Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
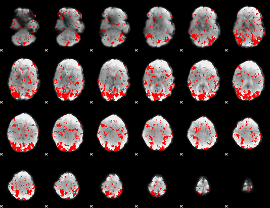 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
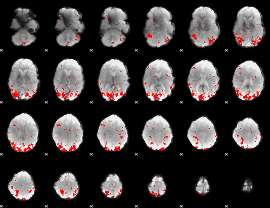 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
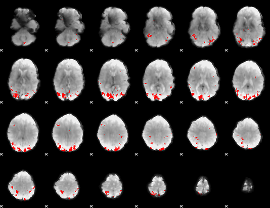 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
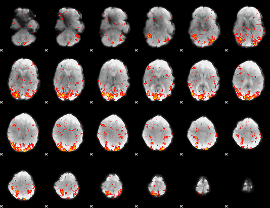 |
GLM Z-score result thresholded by 2.5
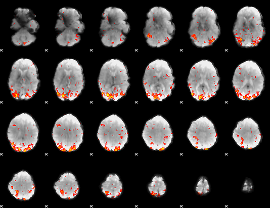 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:053; Jaccard:0.14468
 |
Component:053; Jaccard:0.1885
 |
Component:265; Jaccard:0.23139
 |
Component:265; Jaccard:0.25837
 |
Component:265; Jaccard:0.25399
 |
Component:265; Jaccard:0.23011
 |
Component:265; Jaccard:0.19581
 |
Correlation:0.40497
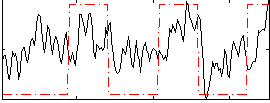 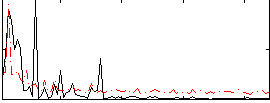 |
Correlation:0.40497
  |
Correlation:-0.081453
  |
Correlation:-0.081453
  |
Correlation:-0.081453
  |
Correlation:-0.081453
  |
Correlation:-0.081453
  |
Component:265; Jaccard:0.13548
 |
Component:265; Jaccard:0.18636
 |
Component:053; Jaccard:0.22271
 |
Component:053; Jaccard:0.23295
 |
Component:053; Jaccard:0.21757
 |
Component:053; Jaccard:0.19004
 |
Component:053; Jaccard:0.15795
 |
Correlation:-0.081453
  |
Correlation:-0.081453
  |
Correlation:0.40497
  |
Correlation:0.40497
  |
Correlation:0.40497
  |
Correlation:0.40497
  |
Correlation:0.40497
  |
Component:221; Jaccard:0.11789
 |
Component:043; Jaccard:0.15318
 |
Component:043; Jaccard:0.18923
 |
Component:043; Jaccard:0.21071
 |
Component:043; Jaccard:0.20583
 |
Component:043; Jaccard:0.18692
 |
Component:043; Jaccard:0.15741
 |
Correlation:0.18799
 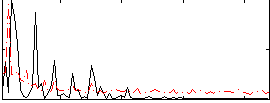 |
Correlation:0.22202
  |
Correlation:0.22202
  |
Correlation:0.22202
  |
Correlation:0.22202
  |
Correlation:0.22202
  |
Correlation:0.22202
  |
Component:006; Jaccard:0.11781
 |
Component:221; Jaccard:0.15052
 |
Component:221; Jaccard:0.17733
 |
Component:221; Jaccard:0.18505
 |
Component:221; Jaccard:0.17312
 |
Component:221; Jaccard:0.14825
 |
Component:145; Jaccard:0.12373
 |
Correlation:-0.043886
  |
Correlation:0.18799
  |
Correlation:0.18799
  |
Correlation:0.18799
  |
Correlation:0.18799
  |
Correlation:0.18799
  |
Correlation:-0.0093769
  |
Component:043; Jaccard:0.11382
 |
Component:384; Jaccard:0.13983
 |
Component:145; Jaccard:0.16257
 |
Component:145; Jaccard:0.17014
 |
Component:145; Jaccard:0.16401
 |
Component:145; Jaccard:0.14702
 |
Component:221; Jaccard:0.12208
 |
Correlation:0.22202
  |
Correlation:-0.18285
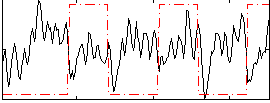  |
Correlation:-0.0093769
  |
Correlation:-0.0093769
  |
Correlation:-0.0093769
  |
Correlation:-0.0093769
  |
Correlation:0.18799
  |
Component:330; Jaccard:0.1122
 |
Component:006; Jaccard:0.13817
 |
Component:384; Jaccard:0.15991
 |
Component:384; Jaccard:0.15911
 |
Component:384; Jaccard:0.14631
 |
Component:248; Jaccard:0.1189
 |
Component:248; Jaccard:0.096011
 |
Correlation:0.2193
  |
Correlation:-0.043886
  |
Correlation:-0.18285
  |
Correlation:-0.18285
  |
Correlation:-0.18285
  |
Correlation:-0.048563
 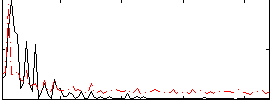 |
Correlation:-0.048563
  |
Component:384; Jaccard:0.11198
 |
Component:145; Jaccard:0.13478
 |
Component:006; Jaccard:0.14938
 |
Component:248; Jaccard:0.15115
 |
Component:248; Jaccard:0.1396
 |
Component:384; Jaccard:0.11785
 |
Component:384; Jaccard:0.094303
 |
Correlation:-0.18285
  |
Correlation:-0.0093769
  |
Correlation:-0.043886
  |
Correlation:-0.048563
  |
Correlation:-0.048563
  |
Correlation:-0.18285
  |
Correlation:-0.18285
  |
Component:368; Jaccard:0.1101
 |
Component:330; Jaccard:0.13144
 |
Component:248; Jaccard:0.14608
 |
Component:006; Jaccard:0.14074
 |
Component:006; Jaccard:0.11632
 |
Component:006; Jaccard:0.089838
 |
Component:006; Jaccard:0.065177
 |
Correlation:-0.03015
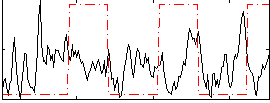 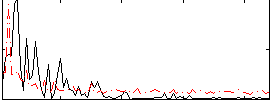 |
Correlation:0.2193
  |
Correlation:-0.048563
  |
Correlation:-0.043886
  |
Correlation:-0.043886
  |
Correlation:-0.043886
  |
Correlation:-0.043886
  |
Component:145; Jaccard:0.10507
 |
Component:248; Jaccard:0.12569
 |
Component:330; Jaccard:0.13521
 |
Component:330; Jaccard:0.12237
 |
Component:330; Jaccard:0.099194
 |
Component:058; Jaccard:0.078942
 |
Component:058; Jaccard:0.063586
 |
Correlation:-0.0093769
  |
Correlation:-0.048563
  |
Correlation:0.2193
  |
Correlation:0.2193
  |
Correlation:0.2193
  |
Correlation:0.49781
  |
Correlation:0.49781
  |
Component:248; Jaccard:0.10029
 |
Component:368; Jaccard:0.12241
 |
Component:368; Jaccard:0.12336
 |
Component:368; Jaccard:0.11311
 |
Component:058; Jaccard:0.09215
 |
Component:330; Jaccard:0.077557
 |
Component:330; Jaccard:0.057149
 |
Correlation:-0.048563
  |
Correlation:-0.03015
  |
Correlation:-0.03015
  |
Correlation:-0.03015
  |
Correlation:0.49781
  |
Correlation:0.2193
  |
Correlation:0.2193
  |
Cope: 02:"SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
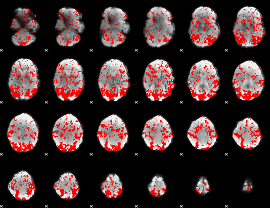 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
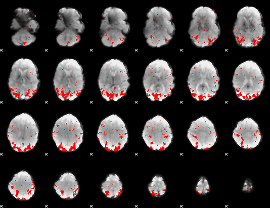 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
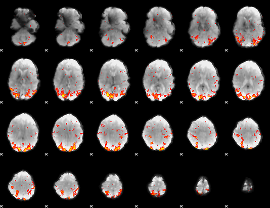 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:265; Jaccard:0.14441
 |
Component:265; Jaccard:0.20327
 |
Component:265; Jaccard:0.26694
 |
Component:265; Jaccard:0.30635
 |
Component:265; Jaccard:0.31544
 |
Component:265; Jaccard:0.29557
 |
Component:265; Jaccard:0.25581
 |
Correlation:0.2606
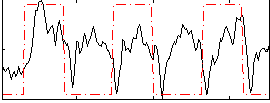  |
Correlation:0.2606
  |
Correlation:0.2606
  |
Correlation:0.2606
  |
Correlation:0.2606
  |
Correlation:0.2606
  |
Correlation:0.2606
  |
Component:006; Jaccard:0.13607
 |
Component:006; Jaccard:0.16714
 |
Component:384; Jaccard:0.19358
 |
Component:384; Jaccard:0.20701
 |
Component:384; Jaccard:0.20288
 |
Component:384; Jaccard:0.17925
 |
Component:043; Jaccard:0.15545
 |
Correlation:0.14686
  |
Correlation:0.14686
  |
Correlation:0.23126
  |
Correlation:0.23126
  |
Correlation:0.23126
  |
Correlation:0.23126
  |
Correlation:-0.048673
  |
Component:384; Jaccard:0.12674
 |
Component:384; Jaccard:0.16452
 |
Component:006; Jaccard:0.19111
 |
Component:043; Jaccard:0.19518
 |
Component:043; Jaccard:0.19367
 |
Component:145; Jaccard:0.17666
 |
Component:145; Jaccard:0.15443
 |
Correlation:0.23126
  |
Correlation:0.23126
  |
Correlation:0.14686
  |
Correlation:-0.048673
  |
Correlation:-0.048673
  |
Correlation:0.17225
  |
Correlation:0.17225
  |
Component:368; Jaccard:0.12622
 |
Component:368; Jaccard:0.14914
 |
Component:145; Jaccard:0.17721
 |
Component:145; Jaccard:0.19258
 |
Component:145; Jaccard:0.18996
 |
Component:043; Jaccard:0.17648
 |
Component:384; Jaccard:0.14977
 |
Correlation:-0.0014883
  |
Correlation:-0.0014883
  |
Correlation:0.17225
  |
Correlation:0.17225
  |
Correlation:0.17225
  |
Correlation:-0.048673
  |
Correlation:0.23126
  |
Component:248; Jaccard:0.11446
 |
Component:145; Jaccard:0.14832
 |
Component:043; Jaccard:0.17694
 |
Component:006; Jaccard:0.19154
 |
Component:248; Jaccard:0.18035
 |
Component:248; Jaccard:0.16411
 |
Component:248; Jaccard:0.14021
 |
Correlation:0.34173
  |
Correlation:0.17225
  |
Correlation:-0.048673
  |
Correlation:0.14686
  |
Correlation:0.34173
  |
Correlation:0.34173
  |
Correlation:0.34173
  |
Component:145; Jaccard:0.11276
 |
Component:248; Jaccard:0.14555
 |
Component:248; Jaccard:0.17557
 |
Component:248; Jaccard:0.1859
 |
Component:006; Jaccard:0.17451
 |
Component:006; Jaccard:0.14283
 |
Component:006; Jaccard:0.11506
 |
Correlation:0.17225
  |
Correlation:0.34173
  |
Correlation:0.34173
  |
Correlation:0.34173
  |
Correlation:0.14686
  |
Correlation:0.14686
  |
Correlation:0.14686
  |
Component:043; Jaccard:0.10851
 |
Component:043; Jaccard:0.14306
 |
Component:368; Jaccard:0.15674
 |
Component:053; Jaccard:0.15
 |
Component:053; Jaccard:0.14703
 |
Component:221; Jaccard:0.12749
 |
Component:221; Jaccard:0.10856
 |
Correlation:-0.048673
  |
Correlation:-0.048673
  |
Correlation:-0.0014883
  |
Correlation:-0.33207
  |
Correlation:-0.33207
  |
Correlation:-0.08303
  |
Correlation:-0.08303
  |
Component:053; Jaccard:0.10519
 |
Component:053; Jaccard:0.12757
 |
Component:053; Jaccard:0.14618
 |
Component:368; Jaccard:0.14773
 |
Component:221; Jaccard:0.13922
 |
Component:053; Jaccard:0.12663
 |
Component:053; Jaccard:0.10493
 |
Correlation:-0.33207
  |
Correlation:-0.33207
  |
Correlation:-0.33207
  |
Correlation:-0.0014883
  |
Correlation:-0.08303
  |
Correlation:-0.33207
  |
Correlation:-0.33207
  |
Component:221; Jaccard:0.101
 |
Component:221; Jaccard:0.12353
 |
Component:221; Jaccard:0.14165
 |
Component:221; Jaccard:0.14709
 |
Component:368; Jaccard:0.12646
 |
Component:368; Jaccard:0.10249
 |
Component:368; Jaccard:0.080305
 |
Correlation:-0.08303
  |
Correlation:-0.08303
  |
Correlation:-0.08303
  |
Correlation:-0.08303
  |
Correlation:-0.0014883
  |
Correlation:-0.0014883
  |
Correlation:-0.0014883
  |
Component:385; Jaccard:0.098099
 |
Component:385; Jaccard:0.10847
 |
Component:385; Jaccard:0.10836
 |
Component:361; Jaccard:0.10403
 |
Component:370; Jaccard:0.094171
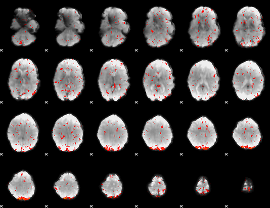 |
Component:370; Jaccard:0.083566
 |
Component:370; Jaccard:0.070538
 |
Correlation:0.081177
  |
Correlation:0.081177
  |
Correlation:0.081177
  |
Correlation:0.081293
  |
Correlation:0.18299
  |
Correlation:0.18299
  |
Correlation:0.18299
  |
Cope: 03:"FACES-SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
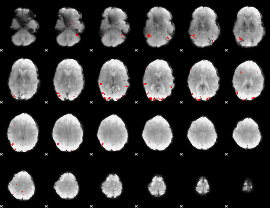 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
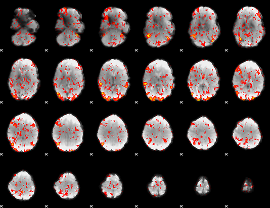 |
GLM Z-score result thresholded by 2
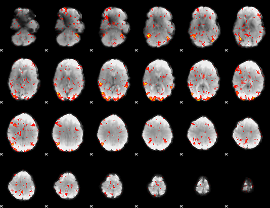 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
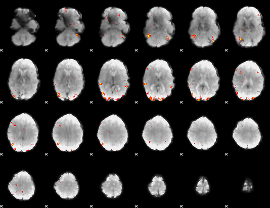 |
GLM Z-score result thresholded by 4
 |
Component:058; Jaccard:0.12885
 |
Component:058; Jaccard:0.17312
 |
Component:058; Jaccard:0.21193
 |
Component:058; Jaccard:0.22198
 |
Component:058; Jaccard:0.19361
 |
Component:058; Jaccard:0.15319
 |
Component:058; Jaccard:0.11666
 |
Correlation:0.52616
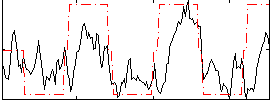 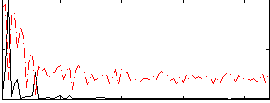 |
Correlation:0.52616
  |
Correlation:0.52616
  |
Correlation:0.52616
  |
Correlation:0.52616
  |
Correlation:0.52616
  |
Correlation:0.52616
  |
Component:053; Jaccard:0.10949
 |
Component:053; Jaccard:0.13274
 |
Component:053; Jaccard:0.15066
 |
Component:053; Jaccard:0.14739
 |
Component:053; Jaccard:0.12807
 |
Component:053; Jaccard:0.1024
 |
Component:053; Jaccard:0.076733
 |
Correlation:0.39978
  |
Correlation:0.39978
  |
Correlation:0.39978
  |
Correlation:0.39978
  |
Correlation:0.39978
  |
Correlation:0.39978
  |
Correlation:0.39978
  |
Component:231; Jaccard:0.10685
 |
Component:231; Jaccard:0.12136
 |
Component:231; Jaccard:0.12349
 |
Component:362; Jaccard:0.11669
 |
Component:362; Jaccard:0.10082
 |
Component:362; Jaccard:0.080176
 |
Component:362; Jaccard:0.059221
 |
Correlation:0.2059
  |
Correlation:0.2059
  |
Correlation:0.2059
  |
Correlation:0.46833
  |
Correlation:0.46833
  |
Correlation:0.46833
  |
Correlation:0.46833
  |
Component:292; Jaccard:0.093324
 |
Component:362; Jaccard:0.11052
 |
Component:362; Jaccard:0.12284
 |
Component:292; Jaccard:0.1127
 |
Component:292; Jaccard:0.093076
 |
Component:292; Jaccard:0.070556
 |
Component:292; Jaccard:0.050324
 |
Correlation:0.46856
  |
Correlation:0.46833
  |
Correlation:0.46833
  |
Correlation:0.46856
  |
Correlation:0.46856
  |
Correlation:0.46856
  |
Correlation:0.46856
  |
Component:362; Jaccard:0.093305
 |
Component:292; Jaccard:0.10879
 |
Component:292; Jaccard:0.11739
 |
Component:231; Jaccard:0.1059
 |
Component:231; Jaccard:0.073229
 |
Component:231; Jaccard:0.049642
 |
Component:390; Jaccard:0.038541
 |
Correlation:0.46833
  |
Correlation:0.46856
  |
Correlation:0.46856
  |
Correlation:0.2059
  |
Correlation:0.2059
  |
Correlation:0.2059
  |
Correlation:0.14213
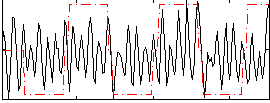  |
Component:330; Jaccard:0.091114
 |
Component:330; Jaccard:0.10122
 |
Component:330; Jaccard:0.10227
 |
Component:330; Jaccard:0.087377
 |
Component:358; Jaccard:0.064936
 |
Component:221; Jaccard:0.047397
 |
Component:149; Jaccard:0.034817
 |
Correlation:0.21012
 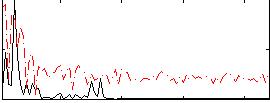 |
Correlation:0.21012
  |
Correlation:0.21012
  |
Correlation:0.21012
  |
Correlation:0.34078
  |
Correlation:0.14701
  |
Correlation:0.47029
  |
Component:358; Jaccard:0.086771
 |
Component:358; Jaccard:0.098661
 |
Component:358; Jaccard:0.099356
 |
Component:358; Jaccard:0.087319
 |
Component:330; Jaccard:0.063428
 |
Component:390; Jaccard:0.04734
 |
Component:221; Jaccard:0.033852
 |
Correlation:0.34078
  |
Correlation:0.34078
  |
Correlation:0.34078
  |
Correlation:0.34078
  |
Correlation:0.21012
  |
Correlation:0.14213
  |
Correlation:0.14701
  |
Component:163; Jaccard:0.08277
 |
Component:163; Jaccard:0.082775
 |
Component:221; Jaccard:0.079094
 |
Component:221; Jaccard:0.074762
 |
Component:221; Jaccard:0.061206
 |
Component:149; Jaccard:0.04693
 |
Component:231; Jaccard:0.031921
 |
Correlation:0.065388
  |
Correlation:0.065388
  |
Correlation:0.14701
  |
Correlation:0.14701
  |
Correlation:0.14701
  |
Correlation:0.47029
  |
Correlation:0.2059
  |
Component:238; Jaccard:0.075088
 |
Component:238; Jaccard:0.079279
 |
Component:238; Jaccard:0.075177
 |
Component:149; Jaccard:0.063745
 |
Component:149; Jaccard:0.058517
 |
Component:358; Jaccard:0.045752
 |
Component:358; Jaccard:0.031208
 |
Correlation:0.34101
  |
Correlation:0.34101
  |
Correlation:0.34101
  |
Correlation:0.47029
  |
Correlation:0.47029
  |
Correlation:0.34078
  |
Correlation:0.34078
  |
Component:307; Jaccard:0.074972
 |
Component:221; Jaccard:0.076819
 |
Component:163; Jaccard:0.073603
 |
Component:238; Jaccard:0.058554
 |
Component:390; Jaccard:0.053571
 |
Component:330; Jaccard:0.04558
 |
Component:330; Jaccard:0.030252
 |
Correlation:0.20114
  |
Correlation:0.14701
  |
Correlation:0.065388
  |
Correlation:0.34101
  |
Correlation:0.14213
  |
Correlation:0.21012
  |
Correlation:0.21012
  |
Cope: 04:"neg_FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
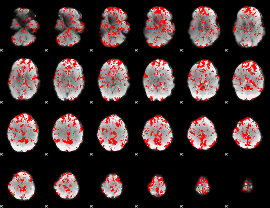 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
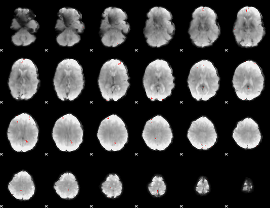 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
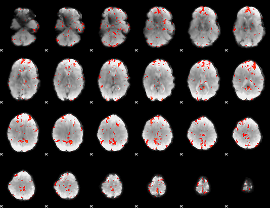 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
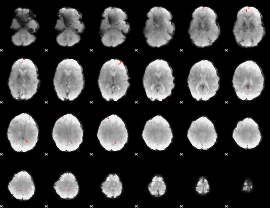 |
Component:178; Jaccard:0.1184
 |
Component:178; Jaccard:0.15329
 |
Component:178; Jaccard:0.17558
 |
Component:178; Jaccard:0.15655
 |
Component:178; Jaccard:0.10791
 |
Component:178; Jaccard:0.054583
 |
Component:178; Jaccard:0.022335
 |
Correlation:0.48396
  |
Correlation:0.48396
  |
Correlation:0.48396
  |
Correlation:0.48396
  |
Correlation:0.48396
  |
Correlation:0.48396
  |
Correlation:0.48396
  |
Component:276; Jaccard:0.091534
 |
Component:276; Jaccard:0.097684
 |
Component:276; Jaccard:0.090927
 |
Component:158; Jaccard:0.066918
 |
Component:372; Jaccard:0.043815
 |
Component:372; Jaccard:0.022544
 |
Component:372; Jaccard:0.011319
 |
Correlation:0.10788
  |
Correlation:0.10788
  |
Correlation:0.10788
  |
Correlation:0.1395
  |
Correlation:0.32857
  |
Correlation:0.32857
  |
Correlation:0.32857
  |
Component:261; Jaccard:0.080215
 |
Component:035; Jaccard:0.079687
 |
Component:158; Jaccard:0.08287
 |
Component:372; Jaccard:0.066829
 |
Component:158; Jaccard:0.042466
 |
Component:158; Jaccard:0.019625
 |
Component:158; Jaccard:0.007972
 |
Correlation:0.17145
  |
Correlation:0.23847
  |
Correlation:0.1395
  |
Correlation:0.32857
  |
Correlation:0.1395
  |
Correlation:0.1395
  |
Correlation:0.1395
  |
Component:035; Jaccard:0.077916
 |
Component:261; Jaccard:0.078813
 |
Component:372; Jaccard:0.080766
 |
Component:276; Jaccard:0.065636
 |
Component:237; Jaccard:0.037092
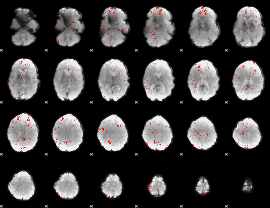 |
Component:276; Jaccard:0.019161
 |
Component:005; Jaccard:0.007509
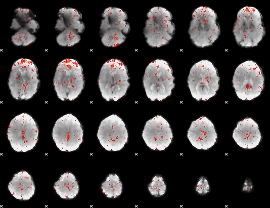 |
Correlation:0.23847
  |
Correlation:0.17145
  |
Correlation:0.32857
  |
Correlation:0.10788
  |
Correlation:0.11845
  |
Correlation:0.10788
  |
Correlation:0.14122
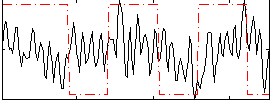 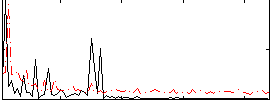 |
Component:055; Jaccard:0.07528
 |
Component:158; Jaccard:0.078667
 |
Component:237; Jaccard:0.071606
 |
Component:237; Jaccard:0.06015
 |
Component:276; Jaccard:0.036957
 |
Component:261; Jaccard:0.017252
 |
Component:205; Jaccard:0.007317
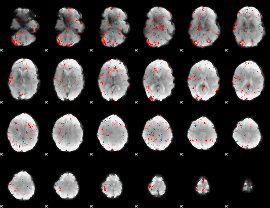 |
Correlation:0.030668
  |
Correlation:0.1395
  |
Correlation:0.11845
  |
Correlation:0.11845
  |
Correlation:0.10788
  |
Correlation:0.17145
  |
Correlation:-0.023007
  |
Component:366; Jaccard:0.068718
 |
Component:372; Jaccard:0.0755
 |
Component:366; Jaccard:0.069107
 |
Component:366; Jaccard:0.05113
 |
Component:261; Jaccard:0.031682
 |
Component:005; Jaccard:0.016442
 |
Component:276; Jaccard:0.007029
 |
Correlation:0.27817
  |
Correlation:0.32857
  |
Correlation:0.27817
  |
Correlation:0.27817
  |
Correlation:0.17145
  |
Correlation:0.14122
  |
Correlation:0.10788
  |
Component:328; Jaccard:0.06776
 |
Component:366; Jaccard:0.073993
 |
Component:261; Jaccard:0.067507
 |
Component:310; Jaccard:0.050341
 |
Component:310; Jaccard:0.031374
 |
Component:237; Jaccard:0.015991
 |
Component:261; Jaccard:0.006885
 |
Correlation:0.28661
  |
Correlation:0.27817
  |
Correlation:0.17145
  |
Correlation:0.10877
  |
Correlation:0.10877
  |
Correlation:0.11845
  |
Correlation:0.17145
  |
Component:158; Jaccard:0.067711
 |
Component:328; Jaccard:0.072767
 |
Component:005; Jaccard:0.066954
 |
Component:005; Jaccard:0.049418
 |
Component:005; Jaccard:0.030408
 |
Component:254; Jaccard:0.014708
 |
Component:254; Jaccard:0.006115
 |
Correlation:0.1395
  |
Correlation:0.28661
  |
Correlation:0.14122
  |
Correlation:0.14122
  |
Correlation:0.14122
  |
Correlation:0.21923
  |
Correlation:0.21923
  |
Component:381; Jaccard:0.066965
 |
Component:381; Jaccard:0.072456
 |
Component:035; Jaccard:0.066882
 |
Component:111; Jaccard:0.047614
 |
Component:035; Jaccard:0.028382
 |
Component:035; Jaccard:0.013619
 |
Component:035; Jaccard:0.006038
 |
Correlation:0.22907
  |
Correlation:0.22907
  |
Correlation:0.23847
  |
Correlation:0.038182
  |
Correlation:0.23847
  |
Correlation:0.23847
  |
Correlation:0.23847
  |
Component:223; Jaccard:0.066812
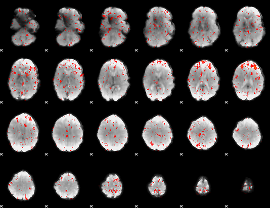 |
Component:310; Jaccard:0.069022
 |
Component:381; Jaccard:0.065785
 |
Component:035; Jaccard:0.047502
 |
Component:366; Jaccard:0.028244
 |
Component:366; Jaccard:0.01358
 |
Component:366; Jaccard:0.005943
 |
Correlation:0.13054
  |
Correlation:0.10877
  |
Correlation:0.22907
  |
Correlation:0.23847
  |
Correlation:0.27817
  |
Correlation:0.27817
  |
Correlation:0.27817
  |
Cope: 05:"neg_SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:292; Jaccard:0.099585
 |
Component:292; Jaccard:0.11761
 |
Component:292; Jaccard:0.12587
 |
Component:292; Jaccard:0.11255
 |
Component:292; Jaccard:0.089997
 |
Component:292; Jaccard:0.06721
 |
Component:292; Jaccard:0.050339
 |
Correlation:0.43696
  |
Correlation:0.43696
  |
Correlation:0.43696
  |
Correlation:0.43696
  |
Correlation:0.43696
  |
Correlation:0.43696
  |
Correlation:0.43696
  |
Component:178; Jaccard:0.099048
 |
Component:178; Jaccard:0.11698
 |
Component:178; Jaccard:0.12315
 |
Component:178; Jaccard:0.10543
 |
Component:149; Jaccard:0.07577
 |
Component:149; Jaccard:0.060192
 |
Component:149; Jaccard:0.044108
 |
Correlation:-0.20997
  |
Correlation:-0.20997
  |
Correlation:-0.20997
  |
Correlation:-0.20997
  |
Correlation:0.49713
  |
Correlation:0.49713
  |
Correlation:0.49713
  |
Component:276; Jaccard:0.093469
 |
Component:276; Jaccard:0.10075
 |
Component:231; Jaccard:0.10129
 |
Component:158; Jaccard:0.08897
 |
Component:178; Jaccard:0.066978
 |
Component:362; Jaccard:0.046508
 |
Component:362; Jaccard:0.035299
 |
Correlation:0.11229
  |
Correlation:0.11229
  |
Correlation:0.25225
  |
Correlation:0.038322
  |
Correlation:-0.20997
  |
Correlation:0.53403
  |
Correlation:0.53403
  |
Component:231; Jaccard:0.090768
 |
Component:231; Jaccard:0.10034
 |
Component:158; Jaccard:0.098986
 |
Component:149; Jaccard:0.081292
 |
Component:158; Jaccard:0.06295
 |
Component:158; Jaccard:0.033103
 |
Component:158; Jaccard:0.015178
 |
Correlation:0.25225
  |
Correlation:0.25225
  |
Correlation:0.038322
  |
Correlation:0.49713
  |
Correlation:0.038322
  |
Correlation:0.038322
  |
Correlation:0.038322
  |
Component:055; Jaccard:0.089352
 |
Component:249; Jaccard:0.091101
 |
Component:276; Jaccard:0.094328
 |
Component:231; Jaccard:0.079544
 |
Component:362; Jaccard:0.060688
 |
Component:178; Jaccard:0.030608
 |
Component:231; Jaccard:0.013697
 |
Correlation:0.12249
  |
Correlation:0.16022
  |
Correlation:0.11229
  |
Correlation:0.25225
  |
Correlation:0.53403
  |
Correlation:-0.20997
  |
Correlation:0.25225
  |
Component:249; Jaccard:0.084527
 |
Component:158; Jaccard:0.089794
 |
Component:249; Jaccard:0.089779
 |
Component:362; Jaccard:0.074944
 |
Component:231; Jaccard:0.05
 |
Component:231; Jaccard:0.025572
 |
Component:205; Jaccard:0.012804
 |
Correlation:0.16022
  |
Correlation:0.038322
  |
Correlation:0.16022
  |
Correlation:0.53403
  |
Correlation:0.25225
  |
Correlation:0.25225
  |
Correlation:0.20348
  |
Component:158; Jaccard:0.075051
 |
Component:055; Jaccard:0.083611
 |
Component:362; Jaccard:0.07875
 |
Component:276; Jaccard:0.072523
 |
Component:276; Jaccard:0.042198
 |
Component:276; Jaccard:0.019901
 |
Component:178; Jaccard:0.010356
 |
Correlation:0.038322
  |
Correlation:0.12249
  |
Correlation:0.53403
  |
Correlation:0.11229
  |
Correlation:0.11229
  |
Correlation:0.11229
  |
Correlation:-0.20997
  |
Component:307; Jaccard:0.072017
 |
Component:111; Jaccard:0.079294
 |
Component:149; Jaccard:0.078504
 |
Component:249; Jaccard:0.069388
 |
Component:249; Jaccard:0.040159
 |
Component:249; Jaccard:0.019667
 |
Component:350; Jaccard:0.009101
 |
Correlation:0.29422
  |
Correlation:-0.045941
  |
Correlation:0.49713
  |
Correlation:0.16022
  |
Correlation:0.16022
  |
Correlation:0.16022
  |
Correlation:0.36266
  |
Component:111; Jaccard:0.071643
 |
Component:193; Jaccard:0.074317
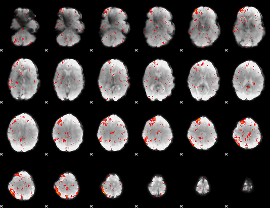 |
Component:111; Jaccard:0.071199
 |
Component:310; Jaccard:0.055669
 |
Component:237; Jaccard:0.034108
 |
Component:205; Jaccard:0.017237
 |
Component:276; Jaccard:0.009071
 |
Correlation:-0.045941
  |
Correlation:-0.023508
  |
Correlation:-0.045941
  |
Correlation:-0.080525
  |
Correlation:-0.088781
  |
Correlation:0.20348
  |
Correlation:0.11229
  |
Component:193; Jaccard:0.070352
 |
Component:362; Jaccard:0.073978
 |
Component:310; Jaccard:0.069386
 |
Component:111; Jaccard:0.052833
 |
Component:310; Jaccard:0.033655
 |
Component:310; Jaccard:0.016265
 |
Component:241; Jaccard:0.008643
 |
Correlation:-0.023508
  |
Correlation:0.53403
  |
Correlation:-0.080525
  |
Correlation:-0.045941
  |
Correlation:-0.080525
  |
Correlation:-0.080525
  |
Correlation:0.1853
  |
Cope: 06:"SHAPES-FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
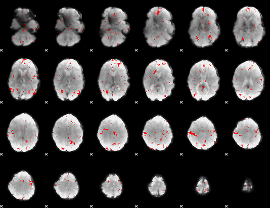 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
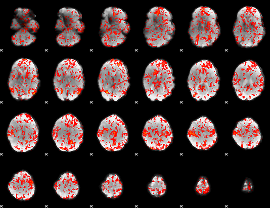 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
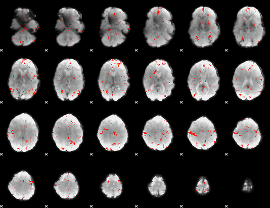 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
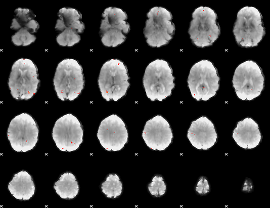 |
Component:301; Jaccard:0.085639
 |
Component:301; Jaccard:0.090101
 |
Component:211; Jaccard:0.084239
 |
Component:211; Jaccard:0.071429
 |
Component:211; Jaccard:0.047595
 |
Component:211; Jaccard:0.022934
 |
Component:370; Jaccard:0.010548
 |
Correlation:0.055019
 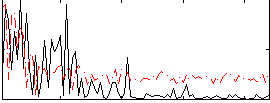 |
Correlation:0.055019
  |
Correlation:0.28925
  |
Correlation:0.28925
  |
Correlation:0.28925
  |
Correlation:0.28925
  |
Correlation:0.20431
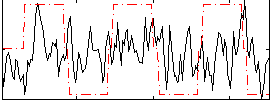  |
Component:279; Jaccard:0.078312
 |
Component:072; Jaccard:0.079957
 |
Component:301; Jaccard:0.079592
 |
Component:072; Jaccard:0.062618
 |
Component:007; Jaccard:0.038505
 |
Component:370; Jaccard:0.020263
 |
Component:384; Jaccard:0.009231
 |
Correlation:0.039724
  |
Correlation:0.42229
  |
Correlation:0.055019
  |
Correlation:0.42229
  |
Correlation:0.35862
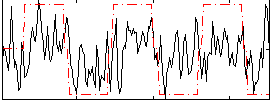  |
Correlation:0.20431
  |
Correlation:0.22462
  |
Component:072; Jaccard:0.073772
 |
Component:279; Jaccard:0.076793
 |
Component:072; Jaccard:0.078881
 |
Component:007; Jaccard:0.059492
 |
Component:072; Jaccard:0.03723
 |
Component:007; Jaccard:0.018554
 |
Component:392; Jaccard:0.008381
 |
Correlation:0.42229
  |
Correlation:0.039724
  |
Correlation:0.42229
  |
Correlation:0.35862
  |
Correlation:0.42229
  |
Correlation:0.35862
  |
Correlation:0.050425
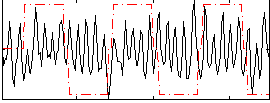 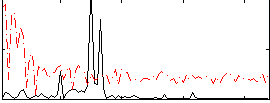 |
Component:095; Jaccard:0.071963
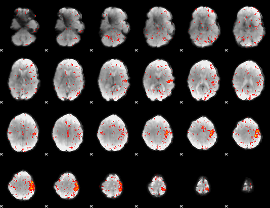 |
Component:211; Jaccard:0.076373
 |
Component:007; Jaccard:0.073689
 |
Component:189; Jaccard:0.057082
 |
Component:370; Jaccard:0.0333
 |
Component:384; Jaccard:0.018454
 |
Component:007; Jaccard:0.008188
 |
Correlation:0.13644
  |
Correlation:0.28925
  |
Correlation:0.35862
  |
Correlation:0.4239
  |
Correlation:0.20431
  |
Correlation:0.22462
  |
Correlation:0.35862
  |
Component:047; Jaccard:0.069226
 |
Component:095; Jaccard:0.071281
 |
Component:189; Jaccard:0.068736
 |
Component:301; Jaccard:0.053317
 |
Component:384; Jaccard:0.033204
 |
Component:072; Jaccard:0.017665
 |
Component:248; Jaccard:0.008085
 |
Correlation:0.32163
  |
Correlation:0.13644
  |
Correlation:0.4239
  |
Correlation:0.055019
  |
Correlation:0.22462
  |
Correlation:0.42229
  |
Correlation:0.2117
  |
Component:320; Jaccard:0.06751
 |
Component:007; Jaccard:0.071218
 |
Component:384; Jaccard:0.062167
 |
Component:384; Jaccard:0.051104
 |
Component:189; Jaccard:0.03168
 |
Component:178; Jaccard:0.014616
 |
Component:211; Jaccard:0.008029
 |
Correlation:0.0076941
  |
Correlation:0.35862
  |
Correlation:0.22462
  |
Correlation:0.22462
  |
Correlation:0.4239
  |
Correlation:0.37639
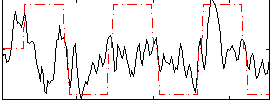  |
Correlation:0.28925
  |
Component:369; Jaccard:0.066674
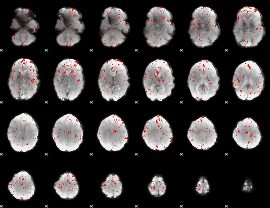 |
Component:189; Jaccard:0.070648
 |
Component:095; Jaccard:0.061122
 |
Component:254; Jaccard:0.047192
 |
Component:301; Jaccard:0.030585
 |
Component:248; Jaccard:0.014591
 |
Component:072; Jaccard:0.007994
 |
Correlation:-0.0031723
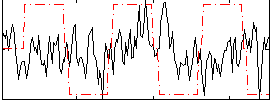  |
Correlation:0.4239
  |
Correlation:0.13644
  |
Correlation:0.12521
  |
Correlation:0.055019
  |
Correlation:0.2117
  |
Correlation:0.42229
  |
Component:348; Jaccard:0.066335
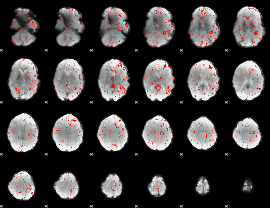 |
Component:381; Jaccard:0.068729
 |
Component:381; Jaccard:0.060604
 |
Component:370; Jaccard:0.046189
 |
Component:381; Jaccard:0.02878
 |
Component:381; Jaccard:0.013969
 |
Component:178; Jaccard:0.007167
 |
Correlation:0.18408
  |
Correlation:0.16614
  |
Correlation:0.16614
  |
Correlation:0.20431
  |
Correlation:0.16614
  |
Correlation:0.16614
  |
Correlation:0.37639
  |
Component:381; Jaccard:0.065888
 |
Component:047; Jaccard:0.0682
 |
Component:254; Jaccard:0.059589
 |
Component:381; Jaccard:0.045884
 |
Component:254; Jaccard:0.028436
 |
Component:392; Jaccard:0.013678
 |
Component:267; Jaccard:0.00679
 |
Correlation:0.16614
  |
Correlation:0.32163
  |
Correlation:0.12521
  |
Correlation:0.16614
  |
Correlation:0.12521
  |
Correlation:0.050425
  |
Correlation:0.091269
  |
Component:261; Jaccard:0.065007
 |
Component:178; Jaccard:0.066868
 |
Component:370; Jaccard:0.058744
 |
Component:178; Jaccard:0.043228
 |
Component:178; Jaccard:0.026881
 |
Component:254; Jaccard:0.013429
 |
Component:047; Jaccard:0.006687
 |
Correlation:0.13999
  |
Correlation:0.37639
  |
Correlation:0.20431
  |
Correlation:0.37639
  |
Correlation:0.37639
  |
Correlation:0.12521
  |
Correlation:0.32163
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































