Task name: RELATIONAL, TR=0.72, totally 232 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"MATCH" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
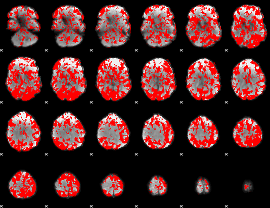 |
GLM binary result thresholded by 1.5
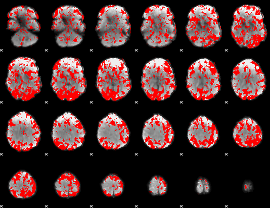 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
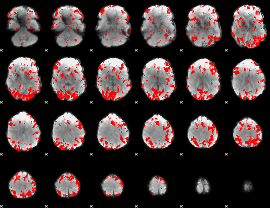 |
GLM binary result thresholded by 3
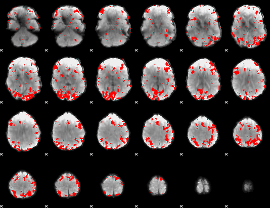 |
GLM binary result thresholded by 3.5
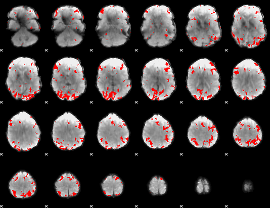 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
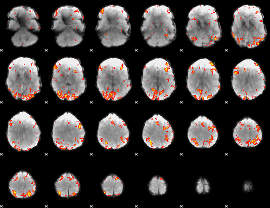 |
GLM Z-score result thresholded by 4
 |
Component:370; Jaccard:0.11204
 |
Component:370; Jaccard:0.14313
 |
Component:370; Jaccard:0.17898
 |
Component:370; Jaccard:0.21471
 |
Component:370; Jaccard:0.24016
 |
Component:370; Jaccard:0.25031
 |
Component:370; Jaccard:0.23501
 |
Correlation:0.21961
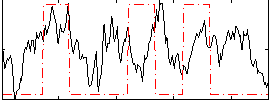 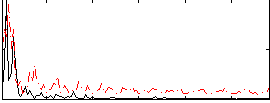 |
Correlation:0.21961
  |
Correlation:0.21961
  |
Correlation:0.21961
  |
Correlation:0.21961
  |
Correlation:0.21961
  |
Correlation:0.21961
  |
Component:048; Jaccard:0.10677
 |
Component:048; Jaccard:0.13375
 |
Component:048; Jaccard:0.16481
 |
Component:317; Jaccard:0.19506
 |
Component:317; Jaccard:0.22187
 |
Component:317; Jaccard:0.23318
 |
Component:317; Jaccard:0.21985
 |
Correlation:0.093265
  |
Correlation:0.093265
  |
Correlation:0.093265
  |
Correlation:0.08457
  |
Correlation:0.08457
  |
Correlation:0.08457
  |
Correlation:0.08457
  |
Component:317; Jaccard:0.098998
 |
Component:317; Jaccard:0.12648
 |
Component:317; Jaccard:0.16054
 |
Component:048; Jaccard:0.19236
 |
Component:048; Jaccard:0.20997
 |
Component:048; Jaccard:0.20825
 |
Component:048; Jaccard:0.18902
 |
Correlation:0.08457
  |
Correlation:0.08457
  |
Correlation:0.08457
  |
Correlation:0.093265
  |
Correlation:0.093265
  |
Correlation:0.093265
  |
Correlation:0.093265
  |
Component:264; Jaccard:0.093427
 |
Component:383; Jaccard:0.10268
 |
Component:287; Jaccard:0.11807
 |
Component:287; Jaccard:0.13626
 |
Component:287; Jaccard:0.14919
 |
Component:287; Jaccard:0.15008
 |
Component:287; Jaccard:0.13955
 |
Correlation:-0.0026473
  |
Correlation:0.17217
  |
Correlation:0.10199
  |
Correlation:0.10199
  |
Correlation:0.10199
  |
Correlation:0.10199
  |
Correlation:0.10199
  |
Component:026; Jaccard:0.089963
 |
Component:264; Jaccard:0.10193
 |
Component:049; Jaccard:0.11486
 |
Component:049; Jaccard:0.12897
 |
Component:049; Jaccard:0.13618
 |
Component:049; Jaccard:0.12846
 |
Component:213; Jaccard:0.12155
 |
Correlation:0.056744
  |
Correlation:-0.0026473
  |
Correlation:0.16738
  |
Correlation:0.16738
  |
Correlation:0.16738
  |
Correlation:0.16738
  |
Correlation:0.22832
  |
Component:383; Jaccard:0.088529
 |
Component:026; Jaccard:0.10066
 |
Component:383; Jaccard:0.11325
 |
Component:383; Jaccard:0.11984
 |
Component:213; Jaccard:0.12268
 |
Component:213; Jaccard:0.12654
 |
Component:049; Jaccard:0.1156
 |
Correlation:0.17217
  |
Correlation:0.056744
  |
Correlation:0.17217
  |
Correlation:0.17217
  |
Correlation:0.22832
  |
Correlation:0.22832
  |
Correlation:0.16738
  |
Component:150; Jaccard:0.084955
 |
Component:150; Jaccard:0.09994
 |
Component:150; Jaccard:0.11154
 |
Component:150; Jaccard:0.11538
 |
Component:383; Jaccard:0.11618
 |
Component:216; Jaccard:0.11329
 |
Component:218; Jaccard:0.10795
 |
Correlation:-0.033429
  |
Correlation:-0.033429
  |
Correlation:-0.033429
  |
Correlation:-0.033429
  |
Correlation:0.17217
  |
Correlation:0.12417
  |
Correlation:0.073289
  |
Component:049; Jaccard:0.08373
 |
Component:287; Jaccard:0.099536
 |
Component:026; Jaccard:0.109
 |
Component:026; Jaccard:0.1147
 |
Component:216; Jaccard:0.11449
 |
Component:218; Jaccard:0.11273
 |
Component:216; Jaccard:0.10451
 |
Correlation:0.16738
  |
Correlation:0.10199
  |
Correlation:0.056744
  |
Correlation:0.056744
  |
Correlation:0.12417
  |
Correlation:0.073289
  |
Correlation:0.12417
  |
Component:287; Jaccard:0.082487
 |
Component:049; Jaccard:0.098703
 |
Component:264; Jaccard:0.10495
 |
Component:213; Jaccard:0.1086
 |
Component:026; Jaccard:0.11447
 |
Component:383; Jaccard:0.10621
 |
Component:383; Jaccard:0.089113
 |
Correlation:0.10199
  |
Correlation:0.16738
  |
Correlation:-0.0026473
  |
Correlation:0.22832
  |
Correlation:0.056744
  |
Correlation:0.17217
  |
Correlation:0.17217
  |
Component:316; Jaccard:0.081066
 |
Component:316; Jaccard:0.087996
 |
Component:216; Jaccard:0.094114
 |
Component:216; Jaccard:0.10695
 |
Component:150; Jaccard:0.11087
 |
Component:026; Jaccard:0.10408
 |
Component:026; Jaccard:0.087585
 |
Correlation:-0.046521
  |
Correlation:-0.046521
  |
Correlation:0.12417
  |
Correlation:0.12417
  |
Correlation:-0.033429
  |
Correlation:0.056744
  |
Correlation:0.056744
  |
Cope: 02:"REL" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
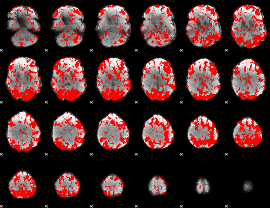 |
GLM binary result thresholded by 1.5
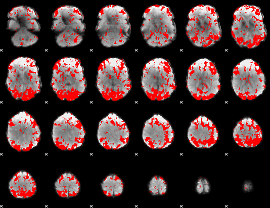 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
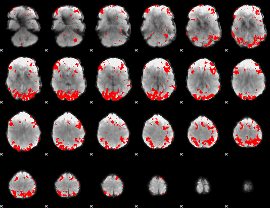 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
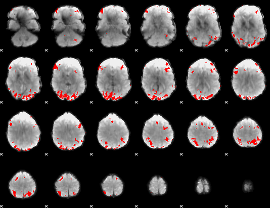 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:370; Jaccard:0.12632
 |
Component:317; Jaccard:0.15957
 |
Component:317; Jaccard:0.20106
 |
Component:317; Jaccard:0.23503
 |
Component:317; Jaccard:0.25095
 |
Component:317; Jaccard:0.24359
 |
Component:317; Jaccard:0.21991
 |
Correlation:0.0038149
  |
Correlation:0.1708
  |
Correlation:0.1708
  |
Correlation:0.1708
  |
Correlation:0.1708
  |
Correlation:0.1708
  |
Correlation:0.1708
  |
Component:048; Jaccard:0.12207
 |
Component:370; Jaccard:0.15887
 |
Component:370; Jaccard:0.19147
 |
Component:370; Jaccard:0.21482
 |
Component:370; Jaccard:0.22228
 |
Component:370; Jaccard:0.21486
 |
Component:370; Jaccard:0.19816
 |
Correlation:-0.0018871
  |
Correlation:0.0038149
  |
Correlation:0.0038149
  |
Correlation:0.0038149
  |
Correlation:0.0038149
  |
Correlation:0.0038149
  |
Correlation:0.0038149
  |
Component:317; Jaccard:0.12187
 |
Component:048; Jaccard:0.15042
 |
Component:048; Jaccard:0.17496
 |
Component:048; Jaccard:0.18849
 |
Component:287; Jaccard:0.20107
 |
Component:287; Jaccard:0.20255
 |
Component:287; Jaccard:0.18918
 |
Correlation:0.1708
  |
Correlation:-0.0018871
  |
Correlation:-0.0018871
  |
Correlation:-0.0018871
  |
Correlation:0.1822
  |
Correlation:0.1822
  |
Correlation:0.1822
  |
Component:287; Jaccard:0.10514
 |
Component:287; Jaccard:0.13087
 |
Component:287; Jaccard:0.15954
 |
Component:287; Jaccard:0.18641
 |
Component:048; Jaccard:0.18733
 |
Component:048; Jaccard:0.17243
 |
Component:213; Jaccard:0.16076
 |
Correlation:0.1822
  |
Correlation:0.1822
  |
Correlation:0.1822
  |
Correlation:0.1822
  |
Correlation:-0.0018871
  |
Correlation:-0.0018871
  |
Correlation:0.16829
  |
Component:150; Jaccard:0.10273
 |
Component:150; Jaccard:0.12072
 |
Component:150; Jaccard:0.13186
 |
Component:213; Jaccard:0.14506
 |
Component:213; Jaccard:0.16022
 |
Component:213; Jaccard:0.16797
 |
Component:048; Jaccard:0.14687
 |
Correlation:0.052634
  |
Correlation:0.052634
  |
Correlation:0.052634
  |
Correlation:0.16829
  |
Correlation:0.16829
  |
Correlation:0.16829
  |
Correlation:-0.0018871
  |
Component:026; Jaccard:0.095908
 |
Component:026; Jaccard:0.1087
 |
Component:213; Jaccard:0.12514
 |
Component:216; Jaccard:0.13931
 |
Component:216; Jaccard:0.14863
 |
Component:216; Jaccard:0.14984
 |
Component:216; Jaccard:0.13954
 |
Correlation:-0.14301
  |
Correlation:-0.14301
  |
Correlation:0.16829
  |
Correlation:0.057112
  |
Correlation:0.057112
  |
Correlation:0.057112
  |
Correlation:0.057112
  |
Component:387; Jaccard:0.092487
 |
Component:387; Jaccard:0.10482
 |
Component:216; Jaccard:0.12069
 |
Component:150; Jaccard:0.13284
 |
Component:282; Jaccard:0.13666
 |
Component:282; Jaccard:0.1371
 |
Component:282; Jaccard:0.12824
 |
Correlation:0.18588
  |
Correlation:0.18588
  |
Correlation:0.057112
  |
Correlation:0.052634
  |
Correlation:0.095533
  |
Correlation:0.095533
  |
Correlation:0.095533
  |
Component:049; Jaccard:0.087624
 |
Component:213; Jaccard:0.10226
 |
Component:026; Jaccard:0.11709
 |
Component:282; Jaccard:0.12931
 |
Component:218; Jaccard:0.1315
 |
Component:218; Jaccard:0.13111
 |
Component:218; Jaccard:0.12084
 |
Correlation:0.033124
  |
Correlation:0.16829
  |
Correlation:-0.14301
  |
Correlation:0.095533
  |
Correlation:0.14526
  |
Correlation:0.14526
  |
Correlation:0.14526
  |
Component:216; Jaccard:0.082411
 |
Component:049; Jaccard:0.10148
 |
Component:282; Jaccard:0.11531
 |
Component:218; Jaccard:0.12299
 |
Component:150; Jaccard:0.12249
 |
Component:049; Jaccard:0.10977
 |
Component:049; Jaccard:0.096843
 |
Correlation:0.057112
  |
Correlation:0.033124
  |
Correlation:0.095533
  |
Correlation:0.14526
  |
Correlation:0.052634
  |
Correlation:0.033124
  |
Correlation:0.033124
  |
Component:213; Jaccard:0.082211
 |
Component:216; Jaccard:0.10118
 |
Component:387; Jaccard:0.11259
 |
Component:026; Jaccard:0.1214
 |
Component:026; Jaccard:0.11716
 |
Component:026; Jaccard:0.10634
 |
Component:026; Jaccard:0.090909
 |
Correlation:0.16829
  |
Correlation:0.057112
  |
Correlation:0.18588
  |
Correlation:-0.14301
  |
Correlation:-0.14301
  |
Correlation:-0.14301
  |
Correlation:-0.14301
  |
Cope: 03:"MATCH-REL" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
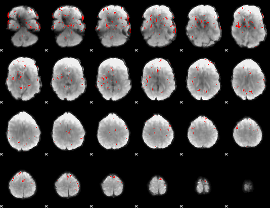 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
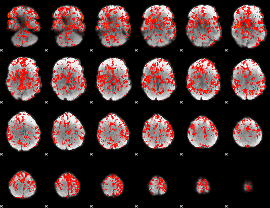 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:247; Jaccard:0.1016
 |
Component:307; Jaccard:0.10049
 |
Component:051; Jaccard:0.12846
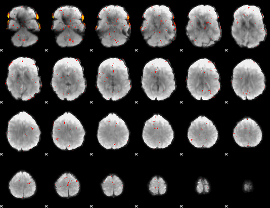 |
Component:051; Jaccard:0.16562
 |
Component:051; Jaccard:0.15455
 |
Component:051; Jaccard:0.10492
 |
Component:051; Jaccard:0.058103
 |
Correlation:0.20131
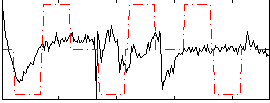  |
Correlation:0.14057
  |
Correlation:0.4662
  |
Correlation:0.4662
  |
Correlation:0.4662
  |
Correlation:0.4662
  |
Correlation:0.4662
  |
Component:286; Jaccard:0.098124
 |
Component:247; Jaccard:0.099408
 |
Component:323; Jaccard:0.10104
 |
Component:323; Jaccard:0.073626
 |
Component:065; Jaccard:0.046978
 |
Component:065; Jaccard:0.037074
 |
Component:065; Jaccard:0.02796
 |
Correlation:0.062238
  |
Correlation:0.20131
  |
Correlation:0.19234
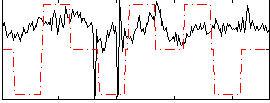  |
Correlation:0.19234
  |
Correlation:0.16184
  |
Correlation:0.16184
  |
Correlation:0.16184
  |
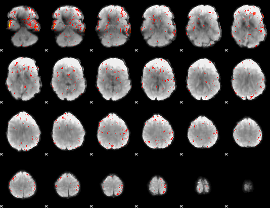
Component:044; Jaccard:0.090359
 |
Component:323; Jaccard:0.096744
 |
Component:307; Jaccard:0.093545
 |
Component:254; Jaccard:0.067074
 |
Component:249; Jaccard:0.043536
 |
Component:249; Jaccard:0.025636
 |
Component:249; Jaccard:0.015362
 |
Correlation:0.28326
  |
Correlation:0.19234
  |
Correlation:0.14057
  |
Correlation:0.21679
  |
Correlation:0.29546
  |
Correlation:0.29546
  |
Correlation:0.29546
  |
Component:307; Jaccard:0.089404
 |
Component:286; Jaccard:0.093716
 |
Component:019; Jaccard:0.089509
 |
Component:002; Jaccard:0.066861
 |
Component:318; Jaccard:0.041098
 |
Component:318; Jaccard:0.01886
 |
Component:146; Jaccard:0.009406
 |
Correlation:0.14057
  |
Correlation:0.062238
  |
Correlation:0.040942
  |
Correlation:0.11612
  |
Correlation:0.27065
  |
Correlation:0.27065
  |
Correlation:0.26493
  |
Component:152; Jaccard:0.08686
 |
Component:019; Jaccard:0.090664
 |
Component:254; Jaccard:0.085697
 |
Component:318; Jaccard:0.065564
 |
Component:323; Jaccard:0.03843
 |
Component:209; Jaccard:0.018056
 |
Component:201; Jaccard:0.007667
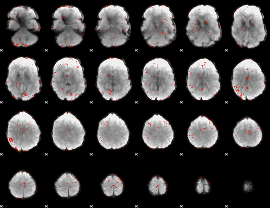 |
Correlation:0.12773
  |
Correlation:0.040942
  |
Correlation:0.21679
  |
Correlation:0.27065
  |
Correlation:0.19234
  |
Correlation:0.10189
  |
Correlation:-0.013347
  |
Component:323; Jaccard:0.081454
 |
Component:254; Jaccard:0.084681
 |
Component:002; Jaccard:0.083782
 |
Component:307; Jaccard:0.06455
 |
Component:146; Jaccard:0.037846
 |
Component:146; Jaccard:0.016112
 |
Component:002; Jaccard:0.007231
 |
Correlation:0.19234
  |
Correlation:0.21679
  |
Correlation:0.11612
  |
Correlation:0.14057
  |
Correlation:0.26493
  |
Correlation:0.26493
  |
Correlation:0.11612
  |
Component:056; Jaccard:0.080454
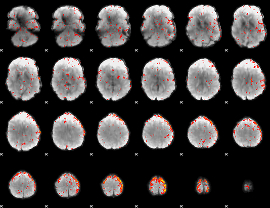 |
Component:056; Jaccard:0.082564
 |
Component:187; Jaccard:0.079191
 |
Component:019; Jaccard:0.06324
 |
Component:002; Jaccard:0.037233
 |
Component:002; Jaccard:0.015514
 |
Component:209; Jaccard:0.006507
 |
Correlation:0.31905
  |
Correlation:0.31905
  |
Correlation:0.11611
  |
Correlation:0.040942
  |
Correlation:0.11612
  |
Correlation:0.11612
  |
Correlation:0.10189
  |
Component:155; Jaccard:0.080291
 |
Component:002; Jaccard:0.081703
 |
Component:278; Jaccard:0.075058
 |
Component:249; Jaccard:0.061131
 |
Component:254; Jaccard:0.033939
 |
Component:052; Jaccard:0.014706
 |
Component:095; Jaccard:0.006235
 |
Correlation:0.26776
  |
Correlation:0.11612
  |
Correlation:0.20006
  |
Correlation:0.29546
  |
Correlation:0.21679
  |
Correlation:0.2236
  |
Correlation:0.02357
  |
Component:019; Jaccard:0.07651
 |
Component:051; Jaccard:0.081093
 |
Component:318; Jaccard:0.073556
 |
Component:209; Jaccard:0.056384
 |
Component:019; Jaccard:0.033309
 |
Component:323; Jaccard:0.013632
 |
Component:279; Jaccard:0.006214
 |
Correlation:0.040942
  |
Correlation:0.4662
  |
Correlation:0.27065
  |
Correlation:0.10189
  |
Correlation:0.040942
  |
Correlation:0.19234
  |
Correlation:-0.010885
  |
Component:149; Jaccard:0.075404
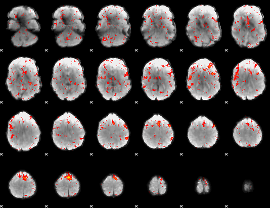 |
Component:044; Jaccard:0.080052
 |
Component:247; Jaccard:0.073058
 |
Component:187; Jaccard:0.05505
 |
Component:209; Jaccard:0.033213
 |
Component:243; Jaccard:0.012867
 |
Component:318; Jaccard:0.005572
 |
Correlation:0.20795
  |
Correlation:0.28326
  |
Correlation:0.20131
  |
Correlation:0.11611
  |
Correlation:0.10189
  |
Correlation:0.1744
  |
Correlation:0.27065
  |
Cope: 04:"REL-MATCH" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
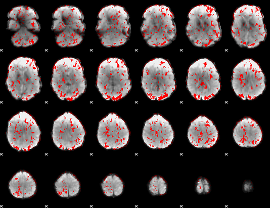 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
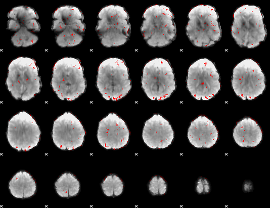 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
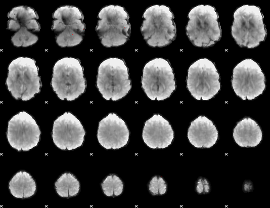 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
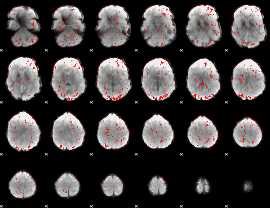 |
GLM Z-score result thresholded by 2
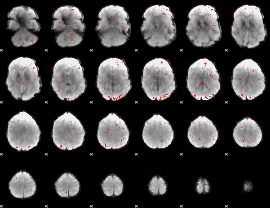 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
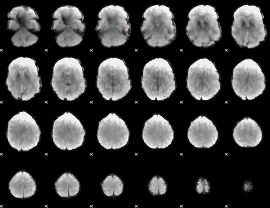 |
GLM Z-score result thresholded by 4
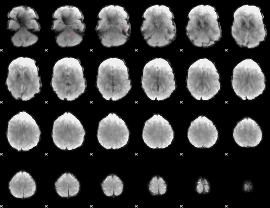 |
Component:351; Jaccard:0.11248
 |
Component:351; Jaccard:0.12007
 |
Component:351; Jaccard:0.097399
 |
Component:226; Jaccard:0.060327
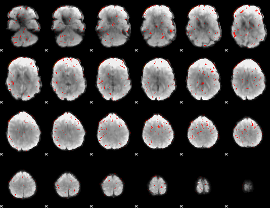 |
Component:226; Jaccard:0.028699
 |
Component:226; Jaccard:0.010675
 |
Component:226; Jaccard:0.003823
 |
Correlation:0.16936
  |
Correlation:0.16936
  |
Correlation:0.16936
  |
Correlation:0.24366
  |
Correlation:0.24366
  |
Correlation:0.24366
  |
Correlation:0.24366
  |
Component:387; Jaccard:0.10817
 |
Component:387; Jaccard:0.10728
 |
Component:226; Jaccard:0.096071
 |
Component:108; Jaccard:0.059065
 |
Component:400; Jaccard:0.027949
 |
Component:108; Jaccard:0.009366
 |
Component:400; Jaccard:0.002489
 |
Correlation:0.10248
  |
Correlation:0.10248
  |
Correlation:0.24366
  |
Correlation:0.16325
  |
Correlation:0.092844
  |
Correlation:0.16325
  |
Correlation:0.092844
  |
Component:151; Jaccard:0.095796
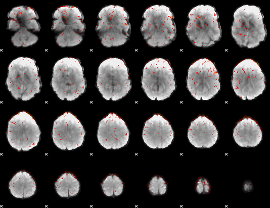 |
Component:151; Jaccard:0.10468
 |
Component:108; Jaccard:0.091971
 |
Component:400; Jaccard:0.057421
 |
Component:108; Jaccard:0.023112
 |
Component:400; Jaccard:0.00849
 |
Component:343; Jaccard:0.002099
 |
Correlation:0.23646
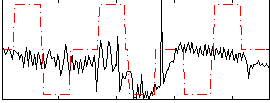 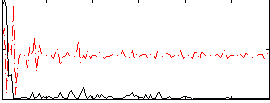 |
Correlation:0.23646
  |
Correlation:0.16325
  |
Correlation:0.092844
  |
Correlation:0.16325
  |
Correlation:0.092844
  |
Correlation:-0.043903
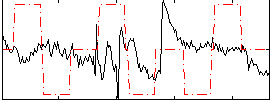  |
Component:226; Jaccard:0.087693
 |
Component:226; Jaccard:0.10152
 |
Component:151; Jaccard:0.086522
 |
Component:351; Jaccard:0.056812
 |
Component:351; Jaccard:0.022205
 |
Component:351; Jaccard:0.005839
 |
Component:393; Jaccard:0.001548
 |
Correlation:0.24366
  |
Correlation:0.24366
  |
Correlation:0.23646
  |
Correlation:0.16936
  |
Correlation:0.16936
  |
Correlation:0.16936
  |
Correlation:-0.1873
  |
Component:144; Jaccard:0.083859
 |
Component:108; Jaccard:0.095652
 |
Component:400; Jaccard:0.084571
 |
Component:151; Jaccard:0.048672
 |
Component:144; Jaccard:0.01817
 |
Component:144; Jaccard:0.005226
 |
Component:068; Jaccard:0.001503
 |
Correlation:0.25245
  |
Correlation:0.16325
  |
Correlation:0.092844
  |
Correlation:0.23646
  |
Correlation:0.25245
  |
Correlation:0.25245
  |
Correlation:0.34614
  |
Component:169; Jaccard:0.077526
 |
Component:144; Jaccard:0.089034
 |
Component:387; Jaccard:0.083415
 |
Component:387; Jaccard:0.046271
 |
Component:387; Jaccard:0.016726
 |
Component:248; Jaccard:0.00484
 |
Component:351; Jaccard:0.001362
 |
Correlation:0.23461
  |
Correlation:0.25245
  |
Correlation:0.10248
  |
Correlation:0.10248
  |
Correlation:0.10248
  |
Correlation:0.050918
  |
Correlation:0.16936
  |
Component:259; Jaccard:0.076928
 |
Component:400; Jaccard:0.086978
 |
Component:144; Jaccard:0.073824
 |
Component:144; Jaccard:0.043586
 |
Component:151; Jaccard:0.015615
 |
Component:343; Jaccard:0.004783
 |
Component:055; Jaccard:0.00132
 |
Correlation:0.23183
  |
Correlation:0.092844
  |
Correlation:0.25245
  |
Correlation:0.25245
  |
Correlation:0.23646
  |
Correlation:-0.043903
  |
Correlation:0.039927
  |
Component:108; Jaccard:0.076781
 |
Component:169; Jaccard:0.083325
 |
Component:169; Jaccard:0.070883
 |
Component:169; Jaccard:0.03832
 |
Component:085; Jaccard:0.01469
 |
Component:387; Jaccard:0.004645
 |
Component:108; Jaccard:0.001308
 |
Correlation:0.16325
  |
Correlation:0.23461
  |
Correlation:0.23461
  |
Correlation:0.23461
  |
Correlation:0.11876
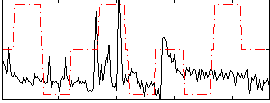  |
Correlation:0.10248
  |
Correlation:0.16325
  |
Component:195; Jaccard:0.075081
 |
Component:195; Jaccard:0.080317
 |
Component:195; Jaccard:0.065264
 |
Component:195; Jaccard:0.037295
 |
Component:195; Jaccard:0.014221
 |
Component:195; Jaccard:0.004519
 |
Component:165; Jaccard:0.001275
 |
Correlation:0.28363
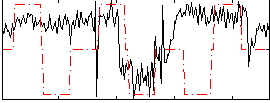  |
Correlation:0.28363
  |
Correlation:0.28363
  |
Correlation:0.28363
  |
Correlation:0.28363
  |
Correlation:0.28363
  |
Correlation:0.10803
  |
Component:268; Jaccard:0.074489
 |
Component:259; Jaccard:0.072813
 |
Component:085; Jaccard:0.056965
 |
Component:085; Jaccard:0.032635
 |
Component:248; Jaccard:0.012816
 |
Component:085; Jaccard:0.004245
 |
Component:206; Jaccard:0.001223
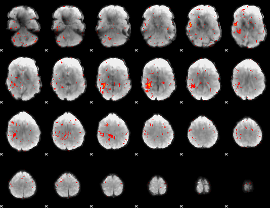 |
Correlation:0.20845
  |
Correlation:0.23183
  |
Correlation:0.11876
  |
Correlation:0.11876
  |
Correlation:0.050918
  |
Correlation:0.11876
  |
Correlation:0.043
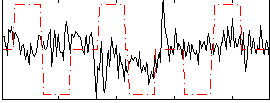  |
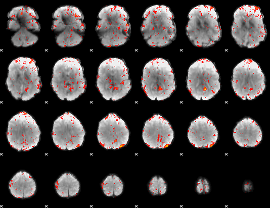
Cope: 05:"neg_MATCH" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
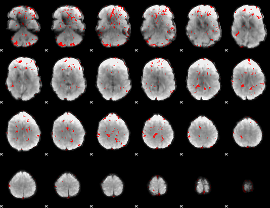 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
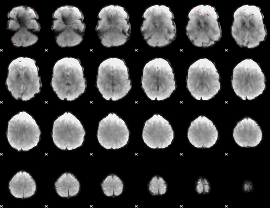 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
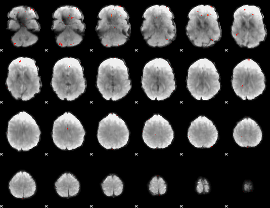 |
GLM Z-score result thresholded by 4
 |
Component:153; Jaccard:0.095518
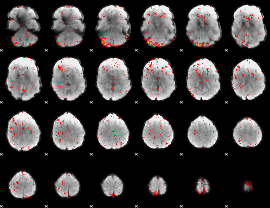 |
Component:153; Jaccard:0.095686
 |
Component:029; Jaccard:0.095527
 |
Component:392; Jaccard:0.086124
 |
Component:392; Jaccard:0.065858
 |
Component:392; Jaccard:0.041233
 |
Component:392; Jaccard:0.021783
 |
Correlation:0.11305
  |
Correlation:0.11305
  |
Correlation:0.12093
  |
Correlation:0.12895
  |
Correlation:0.12895
  |
Correlation:0.12895
  |
Correlation:0.12895
  |
Component:334; Jaccard:0.083647
 |
Component:029; Jaccard:0.089276
 |
Component:392; Jaccard:0.094752
 |
Component:029; Jaccard:0.083194
 |
Component:029; Jaccard:0.054037
 |
Component:029; Jaccard:0.027968
 |
Component:045; Jaccard:0.013901
 |
Correlation:0.16665
  |
Correlation:0.12093
  |
Correlation:0.12895
  |
Correlation:0.12093
  |
Correlation:0.12093
  |
Correlation:0.12093
  |
Correlation:0.06107
  |
Component:102; Jaccard:0.081494
 |
Component:392; Jaccard:0.087761
 |
Component:102; Jaccard:0.084084
 |
Component:102; Jaccard:0.068772
 |
Component:102; Jaccard:0.043817
 |
Component:045; Jaccard:0.025082
 |
Component:029; Jaccard:0.011384
 |
Correlation:-0.17209
  |
Correlation:0.12895
  |
Correlation:-0.17209
  |
Correlation:-0.17209
  |
Correlation:-0.17209
  |
Correlation:0.06107
  |
Correlation:0.12093
  |
Component:029; Jaccard:0.077684
 |
Component:334; Jaccard:0.086753
 |
Component:153; Jaccard:0.079404
 |
Component:334; Jaccard:0.061324
 |
Component:334; Jaccard:0.038635
 |
Component:102; Jaccard:0.023246
 |
Component:102; Jaccard:0.008817
 |
Correlation:0.12093
  |
Correlation:0.16665
  |
Correlation:0.11305
  |
Correlation:0.16665
  |
Correlation:0.16665
  |
Correlation:-0.17209
  |
Correlation:-0.17209
  |
Component:392; Jaccard:0.075139
 |
Component:102; Jaccard:0.086619
 |
Component:334; Jaccard:0.078328
 |
Component:153; Jaccard:0.058244
 |
Component:045; Jaccard:0.035838
 |
Component:334; Jaccard:0.021204
 |
Component:334; Jaccard:0.008703
 |
Correlation:0.12895
  |
Correlation:-0.17209
  |
Correlation:0.16665
  |
Correlation:0.11305
  |
Correlation:0.06107
  |
Correlation:0.16665
  |
Correlation:0.16665
  |
Component:351; Jaccard:0.074645
 |
Component:351; Jaccard:0.072019
 |
Component:088; Jaccard:0.063665
 |
Component:114; Jaccard:0.050386
 |
Component:153; Jaccard:0.033828
 |
Component:088; Jaccard:0.016125
 |
Component:153; Jaccard:0.007028
 |
Correlation:0.13958
  |
Correlation:0.13958
  |
Correlation:-0.14311
  |
Correlation:0.0012366
  |
Correlation:0.11305
  |
Correlation:-0.14311
  |
Correlation:0.11305
  |
Component:194; Jaccard:0.07346
 |
Component:107; Jaccard:0.070113
 |
Component:389; Jaccard:0.06202
 |
Component:107; Jaccard:0.048284
 |
Component:107; Jaccard:0.030683
 |
Component:199; Jaccard:0.015641
 |
Component:005; Jaccard:0.006858
 |
Correlation:0.0062667
  |
Correlation:0.087185
  |
Correlation:0.21219
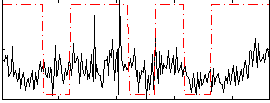  |
Correlation:0.087185
  |
Correlation:0.087185
  |
Correlation:-0.11825
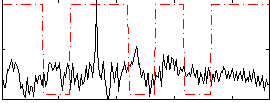  |
Correlation:0.039897
  |
Component:088; Jaccard:0.071557
 |
Component:088; Jaccard:0.069127
 |
Component:107; Jaccard:0.061679
 |
Component:389; Jaccard:0.04668
 |
Component:114; Jaccard:0.030564
 |
Component:107; Jaccard:0.015053
 |
Component:199; Jaccard:0.006837
 |
Correlation:-0.14311
  |
Correlation:-0.14311
  |
Correlation:0.087185
  |
Correlation:0.21219
  |
Correlation:0.0012366
  |
Correlation:0.087185
  |
Correlation:-0.11825
  |
Component:207; Jaccard:0.071227
 |
Component:389; Jaccard:0.066936
 |
Component:114; Jaccard:0.059591
 |
Component:088; Jaccard:0.046478
 |
Component:389; Jaccard:0.030145
 |
Component:153; Jaccard:0.014948
 |
Component:109; Jaccard:0.006581
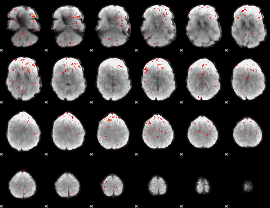 |
Correlation:0.083203
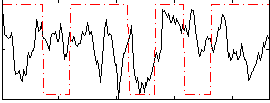  |
Correlation:0.21219
  |
Correlation:0.0012366
  |
Correlation:-0.14311
  |
Correlation:0.21219
  |
Correlation:0.11305
  |
Correlation:0.066954
  |
Component:023; Jaccard:0.069991
 |
Component:207; Jaccard:0.064768
 |
Component:109; Jaccard:0.059158
 |
Component:055; Jaccard:0.045027
 |
Component:055; Jaccard:0.028706
 |
Component:005; Jaccard:0.014849
 |
Component:344; Jaccard:0.006371
 |
Correlation:0.066648
  |
Correlation:0.083203
  |
Correlation:0.066954
  |
Correlation:0.11506
  |
Correlation:0.11506
  |
Correlation:0.039897
  |
Correlation:0.16191
  |
Cope: 06:"neg_REL" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
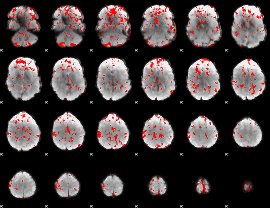 |
GLM binary result thresholded by 2
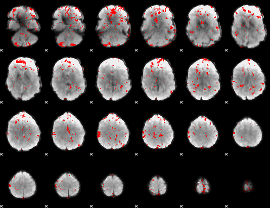 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
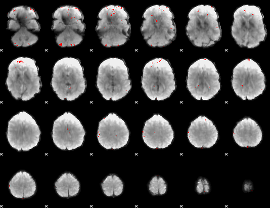 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:308; Jaccard:0.10628
 |
Component:334; Jaccard:0.11181
 |
Component:334; Jaccard:0.11601
 |
Component:334; Jaccard:0.10047
 |
Component:334; Jaccard:0.073417
 |
Component:334; Jaccard:0.043103
 |
Component:334; Jaccard:0.022197
 |
Correlation:0.2482
  |
Correlation:0.19791
  |
Correlation:0.19791
  |
Correlation:0.19791
  |
Correlation:0.19791
  |
Correlation:0.19791
  |
Correlation:0.19791
  |
Component:194; Jaccard:0.10538
 |
Component:308; Jaccard:0.10883
 |
Component:194; Jaccard:0.097413
 |
Component:194; Jaccard:0.074331
 |
Component:102; Jaccard:0.053943
 |
Component:392; Jaccard:0.034056
 |
Component:392; Jaccard:0.020432
 |
Correlation:0.1742
  |
Correlation:0.2482
  |
Correlation:0.1742
  |
Correlation:0.1742
  |
Correlation:0.28119
  |
Correlation:0.283
  |
Correlation:0.283
  |
Component:334; Jaccard:0.098962
 |
Component:194; Jaccard:0.10617
 |
Component:308; Jaccard:0.095587
 |
Component:102; Jaccard:0.072087
 |
Component:392; Jaccard:0.052005
 |
Component:102; Jaccard:0.032402
 |
Component:102; Jaccard:0.014219
 |
Correlation:0.19791
  |
Correlation:0.1742
  |
Correlation:0.2482
  |
Correlation:0.28119
  |
Correlation:0.283
  |
Correlation:0.28119
  |
Correlation:0.28119
  |
Component:181; Jaccard:0.097906
 |
Component:181; Jaccard:0.097544
 |
Component:029; Jaccard:0.085138
 |
Component:029; Jaccard:0.070388
 |
Component:029; Jaccard:0.049672
 |
Component:210; Jaccard:0.027435
 |
Component:210; Jaccard:0.013735
 |
Correlation:0.25287
  |
Correlation:0.25287
  |
Correlation:0.18487
  |
Correlation:0.18487
  |
Correlation:0.18487
  |
Correlation:0.30351
  |
Correlation:0.30351
  |
Component:023; Jaccard:0.08145
 |
Component:088; Jaccard:0.082696
 |
Component:102; Jaccard:0.083865
 |
Component:308; Jaccard:0.070282
 |
Component:194; Jaccard:0.045778
 |
Component:029; Jaccard:0.025216
 |
Component:088; Jaccard:0.010225
 |
Correlation:0.19601
  |
Correlation:0.30093
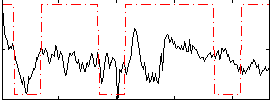  |
Correlation:0.28119
  |
Correlation:0.2482
  |
Correlation:0.1742
  |
Correlation:0.18487
  |
Correlation:0.30093
  |
Component:153; Jaccard:0.080943
 |
Component:153; Jaccard:0.08267
 |
Component:181; Jaccard:0.083615
 |
Component:392; Jaccard:0.064585
 |
Component:210; Jaccard:0.043483
 |
Component:088; Jaccard:0.024177
 |
Component:199; Jaccard:0.010169
 |
Correlation:0.1411
  |
Correlation:0.1411
  |
Correlation:0.25287
  |
Correlation:0.283
  |
Correlation:0.30351
  |
Correlation:0.30093
  |
Correlation:0.22348
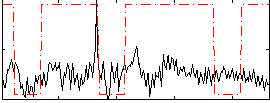  |
Component:088; Jaccard:0.078611
 |
Component:090; Jaccard:0.082015
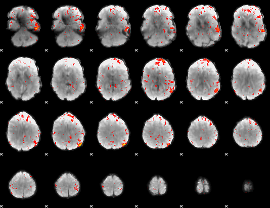 |
Component:088; Jaccard:0.078825
 |
Component:088; Jaccard:0.06293
 |
Component:088; Jaccard:0.042438
 |
Component:005; Jaccard:0.022291
 |
Component:045; Jaccard:0.009442
 |
Correlation:0.30093
  |
Correlation:0.20522
  |
Correlation:0.30093
  |
Correlation:0.30093
  |
Correlation:0.30093
  |
Correlation:0.17032
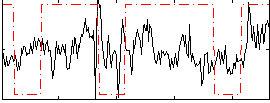  |
Correlation:-0.13829
  |
Component:090; Jaccard:0.0756
 |
Component:102; Jaccard:0.081935
 |
Component:090; Jaccard:0.076254
 |
Component:181; Jaccard:0.06079
 |
Component:308; Jaccard:0.040669
 |
Component:194; Jaccard:0.020332
 |
Component:194; Jaccard:0.008838
 |
Correlation:0.20522
  |
Correlation:0.28119
  |
Correlation:0.20522
  |
Correlation:0.25287
  |
Correlation:0.2482
  |
Correlation:0.1742
  |
Correlation:0.1742
  |
Component:307; Jaccard:0.071666
 |
Component:023; Jaccard:0.081502
 |
Component:153; Jaccard:0.072204
 |
Component:090; Jaccard:0.058875
 |
Component:181; Jaccard:0.038701
 |
Component:199; Jaccard:0.02002
 |
Component:129; Jaccard:0.008756
 |
Correlation:0.093776
  |
Correlation:0.19601
  |
Correlation:0.1411
  |
Correlation:0.20522
  |
Correlation:0.25287
  |
Correlation:0.22348
  |
Correlation:0.035139
  |
Component:102; Jaccard:0.07054
 |
Component:029; Jaccard:0.080154
 |
Component:023; Jaccard:0.071691
 |
Component:210; Jaccard:0.055928
 |
Component:005; Jaccard:0.036647
 |
Component:181; Jaccard:0.019954
 |
Component:005; Jaccard:0.008727
 |
Correlation:0.28119
  |
Correlation:0.18487
  |
Correlation:0.19601
  |
Correlation:0.30351
  |
Correlation:0.17032
  |
Correlation:0.25287
  |
Correlation:0.17032
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































