Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
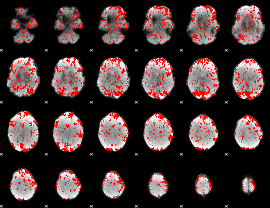 |
GLM binary result thresholded by 1.5
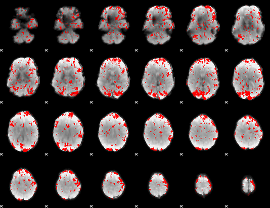 |
GLM binary result thresholded by 2
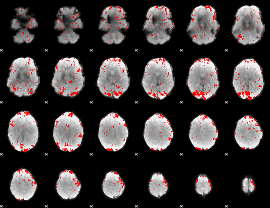 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
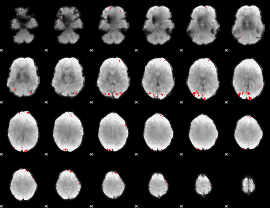 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
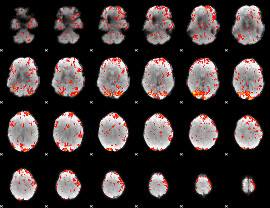 |
GLM Z-score result thresholded by 2
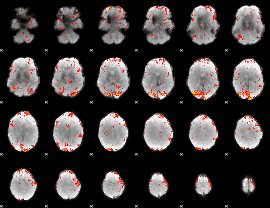 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
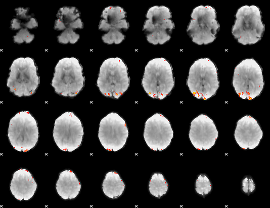 |
Component:357; Jaccard:0.11953
 |
Component:030; Jaccard:0.14951
 |
Component:030; Jaccard:0.17793
 |
Component:030; Jaccard:0.18884
 |
Component:030; Jaccard:0.1773
 |
Component:030; Jaccard:0.14651
 |
Component:030; Jaccard:0.11532
 |
Correlation:-0.10469
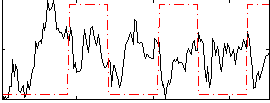  |
Correlation:0.48987
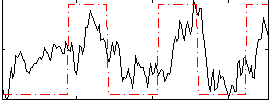 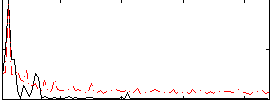 |
Correlation:0.48987
  |
Correlation:0.48987
  |
Correlation:0.48987
  |
Correlation:0.48987
  |
Correlation:0.48987
  |
Component:348; Jaccard:0.11836
 |
Component:357; Jaccard:0.14401
 |
Component:357; Jaccard:0.15783
 |
Component:357; Jaccard:0.15429
 |
Component:357; Jaccard:0.1294
 |
Component:357; Jaccard:0.096702
 |
Component:357; Jaccard:0.068456
 |
Correlation:0.14407
  |
Correlation:-0.10469
  |
Correlation:-0.10469
  |
Correlation:-0.10469
  |
Correlation:-0.10469
  |
Correlation:-0.10469
  |
Correlation:-0.10469
  |
Component:030; Jaccard:0.11396
 |
Component:348; Jaccard:0.13299
 |
Component:372; Jaccard:0.14053
 |
Component:372; Jaccard:0.13383
 |
Component:372; Jaccard:0.10484
 |
Component:372; Jaccard:0.075902
 |
Component:372; Jaccard:0.04931
 |
Correlation:0.48987
  |
Correlation:0.14407
  |
Correlation:0.10431
  |
Correlation:0.10431
  |
Correlation:0.10431
  |
Correlation:0.10431
  |
Correlation:0.10431
  |
Component:372; Jaccard:0.10128
 |
Component:372; Jaccard:0.1257
 |
Component:348; Jaccard:0.13273
 |
Component:348; Jaccard:0.11983
 |
Component:348; Jaccard:0.092389
 |
Component:348; Jaccard:0.064213
 |
Component:348; Jaccard:0.044337
 |
Correlation:0.10431
  |
Correlation:0.10431
  |
Correlation:0.14407
  |
Correlation:0.14407
  |
Correlation:0.14407
  |
Correlation:0.14407
  |
Correlation:0.14407
  |
Component:278; Jaccard:0.097789
 |
Component:044; Jaccard:0.11307
 |
Component:044; Jaccard:0.122
 |
Component:044; Jaccard:0.11132
 |
Component:044; Jaccard:0.089491
 |
Component:044; Jaccard:0.063679
 |
Component:083; Jaccard:0.042961
 |
Correlation:-0.098783
  |
Correlation:0.14661
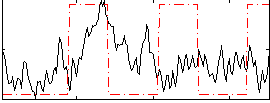  |
Correlation:0.14661
  |
Correlation:0.14661
  |
Correlation:0.14661
  |
Correlation:0.14661
  |
Correlation:0.41145
  |
Component:044; Jaccard:0.096537
 |
Component:278; Jaccard:0.099278
 |
Component:083; Jaccard:0.097449
 |
Component:083; Jaccard:0.091057
 |
Component:083; Jaccard:0.075707
 |
Component:083; Jaccard:0.056104
 |
Component:044; Jaccard:0.038414
 |
Correlation:0.14661
  |
Correlation:-0.098783
  |
Correlation:0.41145
  |
Correlation:0.41145
  |
Correlation:0.41145
  |
Correlation:0.41145
  |
Correlation:0.14661
  |
Component:083; Jaccard:0.08149
 |
Component:083; Jaccard:0.091226
 |
Component:278; Jaccard:0.088932
 |
Component:283; Jaccard:0.07604
 |
Component:283; Jaccard:0.062157
 |
Component:002; Jaccard:0.042475
 |
Component:002; Jaccard:0.032457
 |
Correlation:0.41145
  |
Correlation:0.41145
  |
Correlation:-0.098783
  |
Correlation:-0.13744
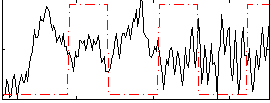  |
Correlation:-0.13744
  |
Correlation:0.18391
  |
Correlation:0.18391
  |
Component:002; Jaccard:0.07999
 |
Component:283; Jaccard:0.08759
 |
Component:283; Jaccard:0.086987
 |
Component:002; Jaccard:0.0695
 |
Component:002; Jaccard:0.058163
 |
Component:283; Jaccard:0.0403
 |
Component:364; Jaccard:0.025896
 |
Correlation:0.18391
  |
Correlation:-0.13744
  |
Correlation:-0.13744
  |
Correlation:0.18391
  |
Correlation:0.18391
  |
Correlation:-0.13744
  |
Correlation:0.15426
  |
Component:283; Jaccard:0.079514
 |
Component:002; Jaccard:0.081489
 |
Component:002; Jaccard:0.077278
 |
Component:362; Jaccard:0.068945
 |
Component:362; Jaccard:0.05362
 |
Component:364; Jaccard:0.038257
 |
Component:283; Jaccard:0.024684
 |
Correlation:-0.13744
  |
Correlation:0.18391
  |
Correlation:0.18391
  |
Correlation:0.13127
  |
Correlation:0.13127
  |
Correlation:0.15426
  |
Correlation:-0.13744
  |
Component:297; Jaccard:0.078124
 |
Component:297; Jaccard:0.078536
 |
Component:362; Jaccard:0.073916
 |
Component:278; Jaccard:0.06759
 |
Component:364; Jaccard:0.051149
 |
Component:362; Jaccard:0.036595
 |
Component:148; Jaccard:0.023083
 |
Correlation:0.10672
  |
Correlation:0.10672
  |
Correlation:0.13127
  |
Correlation:-0.098783
  |
Correlation:0.15426
  |
Correlation:0.13127
  |
Correlation:0.17577
 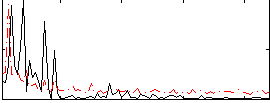 |
Cope: 02:"SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
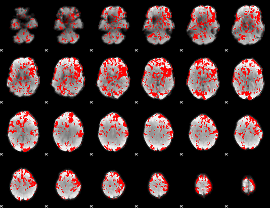 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
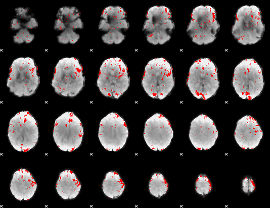 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
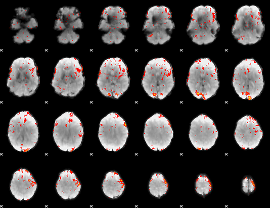 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:357; Jaccard:0.12523
 |
Component:357; Jaccard:0.15692
 |
Component:357; Jaccard:0.18166
 |
Component:357; Jaccard:0.19256
 |
Component:357; Jaccard:0.17898
 |
Component:357; Jaccard:0.14646
 |
Component:357; Jaccard:0.10526
 |
Correlation:0.25366
  |
Correlation:0.25366
  |
Correlation:0.25366
  |
Correlation:0.25366
  |
Correlation:0.25366
  |
Correlation:0.25366
  |
Correlation:0.25366
  |
Component:129; Jaccard:0.11658
 |
Component:129; Jaccard:0.14513
 |
Component:129; Jaccard:0.16525
 |
Component:129; Jaccard:0.16031
 |
Component:129; Jaccard:0.12139
 |
Component:283; Jaccard:0.087977
 |
Component:283; Jaccard:0.055908
 |
Correlation:0.32117
  |
Correlation:0.32117
  |
Correlation:0.32117
  |
Correlation:0.32117
  |
Correlation:0.32117
  |
Correlation:0.40825
  |
Correlation:0.40825
  |
Component:274; Jaccard:0.10738
 |
Component:100; Jaccard:0.1251
 |
Component:100; Jaccard:0.13083
 |
Component:283; Jaccard:0.12891
 |
Component:283; Jaccard:0.11392
 |
Component:129; Jaccard:0.067055
 |
Component:354; Jaccard:0.033194
 |
Correlation:0.063582
  |
Correlation:0.30106
  |
Correlation:0.30106
  |
Correlation:0.40825
  |
Correlation:0.40825
  |
Correlation:0.32117
  |
Correlation:0.28032
  |
Component:100; Jaccard:0.10586
 |
Component:274; Jaccard:0.11992
 |
Component:283; Jaccard:0.1273
 |
Component:100; Jaccard:0.11914
 |
Component:100; Jaccard:0.086221
 |
Component:274; Jaccard:0.054651
 |
Component:274; Jaccard:0.029085
 |
Correlation:0.30106
  |
Correlation:0.063582
  |
Correlation:0.40825
  |
Correlation:0.30106
  |
Correlation:0.30106
  |
Correlation:0.063582
  |
Correlation:0.063582
  |
Component:378; Jaccard:0.10465
 |
Component:378; Jaccard:0.11882
 |
Component:378; Jaccard:0.12259
 |
Component:378; Jaccard:0.11067
 |
Component:378; Jaccard:0.082993
 |
Component:100; Jaccard:0.048002
 |
Component:362; Jaccard:0.029079
 |
Correlation:0.30506
  |
Correlation:0.30506
  |
Correlation:0.30506
  |
Correlation:0.30506
  |
Correlation:0.30506
  |
Correlation:0.30106
  |
Correlation:0.034721
  |
Component:283; Jaccard:0.099776
 |
Component:283; Jaccard:0.11528
 |
Component:274; Jaccard:0.1216
 |
Component:274; Jaccard:0.10707
 |
Component:274; Jaccard:0.082531
 |
Component:354; Jaccard:0.047589
 |
Component:372; Jaccard:0.024092
 |
Correlation:0.40825
  |
Correlation:0.40825
  |
Correlation:0.063582
  |
Correlation:0.063582
  |
Correlation:0.063582
  |
Correlation:0.28032
  |
Correlation:0.043377
  |
Component:278; Jaccard:0.09721
 |
Component:278; Jaccard:0.10039
 |
Component:278; Jaccard:0.096844
 |
Component:324; Jaccard:0.084957
 |
Component:324; Jaccard:0.067268
 |
Component:378; Jaccard:0.045015
 |
Component:002; Jaccard:0.023907
 |
Correlation:0.069559
  |
Correlation:0.069559
  |
Correlation:0.069559
  |
Correlation:0.40643
  |
Correlation:0.40643
  |
Correlation:0.30506
  |
Correlation:-0.14995
  |
Component:022; Jaccard:0.08089
 |
Component:324; Jaccard:0.087505
 |
Component:324; Jaccard:0.090652
 |
Component:372; Jaccard:0.08362
 |
Component:372; Jaccard:0.065074
 |
Component:362; Jaccard:0.04267
 |
Component:366; Jaccard:0.022698
 |
Correlation:0.29132
  |
Correlation:0.40643
  |
Correlation:0.40643
  |
Correlation:0.043377
  |
Correlation:0.043377
  |
Correlation:0.034721
  |
Correlation:0.066535
  |
Component:324; Jaccard:0.078317
 |
Component:294; Jaccard:0.082665
 |
Component:372; Jaccard:0.089877
 |
Component:278; Jaccard:0.079025
 |
Component:362; Jaccard:0.063388
 |
Component:324; Jaccard:0.041417
 |
Component:129; Jaccard:0.02248
 |
Correlation:0.40643
  |
Correlation:0.28794
  |
Correlation:0.043377
  |
Correlation:0.069559
  |
Correlation:0.034721
  |
Correlation:0.40643
  |
Correlation:0.32117
  |
Component:294; Jaccard:0.075566
 |
Component:372; Jaccard:0.081851
 |
Component:294; Jaccard:0.08438
 |
Component:362; Jaccard:0.076057
 |
Component:354; Jaccard:0.061009
 |
Component:372; Jaccard:0.04052
 |
Component:022; Jaccard:0.021945
 |
Correlation:0.28794
  |
Correlation:0.043377
  |
Correlation:0.28794
  |
Correlation:0.034721
  |
Correlation:0.28032
  |
Correlation:0.043377
  |
Correlation:0.29132
  |
Cope: 03:"FACES-SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
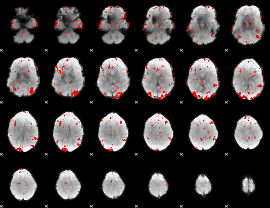 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
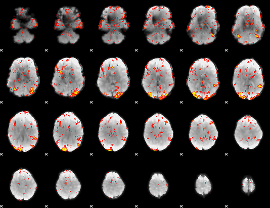 |
GLM Z-score result thresholded by 3
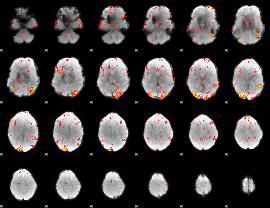 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:239; Jaccard:0.11084
 |
Component:239; Jaccard:0.13288
 |
Component:239; Jaccard:0.15382
 |
Component:030; Jaccard:0.17809
 |
Component:030; Jaccard:0.20122
 |
Component:030; Jaccard:0.21098
 |
Component:030; Jaccard:0.20855
 |
Correlation:0.49468
  |
Correlation:0.49468
  |
Correlation:0.49468
  |
Correlation:0.48754
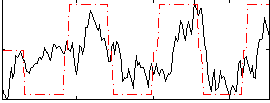 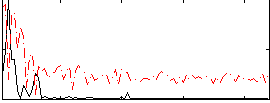 |
Correlation:0.48754
  |
Correlation:0.48754
  |
Correlation:0.48754
  |
Component:343; Jaccard:0.10336
 |
Component:083; Jaccard:0.12636
 |
Component:096; Jaccard:0.1502
 |
Component:096; Jaccard:0.17213
 |
Component:239; Jaccard:0.17497
 |
Component:083; Jaccard:0.17161
 |
Component:083; Jaccard:0.15713
 |
Correlation:0.16479
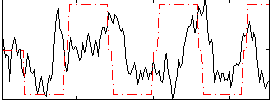  |
Correlation:0.38759
  |
Correlation:0.21643
 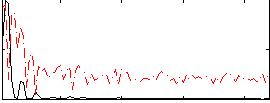 |
Correlation:0.21643
  |
Correlation:0.49468
  |
Correlation:0.38759
  |
Correlation:0.38759
  |
Component:083; Jaccard:0.10232
 |
Component:030; Jaccard:0.12079
 |
Component:030; Jaccard:0.14968
 |
Component:239; Jaccard:0.16884
 |
Component:083; Jaccard:0.17307
 |
Component:239; Jaccard:0.16992
 |
Component:239; Jaccard:0.15454
 |
Correlation:0.38759
  |
Correlation:0.48754
  |
Correlation:0.48754
  |
Correlation:0.49468
  |
Correlation:0.38759
  |
Correlation:0.49468
  |
Correlation:0.49468
  |
Component:040; Jaccard:0.098595
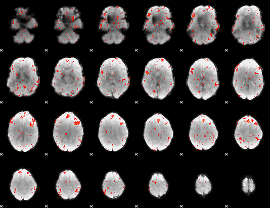 |
Component:096; Jaccard:0.12033
 |
Component:083; Jaccard:0.14665
 |
Component:083; Jaccard:0.16577
 |
Component:096; Jaccard:0.17305
 |
Component:096; Jaccard:0.14738
 |
Component:096; Jaccard:0.10168
 |
Correlation:0.12461
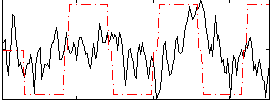  |
Correlation:0.21643
  |
Correlation:0.38759
  |
Correlation:0.38759
  |
Correlation:0.21643
  |
Correlation:0.21643
  |
Correlation:0.21643
  |
Component:030; Jaccard:0.09394
 |
Component:343; Jaccard:0.11315
 |
Component:343; Jaccard:0.11859
 |
Component:138; Jaccard:0.1217
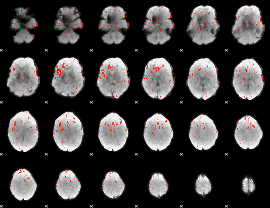 |
Component:138; Jaccard:0.11798
 |
Component:234; Jaccard:0.10454
 |
Component:234; Jaccard:0.092675
 |
Correlation:0.48754
  |
Correlation:0.16479
  |
Correlation:0.16479
  |
Correlation:0.26682
  |
Correlation:0.26682
  |
Correlation:0.29784
  |
Correlation:0.29784
  |
Component:096; Jaccard:0.092691
 |
Component:040; Jaccard:0.10979
 |
Component:040; Jaccard:0.11518
 |
Component:343; Jaccard:0.11274
 |
Component:234; Jaccard:0.10719
 |
Component:138; Jaccard:0.099225
 |
Component:138; Jaccard:0.072972
 |
Correlation:0.21643
  |
Correlation:0.12461
  |
Correlation:0.12461
  |
Correlation:0.16479
  |
Correlation:0.29784
  |
Correlation:0.26682
  |
Correlation:0.26682
  |
Component:297; Jaccard:0.089591
 |
Component:234; Jaccard:0.099989
 |
Component:138; Jaccard:0.11289
 |
Component:040; Jaccard:0.11125
 |
Component:248; Jaccard:0.10296
 |
Component:248; Jaccard:0.085022
 |
Component:248; Jaccard:0.060731
 |
Correlation:0.14934
 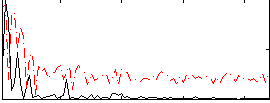 |
Correlation:0.29784
  |
Correlation:0.26682
  |
Correlation:0.12461
  |
Correlation:0.28908
  |
Correlation:0.28908
  |
Correlation:0.28908
  |
Component:234; Jaccard:0.08787
 |
Component:138; Jaccard:0.099508
 |
Component:234; Jaccard:0.10752
 |
Component:234; Jaccard:0.10952
 |
Component:343; Jaccard:0.10031
 |
Component:343; Jaccard:0.082647
 |
Component:343; Jaccard:0.06008
 |
Correlation:0.29784
  |
Correlation:0.26682
  |
Correlation:0.29784
  |
Correlation:0.29784
  |
Correlation:0.16479
  |
Correlation:0.16479
  |
Correlation:0.16479
  |
Component:138; Jaccard:0.083697
 |
Component:297; Jaccard:0.098106
 |
Component:297; Jaccard:0.10098
 |
Component:248; Jaccard:0.10936
 |
Component:040; Jaccard:0.093454
 |
Component:297; Jaccard:0.072041
 |
Component:297; Jaccard:0.055206
 |
Correlation:0.26682
  |
Correlation:0.14934
  |
Correlation:0.14934
  |
Correlation:0.28908
  |
Correlation:0.12461
  |
Correlation:0.14934
  |
Correlation:0.14934
  |
Component:146; Jaccard:0.076457
 |
Component:210; Jaccard:0.086904
 |
Component:248; Jaccard:0.1005
 |
Component:297; Jaccard:0.097168
 |
Component:297; Jaccard:0.087531
 |
Component:040; Jaccard:0.071934
 |
Component:348; Jaccard:0.053677
 |
Correlation:0.20135
  |
Correlation:0.18467
  |
Correlation:0.28908
  |
Correlation:0.14934
  |
Correlation:0.14934
  |
Correlation:0.12461
  |
Correlation:0.12427
  |
Cope: 04:"neg_FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
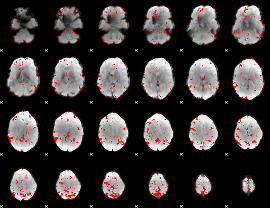 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
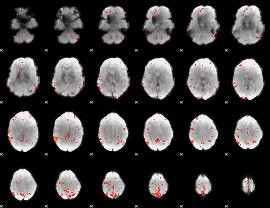 |
GLM Z-score result thresholded by 3.5
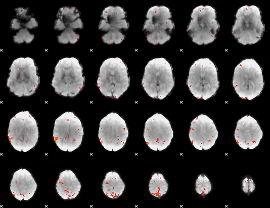 |
GLM Z-score result thresholded by 4
 |
Component:222; Jaccard:0.089345
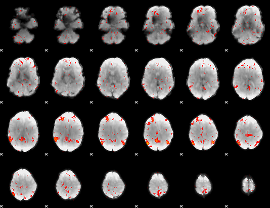 |
Component:222; Jaccard:0.11181
 |
Component:222; Jaccard:0.13465
 |
Component:222; Jaccard:0.14985
 |
Component:222; Jaccard:0.14025
 |
Component:222; Jaccard:0.11109
 |
Component:222; Jaccard:0.077988
 |
Correlation:0.30026
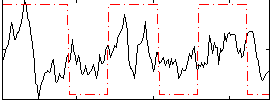  |
Correlation:0.30026
  |
Correlation:0.30026
  |
Correlation:0.30026
  |
Correlation:0.30026
  |
Correlation:0.30026
  |
Correlation:0.30026
  |
Component:189; Jaccard:0.088041
 |
Component:189; Jaccard:0.098201
 |
Component:244; Jaccard:0.11219
 |
Component:244; Jaccard:0.11342
 |
Component:244; Jaccard:0.096066
 |
Component:244; Jaccard:0.064303
 |
Component:244; Jaccard:0.038273
 |
Correlation:0.0043077
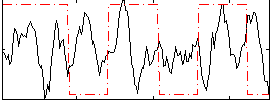  |
Correlation:0.0043077
  |
Correlation:0.17059
  |
Correlation:0.17059
  |
Correlation:0.17059
  |
Correlation:0.17059
  |
Correlation:0.17059
  |
Component:201; Jaccard:0.082699
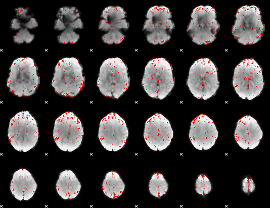 |
Component:244; Jaccard:0.097992
 |
Component:189; Jaccard:0.1046
 |
Component:189; Jaccard:0.10356
 |
Component:189; Jaccard:0.089009
 |
Component:189; Jaccard:0.062379
 |
Component:170; Jaccard:0.036344
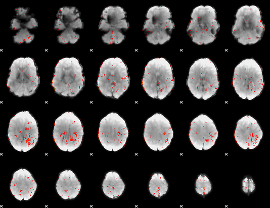 |
Correlation:0.20293
  |
Correlation:0.17059
  |
Correlation:0.0043077
  |
Correlation:0.0043077
  |
Correlation:0.0043077
  |
Correlation:0.0043077
  |
Correlation:0.17577
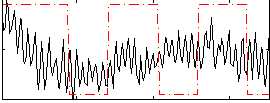  |
Component:244; Jaccard:0.081154
 |
Component:201; Jaccard:0.091014
 |
Component:201; Jaccard:0.09996
 |
Component:201; Jaccard:0.096189
 |
Component:201; Jaccard:0.081679
 |
Component:201; Jaccard:0.059445
 |
Component:141; Jaccard:0.036071
 |
Correlation:0.17059
  |
Correlation:0.20293
  |
Correlation:0.20293
  |
Correlation:0.20293
  |
Correlation:0.20293
  |
Correlation:0.20293
  |
Correlation:0.12686
  |
Component:147; Jaccard:0.072864
 |
Component:147; Jaccard:0.080968
 |
Component:123; Jaccard:0.087976
 |
Component:123; Jaccard:0.090499
 |
Component:123; Jaccard:0.080537
 |
Component:123; Jaccard:0.056367
 |
Component:189; Jaccard:0.035958
 |
Correlation:-0.062921
  |
Correlation:-0.062921
  |
Correlation:0.12175
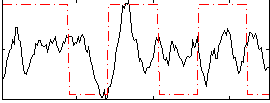  |
Correlation:0.12175
  |
Correlation:0.12175
  |
Correlation:0.12175
  |
Correlation:0.0043077
  |
Component:123; Jaccard:0.070454
 |
Component:123; Jaccard:0.080093
 |
Component:147; Jaccard:0.084218
 |
Component:170; Jaccard:0.085895
 |
Component:170; Jaccard:0.072522
 |
Component:170; Jaccard:0.055835
 |
Component:201; Jaccard:0.034339
 |
Correlation:0.12175
  |
Correlation:0.12175
  |
Correlation:-0.062921
  |
Correlation:0.17577
  |
Correlation:0.17577
  |
Correlation:0.17577
  |
Correlation:0.20293
  |
Component:181; Jaccard:0.069495
 |
Component:181; Jaccard:0.075983
 |
Component:170; Jaccard:0.082645
 |
Component:147; Jaccard:0.081467
 |
Component:141; Jaccard:0.069014
 |
Component:141; Jaccard:0.054741
 |
Component:123; Jaccard:0.032946
 |
Correlation:0.03243
  |
Correlation:0.03243
  |
Correlation:0.17577
  |
Correlation:-0.062921
  |
Correlation:0.12686
  |
Correlation:0.12686
  |
Correlation:0.12175
  |
Component:161; Jaccard:0.067783
 |
Component:170; Jaccard:0.073013
 |
Component:181; Jaccard:0.080636
 |
Component:181; Jaccard:0.078881
 |
Component:147; Jaccard:0.068165
 |
Component:147; Jaccard:0.048453
 |
Component:147; Jaccard:0.029055
 |
Correlation:0.2067
  |
Correlation:0.17577
  |
Correlation:0.03243
  |
Correlation:0.03243
  |
Correlation:-0.062921
  |
Correlation:-0.062921
  |
Correlation:-0.062921
  |
Component:328; Jaccard:0.066888
 |
Component:380; Jaccard:0.07177
 |
Component:012; Jaccard:0.077063
 |
Component:141; Jaccard:0.07651
 |
Component:181; Jaccard:0.064671
 |
Component:012; Jaccard:0.04632
 |
Component:336; Jaccard:0.028621
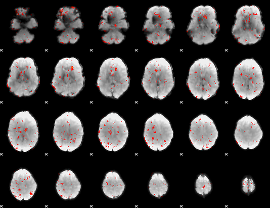 |
Correlation:0.28865
  |
Correlation:0.21076
  |
Correlation:0.15931
  |
Correlation:0.12686
  |
Correlation:0.03243
  |
Correlation:0.15931
  |
Correlation:0.1658
  |
Component:380; Jaccard:0.066847
 |
Component:336; Jaccard:0.07107
 |
Component:336; Jaccard:0.076783
 |
Component:012; Jaccard:0.076335
 |
Component:012; Jaccard:0.063436
 |
Component:181; Jaccard:0.044715
 |
Component:060; Jaccard:0.027842
 |
Correlation:0.21076
  |
Correlation:0.1658
  |
Correlation:0.1658
  |
Correlation:0.15931
  |
Correlation:0.15931
  |
Correlation:0.03243
  |
Correlation:0.14692
  |
Cope: 05:"neg_SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:264; Jaccard:0.084946
 |
Component:210; Jaccard:0.10674
 |
Component:210; Jaccard:0.13894
 |
Component:210; Jaccard:0.17124
 |
Component:210; Jaccard:0.17746
 |
Component:210; Jaccard:0.13743
 |
Component:210; Jaccard:0.068261
 |
Correlation:0.021201
  |
Correlation:0.30435
  |
Correlation:0.30435
  |
Correlation:0.30435
  |
Correlation:0.30435
  |
Correlation:0.30435
  |
Correlation:0.30435
  |
Component:147; Jaccard:0.083655
 |
Component:224; Jaccard:0.1015
 |
Component:224; Jaccard:0.11923
 |
Component:096; Jaccard:0.13489
 |
Component:096; Jaccard:0.13605
 |
Component:096; Jaccard:0.10338
 |
Component:234; Jaccard:0.066525
 |
Correlation:0.23446
  |
Correlation:0.21707
  |
Correlation:0.21707
  |
Correlation:0.25295
  |
Correlation:0.25295
  |
Correlation:0.25295
  |
Correlation:0.29737
  |
Component:224; Jaccard:0.0823
 |
Component:264; Jaccard:0.098219
 |
Component:096; Jaccard:0.11423
 |
Component:224; Jaccard:0.12809
 |
Component:224; Jaccard:0.11174
 |
Component:234; Jaccard:0.085844
 |
Component:239; Jaccard:0.06403
 |
Correlation:0.21707
  |
Correlation:0.021201
  |
Correlation:0.25295
  |
Correlation:0.21707
  |
Correlation:0.21707
  |
Correlation:0.29737
  |
Correlation:0.38641
  |
Component:244; Jaccard:0.07976
 |
Component:244; Jaccard:0.093463
 |
Component:264; Jaccard:0.10575
 |
Component:264; Jaccard:0.10465
 |
Component:239; Jaccard:0.095637
 |
Component:239; Jaccard:0.078895
 |
Component:096; Jaccard:0.045528
 |
Correlation:-0.020368
  |
Correlation:-0.020368
  |
Correlation:0.021201
  |
Correlation:0.021201
  |
Correlation:0.38641
  |
Correlation:0.38641
  |
Correlation:0.25295
  |
Component:210; Jaccard:0.078388
 |
Component:147; Jaccard:0.091381
 |
Component:244; Jaccard:0.101
 |
Component:239; Jaccard:0.10315
 |
Component:234; Jaccard:0.094532
 |
Component:224; Jaccard:0.067577
 |
Component:222; Jaccard:0.040552
 |
Correlation:0.30435
  |
Correlation:0.23446
  |
Correlation:-0.020368
  |
Correlation:0.38641
  |
Correlation:0.29737
  |
Correlation:0.21707
  |
Correlation:-0.13979
  |
Component:239; Jaccard:0.077921
 |
Component:239; Jaccard:0.08983
 |
Component:239; Jaccard:0.09848
 |
Component:234; Jaccard:0.099053
 |
Component:264; Jaccard:0.090104
 |
Component:264; Jaccard:0.062971
 |
Component:293; Jaccard:0.040511
 |
Correlation:0.38641
  |
Correlation:0.38641
  |
Correlation:0.38641
  |
Correlation:0.29737
  |
Correlation:0.021201
  |
Correlation:0.021201
  |
Correlation:0.38308
  |
Component:293; Jaccard:0.077843
 |
Component:096; Jaccard:0.089681
 |
Component:293; Jaccard:0.095257
 |
Component:244; Jaccard:0.096433
 |
Component:244; Jaccard:0.081564
 |
Component:305; Jaccard:0.062606
 |
Component:244; Jaccard:0.035907
 |
Correlation:0.38308
  |
Correlation:0.25295
  |
Correlation:0.38308
  |
Correlation:-0.020368
  |
Correlation:-0.020368
  |
Correlation:0.098438
  |
Correlation:-0.020368
  |
Component:189; Jaccard:0.075692
 |
Component:293; Jaccard:0.087656
 |
Component:234; Jaccard:0.09439
 |
Component:293; Jaccard:0.090559
 |
Component:305; Jaccard:0.079817
 |
Component:359; Jaccard:0.059801
 |
Component:305; Jaccard:0.034921
 |
Correlation:-0.019672
  |
Correlation:0.38308
  |
Correlation:0.29737
  |
Correlation:0.38308
  |
Correlation:0.098438
  |
Correlation:0.18375
  |
Correlation:0.098438
  |
Component:201; Jaccard:0.075374
 |
Component:234; Jaccard:0.083749
 |
Component:147; Jaccard:0.092183
 |
Component:305; Jaccard:0.085816
 |
Component:359; Jaccard:0.079456
 |
Component:293; Jaccard:0.059185
 |
Component:359; Jaccard:0.034755
 |
Correlation:-0.034789
  |
Correlation:0.29737
  |
Correlation:0.23446
  |
Correlation:0.098438
  |
Correlation:0.18375
  |
Correlation:0.38308
  |
Correlation:0.18375
  |
Component:234; Jaccard:0.072448
 |
Component:201; Jaccard:0.079449
 |
Component:222; Jaccard:0.082421
 |
Component:359; Jaccard:0.085279
 |
Component:293; Jaccard:0.077731
 |
Component:244; Jaccard:0.058495
 |
Component:036; Jaccard:0.034483
 |
Correlation:0.29737
  |
Correlation:-0.034789
  |
Correlation:-0.13979
  |
Correlation:0.18375
  |
Correlation:0.38308
  |
Correlation:-0.020368
  |
Correlation:0.34515
  |
Cope: 06:"SHAPES-FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
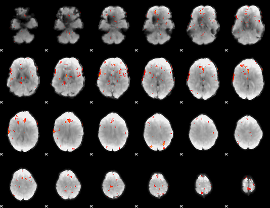 |
Component:328; Jaccard:0.10632
 |
Component:100; Jaccard:0.13659
 |
Component:100; Jaccard:0.18
 |
Component:100; Jaccard:0.22723
 |
Component:100; Jaccard:0.266
 |
Component:100; Jaccard:0.26689
 |
Component:100; Jaccard:0.21841
 |
Correlation:0.41339
  |
Correlation:0.29906
  |
Correlation:0.29906
  |
Correlation:0.29906
  |
Correlation:0.29906
  |
Correlation:0.29906
  |
Correlation:0.29906
  |
Component:100; Jaccard:0.10252
 |
Component:328; Jaccard:0.12299
 |
Component:328; Jaccard:0.13771
 |
Component:129; Jaccard:0.14644
 |
Component:129; Jaccard:0.15695
 |
Component:129; Jaccard:0.15128
 |
Component:129; Jaccard:0.12155
 |
Correlation:0.29906
  |
Correlation:0.41339
  |
Correlation:0.41339
  |
Correlation:0.27379
  |
Correlation:0.27379
  |
Correlation:0.27379
  |
Correlation:0.27379
  |
Component:022; Jaccard:0.10136
 |
Component:022; Jaccard:0.11531
 |
Component:129; Jaccard:0.12678
 |
Component:324; Jaccard:0.14114
 |
Component:324; Jaccard:0.14716
 |
Component:324; Jaccard:0.14005
 |
Component:324; Jaccard:0.11015
 |
Correlation:0.28663
  |
Correlation:0.28663
  |
Correlation:0.27379
  |
Correlation:0.37295
  |
Correlation:0.37295
  |
Correlation:0.37295
  |
Correlation:0.37295
  |
Component:266; Jaccard:0.086304
 |
Component:129; Jaccard:0.10633
 |
Component:022; Jaccard:0.12279
 |
Component:328; Jaccard:0.14003
 |
Component:328; Jaccard:0.13184
 |
Component:051; Jaccard:0.11777
 |
Component:051; Jaccard:0.094039
 |
Correlation:0.20565
  |
Correlation:0.27379
  |
Correlation:0.28663
  |
Correlation:0.41339
  |
Correlation:0.41339
  |
Correlation:0.34815
  |
Correlation:0.34815
  |
Component:129; Jaccard:0.08592
 |
Component:324; Jaccard:0.09938
 |
Component:324; Jaccard:0.12001
 |
Component:022; Jaccard:0.12098
 |
Component:051; Jaccard:0.12409
 |
Component:328; Jaccard:0.10848
 |
Component:328; Jaccard:0.077629
 |
Correlation:0.27379
  |
Correlation:0.37295
  |
Correlation:0.37295
  |
Correlation:0.28663
  |
Correlation:0.34815
  |
Correlation:0.41339
  |
Correlation:0.41339
  |
Component:324; Jaccard:0.080425
 |
Component:051; Jaccard:0.092897
 |
Component:051; Jaccard:0.10774
 |
Component:051; Jaccard:0.11991
 |
Component:022; Jaccard:0.10644
 |
Component:298; Jaccard:0.091348
 |
Component:298; Jaccard:0.076623
 |
Correlation:0.37295
  |
Correlation:0.34815
  |
Correlation:0.34815
  |
Correlation:0.34815
  |
Correlation:0.28663
  |
Correlation:0.28863
  |
Correlation:0.28863
  |
Component:123; Jaccard:0.078834
 |
Component:266; Jaccard:0.089848
 |
Component:304; Jaccard:0.089888
 |
Component:304; Jaccard:0.094751
 |
Component:199; Jaccard:0.098291
 |
Component:199; Jaccard:0.0907
 |
Component:199; Jaccard:0.071609
 |
Correlation:0.15217
  |
Correlation:0.20565
  |
Correlation:0.12526
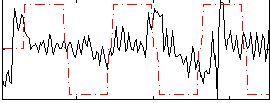 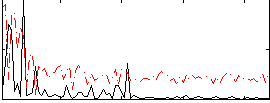 |
Correlation:0.12526
  |
Correlation:0.23386
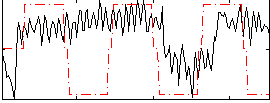  |
Correlation:0.23386
  |
Correlation:0.23386
  |
Component:051; Jaccard:0.078254
 |
Component:123; Jaccard:0.085632
 |
Component:266; Jaccard:0.08921
 |
Component:199; Jaccard:0.093314
 |
Component:304; Jaccard:0.093386
 |
Component:304; Jaccard:0.08536
 |
Component:188; Jaccard:0.071419
 |
Correlation:0.34815
  |
Correlation:0.15217
  |
Correlation:0.20565
  |
Correlation:0.23386
  |
Correlation:0.12526
  |
Correlation:0.12526
  |
Correlation:0.24121
  |
Component:033; Jaccard:0.072078
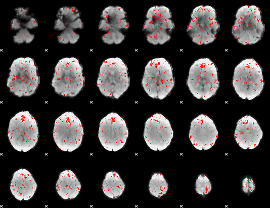 |
Component:378; Jaccard:0.079106
 |
Component:378; Jaccard:0.086842
 |
Component:378; Jaccard:0.090981
 |
Component:298; Jaccard:0.092526
 |
Component:022; Jaccard:0.083501
 |
Component:304; Jaccard:0.065154
 |
Correlation:0.45828
  |
Correlation:0.28539
  |
Correlation:0.28539
  |
Correlation:0.28539
  |
Correlation:0.28863
  |
Correlation:0.28663
  |
Correlation:0.12526
  |
Component:097; Jaccard:0.071926
 |
Component:304; Jaccard:0.079007
 |
Component:123; Jaccard:0.086292
 |
Component:298; Jaccard:0.087362
 |
Component:378; Jaccard:0.088744
 |
Component:188; Jaccard:0.082554
 |
Component:378; Jaccard:0.05876
 |
Correlation:0.20932
  |
Correlation:0.12526
  |
Correlation:0.15217
  |
Correlation:0.28863
  |
Correlation:0.28539
  |
Correlation:0.24121
  |
Correlation:0.28539
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































