thresholded by 2
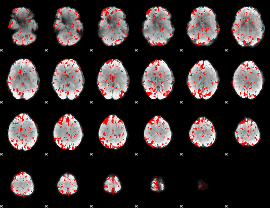 |
GLM binary result thresholded by 2.5
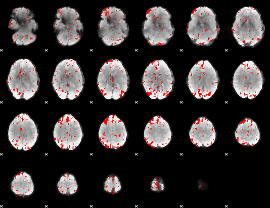 |
GLM binary result thresholded by 3
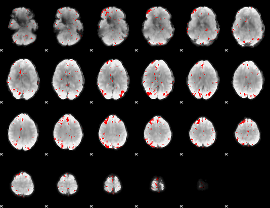 |
GLM binary result thresholded by 3.5
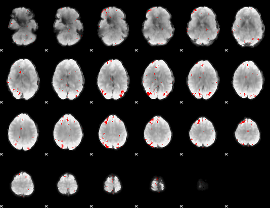 |
GLM binary result thresholded by 4
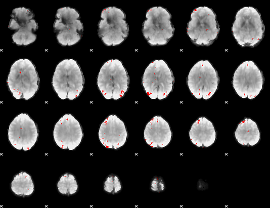 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
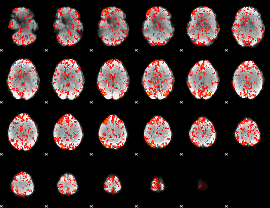 |
GLM Z-score result thresholded by 2
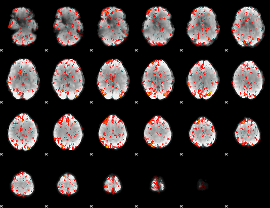 |
GLM Z-score result thresholded by 2.5
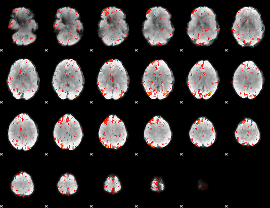 |
GLM Z-score result thresholded by 3
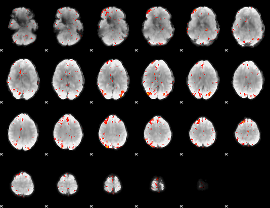 |
GLM Z-score result thresholded by 3.5
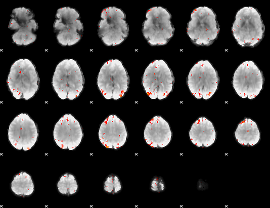 |
GLM Z-score result thresholded by 4
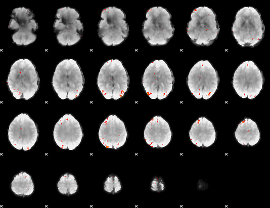 |
Component:034; Jaccard:0.10792
 |
Component:034; Jaccard:0.12988
 |
Component:204; Jaccard:0.1508
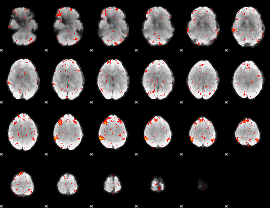 |
Component:204; Jaccard:0.15366
 |
Component:204; Jaccard:0.12998
 |
Component:034; Jaccard:0.085951
 |
Component:034; Jaccard:0.049704
 |
Correlation:0.42243
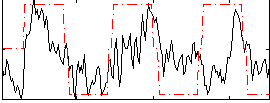 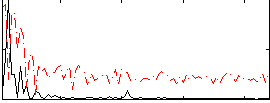 |
Correlation:0.42243
  |
Correlation:0.40429
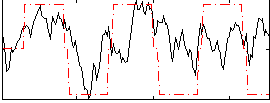 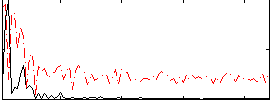 |
Correlation:0.40429
  |
Correlation:0.40429
  |
Correlation:0.42243
  |
Correlation:0.42243
  |
Component:259; Jaccard:0.10192
 |
Component:204; Jaccard:0.12585
 |
Component:034; Jaccard:0.14592
 |
Component:034; Jaccard:0.14501
 |
Component:034; Jaccard:0.1215
 |
Component:204; Jaccard:0.084185
 |
Component:265; Jaccard:0.046896
 |
Correlation:0.36961
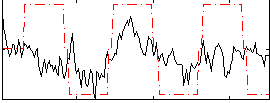 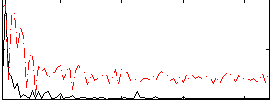 |
Correlation:0.40429
  |
Correlation:0.42243
  |
Correlation:0.42243
  |
Correlation:0.42243
  |
Correlation:0.40429
  |
Correlation:0.24773
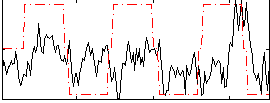 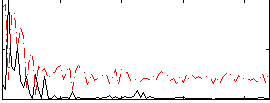 |
Component:264; Jaccard:0.10034
 |
Component:264; Jaccard:0.11883
 |
Component:264; Jaccard:0.13079
 |
Component:264; Jaccard:0.11975
 |
Component:264; Jaccard:0.090385
 |
Component:265; Jaccard:0.070189
 |
Component:204; Jaccard:0.045612
 |
Correlation:0.2411
  |
Correlation:0.2411
  |
Correlation:0.2411
  |
Correlation:0.2411
  |
Correlation:0.2411
  |
Correlation:0.24773
  |
Correlation:0.40429
  |
Component:204; Jaccard:0.099633
 |
Component:259; Jaccard:0.11758
 |
Component:259; Jaccard:0.12581
 |
Component:259; Jaccard:0.11644
 |
Component:011; Jaccard:0.088793
 |
Component:011; Jaccard:0.068311
 |
Component:011; Jaccard:0.043149
 |
Correlation:0.40429
  |
Correlation:0.36961
  |
Correlation:0.36961
  |
Correlation:0.36961
  |
Correlation:0.24648
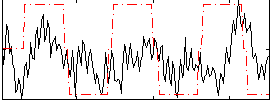  |
Correlation:0.24648
  |
Correlation:0.24648
  |
Component:029; Jaccard:0.096922
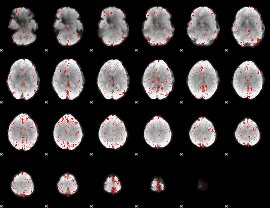 |
Component:029; Jaccard:0.11062
 |
Component:029; Jaccard:0.11524
 |
Component:029; Jaccard:0.10382
 |
Component:265; Jaccard:0.088615
 |
Component:368; Jaccard:0.060125
 |
Component:368; Jaccard:0.036607
 |
Correlation:0.19559
  |
Correlation:0.19559
  |
Correlation:0.19559
  |
Correlation:0.19559
  |
Correlation:0.24773
  |
Correlation:0.29327
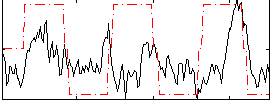 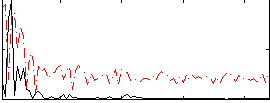 |
Correlation:0.29327
  |
Component:183; Jaccard:0.091386
 |
Component:222; Jaccard:0.10226
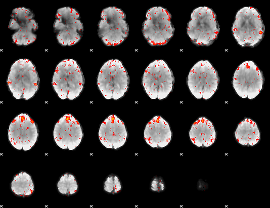 |
Component:222; Jaccard:0.10551
 |
Component:011; Jaccard:0.10033
 |
Component:259; Jaccard:0.088162
 |
Component:264; Jaccard:0.054615
 |
Component:259; Jaccard:0.030326
 |
Correlation:0.066101
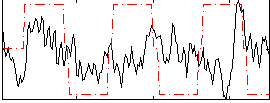  |
Correlation:0.096719
  |
Correlation:0.096719
  |
Correlation:0.24648
  |
Correlation:0.36961
  |
Correlation:0.2411
  |
Correlation:0.36961
  |
Component:222; Jaccard:0.090507
 |
Component:183; Jaccard:0.098107
 |
Component:265; Jaccard:0.098825
 |
Component:265; Jaccard:0.099283
 |
Component:368; Jaccard:0.083504
 |
Component:259; Jaccard:0.052352
 |
Component:264; Jaccard:0.029395
 |
Correlation:0.096719
  |
Correlation:0.066101
  |
Correlation:0.24773
  |
Correlation:0.24773
  |
Correlation:0.29327
  |
Correlation:0.36961
  |
Correlation:0.2411
  |
Component:319; Jaccard:0.084653
 |
Component:189; Jaccard:0.09328
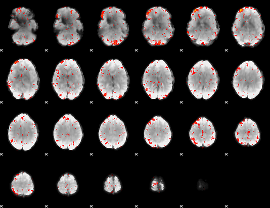 |
Component:011; Jaccard:0.098495
 |
Component:368; Jaccard:0.096567
 |
Component:189; Jaccard:0.075814
 |
Component:189; Jaccard:0.049656
 |
Component:282; Jaccard:0.029044
 |
Correlation:0.23718
 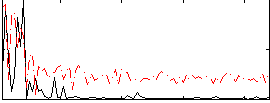 |
Correlation:-0.056903
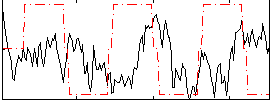 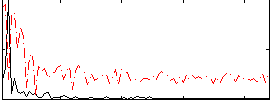 |
Correlation:0.24648
  |
Correlation:0.29327
  |
Correlation:-0.056903
  |
Correlation:-0.056903
  |
Correlation:0.33235
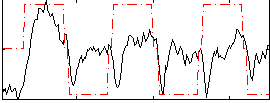  |
Component:342; Jaccard:0.084625
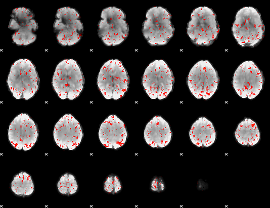 |
Component:319; Jaccard:0.092276
 |
Component:189; Jaccard:0.098004
 |
Component:222; Jaccard:0.095131
 |
Component:029; Jaccard:0.072855
 |
Component:282; Jaccard:0.048146
 |
Component:189; Jaccard:0.027744
 |
Correlation:0.25547
  |
Correlation:0.23718
  |
Correlation:-0.056903
  |
Correlation:0.096719
  |
Correlation:0.19559
  |
Correlation:0.33235
  |
Correlation:-0.056903
  |
Component:189; Jaccard:0.084472
 |
Component:342; Jaccard:0.091195
 |
Component:183; Jaccard:0.095788
 |
Component:189; Jaccard:0.093649
 |
Component:282; Jaccard:0.0653
 |
Component:252; Jaccard:0.044648
 |
Component:252; Jaccard:0.02663
 |
Correlation:-0.056903
  |
Correlation:0.25547
  |
Correlation:0.066101
  |
Correlation:-0.056903
  |
Correlation:0.33235
  |
Correlation:0.18811
  |
Correlation:0.18811
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































