Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:215; Jaccard:0.14057
 |
Component:215; Jaccard:0.17218
 |
Component:215; Jaccard:0.20527
 |
Component:215; Jaccard:0.23033
 |
Component:215; Jaccard:0.23355
 |
Component:215; Jaccard:0.209
 |
Component:215; Jaccard:0.16091
 |
Correlation:0.096131
  |
Correlation:0.096131
  |
Correlation:0.096131
  |
Correlation:0.096131
  |
Correlation:0.096131
  |
Correlation:0.096131
  |
Correlation:0.096131
  |
Component:120; Jaccard:0.129
 |
Component:120; Jaccard:0.15177
 |
Component:120; Jaccard:0.17199
 |
Component:120; Jaccard:0.18275
 |
Component:222; Jaccard:0.18198
 |
Component:222; Jaccard:0.17102
 |
Component:222; Jaccard:0.14189
 |
Correlation:0.16123
  |
Correlation:0.16123
  |
Correlation:0.16123
  |
Correlation:0.16123
  |
Correlation:0.17988
  |
Correlation:0.17988
  |
Correlation:0.17988
  |
Component:222; Jaccard:0.11064
 |
Component:222; Jaccard:0.13442
 |
Component:222; Jaccard:0.15768
 |
Component:222; Jaccard:0.17463
 |
Component:120; Jaccard:0.17664
 |
Component:120; Jaccard:0.1483
 |
Component:256; Jaccard:0.11977
 |
Correlation:0.17988
  |
Correlation:0.17988
  |
Correlation:0.17988
  |
Correlation:0.17988
  |
Correlation:0.16123
  |
Correlation:0.16123
  |
Correlation:0.2218
  |
Component:281; Jaccard:0.10604
 |
Component:281; Jaccard:0.12263
 |
Component:256; Jaccard:0.14024
 |
Component:256; Jaccard:0.15308
 |
Component:256; Jaccard:0.15528
 |
Component:256; Jaccard:0.14489
 |
Component:357; Jaccard:0.11569
 |
Correlation:0.31949
  |
Correlation:0.31949
  |
Correlation:0.2218
  |
Correlation:0.2218
  |
Correlation:0.2218
  |
Correlation:0.2218
  |
Correlation:0.28165
  |
Component:211; Jaccard:0.10561
 |
Component:256; Jaccard:0.12194
 |
Component:357; Jaccard:0.13926
 |
Component:357; Jaccard:0.15129
 |
Component:357; Jaccard:0.15264
 |
Component:357; Jaccard:0.14057
 |
Component:120; Jaccard:0.10849
 |
Correlation:0.14299
  |
Correlation:0.2218
  |
Correlation:0.28165
  |
Correlation:0.28165
  |
Correlation:0.28165
  |
Correlation:0.28165
  |
Correlation:0.16123
  |
Component:256; Jaccard:0.10267
 |
Component:357; Jaccard:0.12192
 |
Component:281; Jaccard:0.13773
 |
Component:281; Jaccard:0.14777
 |
Component:281; Jaccard:0.14721
 |
Component:281; Jaccard:0.12831
 |
Component:150; Jaccard:0.10367
 |
Correlation:0.2218
  |
Correlation:0.28165
  |
Correlation:0.31949
  |
Correlation:0.31949
  |
Correlation:0.31949
  |
Correlation:0.31949
  |
Correlation:0.036421
  |
Component:357; Jaccard:0.10206
 |
Component:211; Jaccard:0.11575
 |
Component:211; Jaccard:0.12247
 |
Component:150; Jaccard:0.12139
 |
Component:150; Jaccard:0.12838
 |
Component:150; Jaccard:0.1227
 |
Component:281; Jaccard:0.097672
 |
Correlation:0.28165
  |
Correlation:0.14299
  |
Correlation:0.14299
  |
Correlation:0.036421
  |
Correlation:0.036421
  |
Correlation:0.036421
  |
Correlation:0.31949
  |
Component:114; Jaccard:0.09562
 |
Component:114; Jaccard:0.10679
 |
Component:344; Jaccard:0.11465
 |
Component:344; Jaccard:0.12123
 |
Component:344; Jaccard:0.11805
 |
Component:033; Jaccard:0.10644
 |
Component:033; Jaccard:0.085352
 |
Correlation:0.0024453
  |
Correlation:0.0024453
  |
Correlation:0.59866
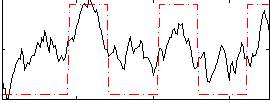  |
Correlation:0.59866
  |
Correlation:0.59866
  |
Correlation:0.34588
  |
Correlation:0.34588
  |
Component:372; Jaccard:0.094166
 |
Component:344; Jaccard:0.10406
 |
Component:114; Jaccard:0.11137
 |
Component:211; Jaccard:0.1208
 |
Component:033; Jaccard:0.11341
 |
Component:344; Jaccard:0.10368
 |
Component:344; Jaccard:0.082799
 |
Correlation:0.12439
  |
Correlation:0.59866
  |
Correlation:0.0024453
  |
Correlation:0.14299
  |
Correlation:0.34588
  |
Correlation:0.59866
  |
Correlation:0.59866
  |
Component:257; Jaccard:0.093625
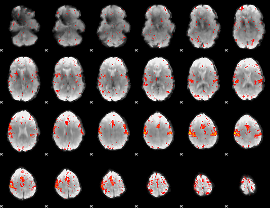 |
Component:257; Jaccard:0.10397
 |
Component:257; Jaccard:0.11024
 |
Component:257; Jaccard:0.11043
 |
Component:211; Jaccard:0.1052
 |
Component:255; Jaccard:0.085936
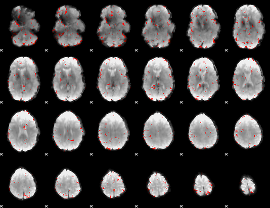 |
Component:255; Jaccard:0.07781
 |
Correlation:0.14328
  |
Correlation:0.14328
  |
Correlation:0.14328
  |
Correlation:0.14328
  |
Correlation:0.14299
  |
Correlation:0.21226
  |
Correlation:0.21226
  |
Cope: 02:"SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
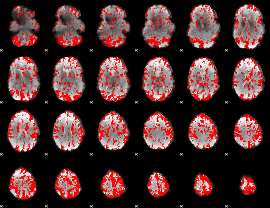 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
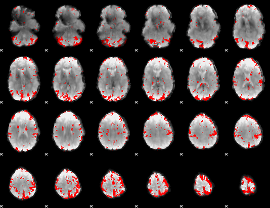 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
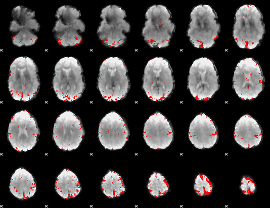 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:215; Jaccard:0.14221
 |
Component:215; Jaccard:0.18096
 |
Component:215; Jaccard:0.22723
 |
Component:215; Jaccard:0.27014
 |
Component:215; Jaccard:0.29274
 |
Component:215; Jaccard:0.28218
 |
Component:215; Jaccard:0.23994
 |
Correlation:0.27356
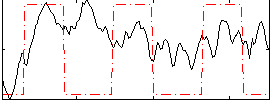 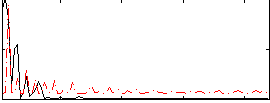 |
Correlation:0.27356
  |
Correlation:0.27356
  |
Correlation:0.27356
  |
Correlation:0.27356
  |
Correlation:0.27356
  |
Correlation:0.27356
  |
Component:120; Jaccard:0.12462
 |
Component:120; Jaccard:0.1493
 |
Component:222; Jaccard:0.18065
 |
Component:222; Jaccard:0.21558
 |
Component:222; Jaccard:0.23857
 |
Component:222; Jaccard:0.24268
 |
Component:222; Jaccard:0.22171
 |
Correlation:0.084684
  |
Correlation:0.084684
  |
Correlation:0.18585
  |
Correlation:0.18585
  |
Correlation:0.18585
  |
Correlation:0.18585
  |
Correlation:0.18585
  |
Component:211; Jaccard:0.11416
 |
Component:222; Jaccard:0.14435
 |
Component:120; Jaccard:0.17415
 |
Component:120; Jaccard:0.18871
 |
Component:120; Jaccard:0.18673
 |
Component:120; Jaccard:0.16739
 |
Component:120; Jaccard:0.13667
 |
Correlation:0.13476
  |
Correlation:0.18585
  |
Correlation:0.084684
  |
Correlation:0.084684
  |
Correlation:0.084684
  |
Correlation:0.084684
  |
Correlation:0.084684
  |
Component:222; Jaccard:0.11284
 |
Component:211; Jaccard:0.13154
 |
Component:211; Jaccard:0.14826
 |
Component:161; Jaccard:0.16126
 |
Component:161; Jaccard:0.16728
 |
Component:161; Jaccard:0.15966
 |
Component:161; Jaccard:0.13639
 |
Correlation:0.18585
  |
Correlation:0.13476
  |
Correlation:0.13476
  |
Correlation:0.2941
  |
Correlation:0.2941
  |
Correlation:0.2941
  |
Correlation:0.2941
  |
Component:230; Jaccard:0.1069
 |
Component:161; Jaccard:0.12648
 |
Component:161; Jaccard:0.14668
 |
Component:357; Jaccard:0.15682
 |
Component:357; Jaccard:0.16064
 |
Component:357; Jaccard:0.15258
 |
Component:357; Jaccard:0.12603
 |
Correlation:0.17615
  |
Correlation:0.2941
  |
Correlation:0.2941
  |
Correlation:0.089
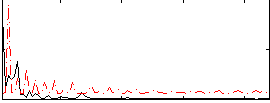  |
Correlation:0.089
  |
Correlation:0.089
  |
Correlation:0.089
  |
Component:161; Jaccard:0.1063
 |
Component:114; Jaccard:0.12559
 |
Component:114; Jaccard:0.14427
 |
Component:114; Jaccard:0.15428
 |
Component:114; Jaccard:0.1509
 |
Component:114; Jaccard:0.13813
 |
Component:281; Jaccard:0.1146
 |
Correlation:0.2941
  |
Correlation:0.24447
  |
Correlation:0.24447
  |
Correlation:0.24447
  |
Correlation:0.24447
  |
Correlation:0.24447
  |
Correlation:0.016305
  |
Component:114; Jaccard:0.10526
 |
Component:357; Jaccard:0.12048
 |
Component:357; Jaccard:0.14227
 |
Component:211; Jaccard:0.15298
 |
Component:211; Jaccard:0.1493
 |
Component:281; Jaccard:0.13434
 |
Component:114; Jaccard:0.11364
 |
Correlation:0.24447
  |
Correlation:0.089
  |
Correlation:0.089
  |
Correlation:0.13476
  |
Correlation:0.13476
  |
Correlation:0.016305
  |
Correlation:0.24447
  |
Component:281; Jaccard:0.10079
 |
Component:230; Jaccard:0.11909
 |
Component:281; Jaccard:0.12965
 |
Component:281; Jaccard:0.14071
 |
Component:281; Jaccard:0.14223
 |
Component:211; Jaccard:0.132
 |
Component:256; Jaccard:0.10909
 |
Correlation:0.016305
  |
Correlation:0.17615
  |
Correlation:0.016305
  |
Correlation:0.016305
  |
Correlation:0.016305
  |
Correlation:0.13476
  |
Correlation:-0.01182
  |
Component:251; Jaccard:0.10048
 |
Component:281; Jaccard:0.11542
 |
Component:230; Jaccard:0.12708
 |
Component:256; Jaccard:0.13171
 |
Component:256; Jaccard:0.13312
 |
Component:256; Jaccard:0.12653
 |
Component:211; Jaccard:0.10402
 |
Correlation:0.15153
  |
Correlation:0.016305
  |
Correlation:0.17615
  |
Correlation:-0.01182
  |
Correlation:-0.01182
  |
Correlation:-0.01182
  |
Correlation:0.13476
  |
Component:357; Jaccard:0.10038
 |
Component:251; Jaccard:0.11377
 |
Component:251; Jaccard:0.12404
 |
Component:251; Jaccard:0.13021
 |
Component:251; Jaccard:0.12655
 |
Component:251; Jaccard:0.11127
 |
Component:150; Jaccard:0.091409
 |
Correlation:0.089
  |
Correlation:0.15153
  |
Correlation:0.15153
  |
Correlation:0.15153
  |
Correlation:0.15153
  |
Correlation:0.15153
  |
Correlation:0.24353
  |
Cope: 03:"FACES-SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
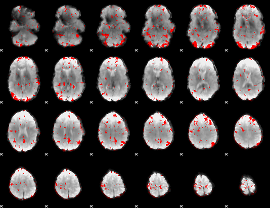 |
GLM binary result thresholded by 2.5
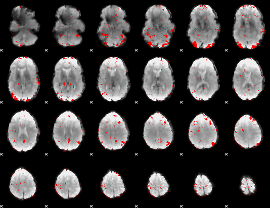 |
GLM binary result thresholded by 3
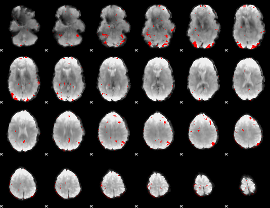 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:191; Jaccard:0.11915
 |
Component:191; Jaccard:0.14948
 |
Component:191; Jaccard:0.17271
 |
Component:191; Jaccard:0.17486
 |
Component:191; Jaccard:0.15795
 |
Component:191; Jaccard:0.13258
 |
Component:191; Jaccard:0.10202
 |
Correlation:0.5746
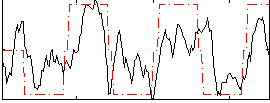 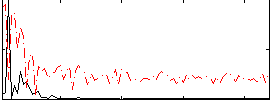 |
Correlation:0.5746
  |
Correlation:0.5746
  |
Correlation:0.5746
  |
Correlation:0.5746
  |
Correlation:0.5746
  |
Correlation:0.5746
  |
Component:373; Jaccard:0.11276
 |
Component:162; Jaccard:0.11509
 |
Component:334; Jaccard:0.13024
 |
Component:334; Jaccard:0.13397
 |
Component:334; Jaccard:0.12359
 |
Component:334; Jaccard:0.09598
 |
Component:334; Jaccard:0.071001
 |
Correlation:0.14566
  |
Correlation:0.20328
  |
Correlation:0.12674
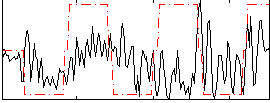  |
Correlation:0.12674
  |
Correlation:0.12674
  |
Correlation:0.12674
  |
Correlation:0.12674
  |
Component:162; Jaccard:0.1033
 |
Component:373; Jaccard:0.1146
 |
Component:091; Jaccard:0.11716
 |
Component:091; Jaccard:0.11679
 |
Component:091; Jaccard:0.10328
 |
Component:091; Jaccard:0.082338
 |
Component:091; Jaccard:0.059224
 |
Correlation:0.20328
  |
Correlation:0.14566
  |
Correlation:0.095954
  |
Correlation:0.095954
  |
Correlation:0.095954
  |
Correlation:0.095954
  |
Correlation:0.095954
  |
Component:344; Jaccard:0.096973
 |
Component:334; Jaccard:0.11272
 |
Component:162; Jaccard:0.11255
 |
Component:162; Jaccard:0.098141
 |
Component:344; Jaccard:0.083369
 |
Component:344; Jaccard:0.066955
 |
Component:344; Jaccard:0.048926
 |
Correlation:0.4801
  |
Correlation:0.12674
  |
Correlation:0.20328
  |
Correlation:0.20328
  |
Correlation:0.4801
  |
Correlation:0.4801
  |
Correlation:0.4801
  |
Component:375; Jaccard:0.093298
 |
Component:091; Jaccard:0.10564
 |
Component:344; Jaccard:0.10351
 |
Component:344; Jaccard:0.096128
 |
Component:162; Jaccard:0.076893
 |
Component:315; Jaccard:0.060139
 |
Component:315; Jaccard:0.043023
 |
Correlation:0.31634
  |
Correlation:0.095954
  |
Correlation:0.4801
  |
Correlation:0.4801
  |
Correlation:0.20328
  |
Correlation:0.19055
  |
Correlation:0.19055
  |
Component:334; Jaccard:0.091315
 |
Component:344; Jaccard:0.10455
 |
Component:373; Jaccard:0.10189
 |
Component:391; Jaccard:0.0872
 |
Component:315; Jaccard:0.07332
 |
Component:162; Jaccard:0.052744
 |
Component:321; Jaccard:0.035097
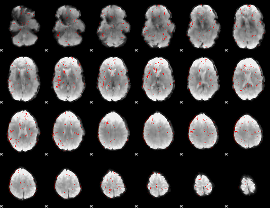 |
Correlation:0.12674
  |
Correlation:0.4801
  |
Correlation:0.14566
  |
Correlation:0.34465
  |
Correlation:0.19055
  |
Correlation:0.20328
  |
Correlation:0.064807
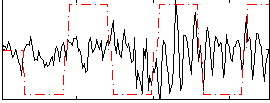  |
Component:091; Jaccard:0.089151
 |
Component:375; Jaccard:0.093933
 |
Component:391; Jaccard:0.092886
 |
Component:315; Jaccard:0.079842
 |
Component:391; Jaccard:0.071255
 |
Component:391; Jaccard:0.051741
 |
Component:393; Jaccard:0.034655
 |
Correlation:0.095954
  |
Correlation:0.31634
  |
Correlation:0.34465
  |
Correlation:0.19055
  |
Correlation:0.34465
  |
Correlation:0.34465
  |
Correlation:0.38371
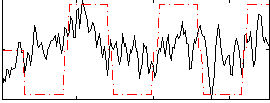  |
Component:033; Jaccard:0.085299
 |
Component:391; Jaccard:0.086969
 |
Component:375; Jaccard:0.087675
 |
Component:373; Jaccard:0.079452
 |
Component:321; Jaccard:0.064289
 |
Component:321; Jaccard:0.04997
 |
Component:104; Jaccard:0.034365
 |
Correlation:0.17873
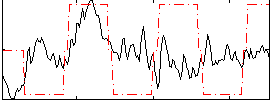  |
Correlation:0.34465
  |
Correlation:0.31634
  |
Correlation:0.14566
  |
Correlation:0.064807
  |
Correlation:0.064807
  |
Correlation:0.42285
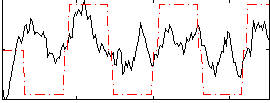  |
Component:140; Jaccard:0.078736
 |
Component:033; Jaccard:0.085113
 |
Component:393; Jaccard:0.082903
 |
Component:321; Jaccard:0.079204
 |
Component:393; Jaccard:0.061626
 |
Component:393; Jaccard:0.047124
 |
Component:391; Jaccard:0.034018
 |
Correlation:0.083313
  |
Correlation:0.17873
  |
Correlation:0.38371
  |
Correlation:0.064807
  |
Correlation:0.38371
  |
Correlation:0.38371
  |
Correlation:0.34465
  |
Component:391; Jaccard:0.078191
 |
Component:393; Jaccard:0.080976
 |
Component:321; Jaccard:0.080613
 |
Component:393; Jaccard:0.074027
 |
Component:104; Jaccard:0.058255
 |
Component:104; Jaccard:0.045098
 |
Component:162; Jaccard:0.03354
 |
Correlation:0.34465
  |
Correlation:0.38371
  |
Correlation:0.064807
  |
Correlation:0.38371
  |
Correlation:0.42285
  |
Correlation:0.42285
  |
Correlation:0.20328
  |
Cope: 04:"neg_FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
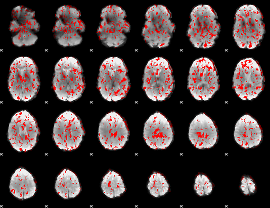 |
GLM binary result thresholded by 1.5
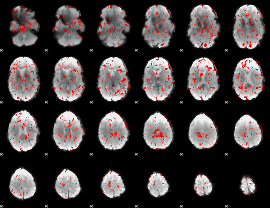 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
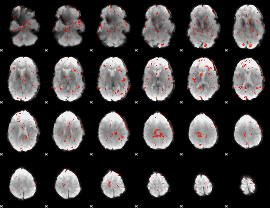 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
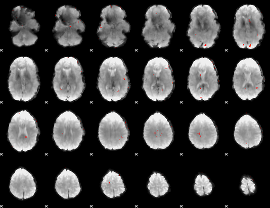 |
GLM Z-score result thresholded by 4
 |
Component:282; Jaccard:0.077675
 |
Component:023; Jaccard:0.091232
 |
Component:214; Jaccard:0.10621
 |
Component:214; Jaccard:0.11271
 |
Component:214; Jaccard:0.081943
 |
Component:214; Jaccard:0.041377
 |
Component:214; Jaccard:0.015718
 |
Correlation:-0.090725
  |
Correlation:-0.13937
  |
Correlation:0.13578
  |
Correlation:0.13578
  |
Correlation:0.13578
  |
Correlation:0.13578
  |
Correlation:0.13578
  |
Component:131; Jaccard:0.076532
 |
Component:282; Jaccard:0.08529
 |
Component:023; Jaccard:0.10211
 |
Component:023; Jaccard:0.089868
 |
Component:023; Jaccard:0.053586
 |
Component:023; Jaccard:0.019711
 |
Component:380; Jaccard:0.006881
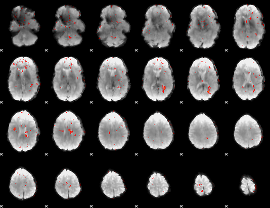 |
Correlation:0.018906
  |
Correlation:-0.090725
  |
Correlation:-0.13937
  |
Correlation:-0.13937
  |
Correlation:-0.13937
  |
Correlation:-0.13937
  |
Correlation:0.14735
  |
Component:252; Jaccard:0.076474
 |
Component:214; Jaccard:0.083442
 |
Component:282; Jaccard:0.078067
 |
Component:009; Jaccard:0.06013
 |
Component:009; Jaccard:0.035418
 |
Component:009; Jaccard:0.017383
 |
Component:023; Jaccard:0.006033
 |
Correlation:-0.13752
  |
Correlation:0.13578
  |
Correlation:-0.090725
  |
Correlation:-0.088441
  |
Correlation:-0.088441
  |
Correlation:-0.088441
  |
Correlation:-0.13937
  |
Component:023; Jaccard:0.072417
 |
Component:278; Jaccard:0.073599
 |
Component:278; Jaccard:0.067707
 |
Component:282; Jaccard:0.058279
 |
Component:282; Jaccard:0.032905
 |
Component:380; Jaccard:0.017198
 |
Component:009; Jaccard:0.005971
 |
Correlation:-0.13937
  |
Correlation:-0.26586
  |
Correlation:-0.26586
  |
Correlation:-0.090725
  |
Correlation:-0.090725
  |
Correlation:0.14735
  |
Correlation:-0.088441
  |
Component:040; Jaccard:0.071259
 |
Component:131; Jaccard:0.0729
 |
Component:009; Jaccard:0.066987
 |
Component:278; Jaccard:0.051618
 |
Component:380; Jaccard:0.032838
 |
Component:094; Jaccard:0.013975
 |
Component:040; Jaccard:0.005642
 |
Correlation:0.09619
  |
Correlation:0.018906
  |
Correlation:-0.088441
  |
Correlation:-0.26586
  |
Correlation:0.14735
  |
Correlation:0.24706
  |
Correlation:0.09619
  |
Component:105; Jaccard:0.070268
 |
Component:095; Jaccard:0.069787
 |
Component:095; Jaccard:0.059589
 |
Component:094; Jaccard:0.047417
 |
Component:094; Jaccard:0.031583
 |
Component:282; Jaccard:0.013349
 |
Component:282; Jaccard:0.004835
 |
Correlation:-0.046732
  |
Correlation:0.12194
  |
Correlation:0.12194
  |
Correlation:0.24706
  |
Correlation:0.24706
  |
Correlation:-0.090725
  |
Correlation:-0.090725
  |
Component:095; Jaccard:0.070248
 |
Component:040; Jaccard:0.069442
 |
Component:131; Jaccard:0.058767
 |
Component:380; Jaccard:0.04688
 |
Component:278; Jaccard:0.028764
 |
Component:040; Jaccard:0.013082
 |
Component:306; Jaccard:0.004446
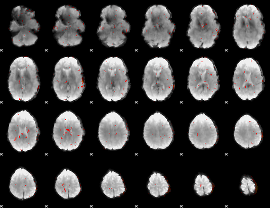 |
Correlation:0.12194
  |
Correlation:0.09619
  |
Correlation:0.018906
  |
Correlation:0.14735
  |
Correlation:-0.26586
  |
Correlation:0.09619
  |
Correlation:-0.23349
  |
Component:278; Jaccard:0.06974
 |
Component:252; Jaccard:0.06888
 |
Component:181; Jaccard:0.058767
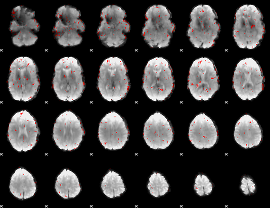 |
Component:181; Jaccard:0.044026
 |
Component:040; Jaccard:0.028129
 |
Component:278; Jaccard:0.012987
 |
Component:278; Jaccard:0.00443
 |
Correlation:-0.26586
  |
Correlation:-0.13752
  |
Correlation:-0.042503
  |
Correlation:-0.042503
  |
Correlation:0.09619
  |
Correlation:-0.26586
  |
Correlation:-0.26586
  |
Component:364; Jaccard:0.069732
 |
Component:105; Jaccard:0.067542
 |
Component:040; Jaccard:0.057859
 |
Component:040; Jaccard:0.043897
 |
Component:306; Jaccard:0.027044
 |
Component:021; Jaccard:0.011711
 |
Component:364; Jaccard:0.004136
 |
Correlation:-0.14669
  |
Correlation:-0.046732
  |
Correlation:0.09619
  |
Correlation:0.09619
  |
Correlation:-0.23349
  |
Correlation:0.10557
 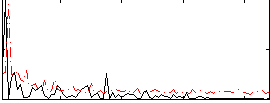 |
Correlation:-0.14669
  |
Component:080; Jaccard:0.066092
 |
Component:364; Jaccard:0.067454
 |
Component:394; Jaccard:0.056394
 |
Component:061; Jaccard:0.04116
 |
Component:021; Jaccard:0.026057
 |
Component:364; Jaccard:0.011074
 |
Component:147; Jaccard:0.003949
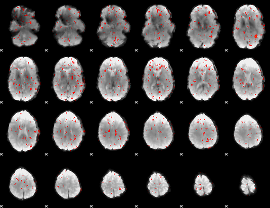 |
Correlation:0.23099
  |
Correlation:-0.14669
  |
Correlation:0.079244
  |
Correlation:0.0085076
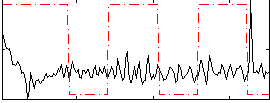  |
Correlation:0.10557
  |
Correlation:-0.14669
  |
Correlation:0.032422
  |
Cope: 05:"neg_SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:278; Jaccard:0.087586
 |
Component:023; Jaccard:0.11423
 |
Component:023; Jaccard:0.13451
 |
Component:023; Jaccard:0.13097
 |
Component:023; Jaccard:0.10356
 |
Component:023; Jaccard:0.066024
 |
Component:023; Jaccard:0.037859
 |
Correlation:0.33137
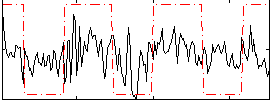 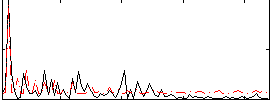 |
Correlation:0.22989
  |
Correlation:0.22989
  |
Correlation:0.22989
  |
Correlation:0.22989
  |
Correlation:0.22989
  |
Correlation:0.22989
  |
Component:023; Jaccard:0.087546
 |
Component:278; Jaccard:0.096753
 |
Component:214; Jaccard:0.096472
 |
Component:214; Jaccard:0.098277
 |
Component:214; Jaccard:0.079123
 |
Component:214; Jaccard:0.048458
 |
Component:214; Jaccard:0.022566
 |
Correlation:0.22989
  |
Correlation:0.33137
  |
Correlation:0.17356
  |
Correlation:0.17356
  |
Correlation:0.17356
  |
Correlation:0.17356
  |
Correlation:0.17356
  |
Component:282; Jaccard:0.081625
 |
Component:334; Jaccard:0.088115
 |
Component:278; Jaccard:0.093444
 |
Component:334; Jaccard:0.083915
 |
Component:247; Jaccard:0.064586
 |
Component:334; Jaccard:0.040478
 |
Component:334; Jaccard:0.020744
 |
Correlation:0.2142
  |
Correlation:0.14489
  |
Correlation:0.33137
  |
Correlation:0.14489
  |
Correlation:0.38118
  |
Correlation:0.14489
  |
Correlation:0.14489
  |
Component:181; Jaccard:0.080194
 |
Component:282; Jaccard:0.084636
 |
Component:334; Jaccard:0.092168
 |
Component:247; Jaccard:0.082051
 |
Component:334; Jaccard:0.062988
 |
Component:247; Jaccard:0.039394
 |
Component:315; Jaccard:0.020679
 |
Correlation:0.085885
  |
Correlation:0.2142
  |
Correlation:0.14489
  |
Correlation:0.38118
  |
Correlation:0.14489
  |
Correlation:0.38118
  |
Correlation:0.12238
 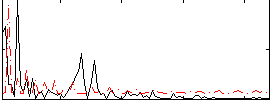 |
Component:334; Jaccard:0.078736
 |
Component:181; Jaccard:0.084571
 |
Component:247; Jaccard:0.087982
 |
Component:278; Jaccard:0.074867
 |
Component:009; Jaccard:0.056729
 |
Component:315; Jaccard:0.037137
 |
Component:247; Jaccard:0.020333
 |
Correlation:0.14489
  |
Correlation:0.085885
  |
Correlation:0.38118
  |
Correlation:0.33137
  |
Correlation:0.25019
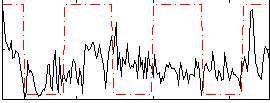  |
Correlation:0.12238
  |
Correlation:0.38118
  |
Component:191; Jaccard:0.075076
 |
Component:247; Jaccard:0.082606
 |
Component:009; Jaccard:0.081478
 |
Component:009; Jaccard:0.071547
 |
Component:306; Jaccard:0.05516
 |
Component:306; Jaccard:0.035512
 |
Component:048; Jaccard:0.018391
 |
Correlation:0.47041
  |
Correlation:0.38118
  |
Correlation:0.25019
  |
Correlation:0.25019
  |
Correlation:0.41308
  |
Correlation:0.41308
  |
Correlation:0.19935
  |
Component:247; Jaccard:0.0707
 |
Component:214; Jaccard:0.079829
 |
Component:181; Jaccard:0.081167
 |
Component:315; Jaccard:0.06679
 |
Component:315; Jaccard:0.052824
 |
Component:009; Jaccard:0.033895
 |
Component:306; Jaccard:0.01826
 |
Correlation:0.38118
  |
Correlation:0.17356
  |
Correlation:0.085885
  |
Correlation:0.12238
  |
Correlation:0.12238
  |
Correlation:0.25019
  |
Correlation:0.41308
  |
Component:279; Jaccard:0.06996
 |
Component:009; Jaccard:0.077814
 |
Component:282; Jaccard:0.077879
 |
Component:181; Jaccard:0.066116
 |
Component:278; Jaccard:0.051301
 |
Component:048; Jaccard:0.031331
 |
Component:009; Jaccard:0.017348
 |
Correlation:0.16753
  |
Correlation:0.25019
  |
Correlation:0.2142
  |
Correlation:0.085885
  |
Correlation:0.33137
  |
Correlation:0.19935
  |
Correlation:0.25019
  |
Component:091; Jaccard:0.069838
 |
Component:191; Jaccard:0.075787
 |
Component:315; Jaccard:0.074076
 |
Component:306; Jaccard:0.065761
 |
Component:048; Jaccard:0.045727
 |
Component:091; Jaccard:0.027506
 |
Component:191; Jaccard:0.016264
 |
Correlation:0.029485
  |
Correlation:0.47041
  |
Correlation:0.12238
  |
Correlation:0.41308
  |
Correlation:0.19935
  |
Correlation:0.029485
  |
Correlation:0.47041
  |
Component:315; Jaccard:0.069376
 |
Component:315; Jaccard:0.073856
 |
Component:191; Jaccard:0.068726
 |
Component:282; Jaccard:0.061071
 |
Component:091; Jaccard:0.043274
 |
Component:380; Jaccard:0.026626
 |
Component:391; Jaccard:0.014005
 |
Correlation:0.12238
  |
Correlation:0.12238
  |
Correlation:0.47041
  |
Correlation:0.2142
  |
Correlation:0.029485
  |
Correlation:0.1897
  |
Correlation:0.28296
 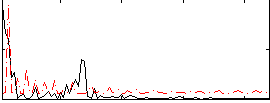 |
Cope: 06:"SHAPES-FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
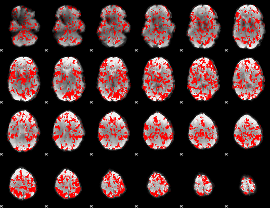 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:108; Jaccard:0.14032
 |
Component:108; Jaccard:0.15624
 |
Component:108; Jaccard:0.14719
 |
Component:106; Jaccard:0.12215
 |
Component:106; Jaccard:0.08002
 |
Component:106; Jaccard:0.040744
 |
Component:106; Jaccard:0.01583
 |
Correlation:0.17928
  |
Correlation:0.17928
  |
Correlation:0.17928
  |
Correlation:0.17328
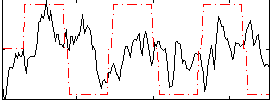  |
Correlation:0.17328
  |
Correlation:0.17328
  |
Correlation:0.17328
  |
Component:187; Jaccard:0.11456
 |
Component:106; Jaccard:0.13273
 |
Component:106; Jaccard:0.1397
 |
Component:108; Jaccard:0.11106
 |
Component:161; Jaccard:0.067772
 |
Component:085; Jaccard:0.035222
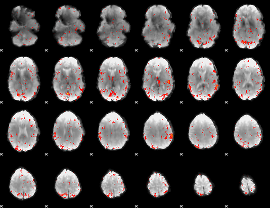 |
Component:085; Jaccard:0.013332
 |
Correlation:0.17733
  |
Correlation:0.17328
  |
Correlation:0.17328
  |
Correlation:0.17928
  |
Correlation:0.1608
  |
Correlation:0.13397
  |
Correlation:0.13397
  |
Component:106; Jaccard:0.11394
 |
Component:161; Jaccard:0.12562
 |
Component:161; Jaccard:0.1247
 |
Component:161; Jaccard:0.10569
 |
Component:085; Jaccard:0.067681
 |
Component:161; Jaccard:0.032244
 |
Component:108; Jaccard:0.013256
 |
Correlation:0.17328
  |
Correlation:0.1608
  |
Correlation:0.1608
  |
Correlation:0.1608
  |
Correlation:0.13397
  |
Correlation:0.1608
  |
Correlation:0.17928
  |
Component:161; Jaccard:0.11309
 |
Component:187; Jaccard:0.1225
 |
Component:085; Jaccard:0.11672
 |
Component:085; Jaccard:0.10139
 |
Component:108; Jaccard:0.066657
 |
Component:108; Jaccard:0.030911
 |
Component:161; Jaccard:0.011832
 |
Correlation:0.1608
  |
Correlation:0.17733
  |
Correlation:0.13397
  |
Correlation:0.13397
  |
Correlation:0.17928
  |
Correlation:0.17928
  |
Correlation:0.1608
  |
Component:230; Jaccard:0.11038
 |
Component:230; Jaccard:0.11465
 |
Component:187; Jaccard:0.1156
 |
Component:187; Jaccard:0.091014
 |
Component:187; Jaccard:0.052029
 |
Component:187; Jaccard:0.023865
 |
Component:328; Jaccard:0.010302
 |
Correlation:0.0101
  |
Correlation:0.0101
  |
Correlation:0.17733
  |
Correlation:0.17733
  |
Correlation:0.17733
  |
Correlation:0.17733
  |
Correlation:0.30845
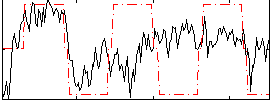  |
Component:085; Jaccard:0.0933
 |
Component:085; Jaccard:0.10975
 |
Component:230; Jaccard:0.10547
 |
Component:230; Jaccard:0.076886
 |
Component:230; Jaccard:0.045264
 |
Component:286; Jaccard:0.021138
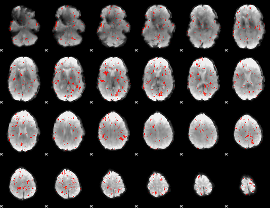 |
Component:187; Jaccard:0.009144
 |
Correlation:0.13397
  |
Correlation:0.13397
  |
Correlation:0.0101
  |
Correlation:0.0101
  |
Correlation:0.0101
  |
Correlation:0.28574
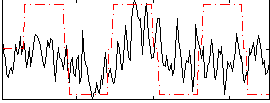  |
Correlation:0.17733
  |
Component:400; Jaccard:0.090967
 |
Component:251; Jaccard:0.092121
 |
Component:251; Jaccard:0.087066
 |
Component:286; Jaccard:0.068858
 |
Component:286; Jaccard:0.04441
 |
Component:230; Jaccard:0.020891
 |
Component:251; Jaccard:0.008522
 |
Correlation:0.19536
  |
Correlation:0.03972
  |
Correlation:0.03972
  |
Correlation:0.28574
  |
Correlation:0.28574
  |
Correlation:0.0101
  |
Correlation:0.03972
  |
Component:251; Jaccard:0.088792
 |
Component:400; Jaccard:0.08975
 |
Component:286; Jaccard:0.081864
 |
Component:251; Jaccard:0.06825
 |
Component:251; Jaccard:0.041609
 |
Component:040; Jaccard:0.020073
 |
Component:040; Jaccard:0.008041
 |
Correlation:0.03972
  |
Correlation:0.19536
  |
Correlation:0.28574
  |
Correlation:0.03972
  |
Correlation:0.03972
  |
Correlation:0.19656
  |
Correlation:0.19656
  |
Component:226; Jaccard:0.083682
 |
Component:226; Jaccard:0.085038
 |
Component:202; Jaccard:0.081102
 |
Component:202; Jaccard:0.061263
 |
Component:040; Jaccard:0.041542
 |
Component:251; Jaccard:0.019461
 |
Component:286; Jaccard:0.00793
 |
Correlation:0.11374
 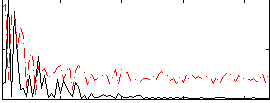 |
Correlation:0.11374
  |
Correlation:0.13848
  |
Correlation:0.13848
  |
Correlation:0.19656
  |
Correlation:0.03972
  |
Correlation:0.28574
  |
Component:169; Jaccard:0.083285
 |
Component:202; Jaccard:0.084215
 |
Component:226; Jaccard:0.078905
 |
Component:040; Jaccard:0.059974
 |
Component:090; Jaccard:0.037799
 |
Component:082; Jaccard:0.018947
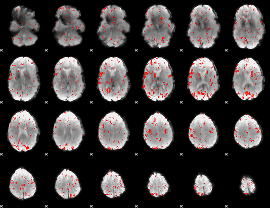 |
Component:094; Jaccard:0.007531
 |
Correlation:0.12226
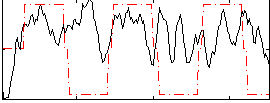  |
Correlation:0.13848
  |
Correlation:0.11374
  |
Correlation:0.19656
  |
Correlation:0.17113
  |
Correlation:0.19091
  |
Correlation:0.21341
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































