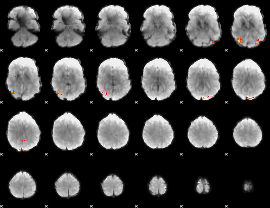
Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"FACES" BACK
|
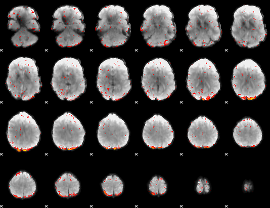
GLM result group-wise
 |
GLM binary result thresholded by 1
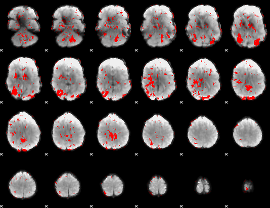 |
GLM binary result thresholded by 1.5
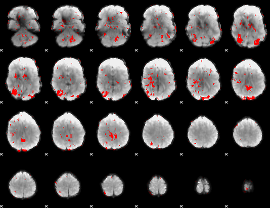 |
GLM binary result thresholded by 2
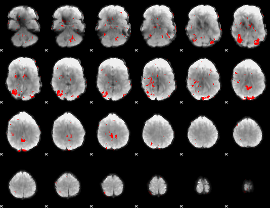 |
GLM binary result thresholded by 2.5
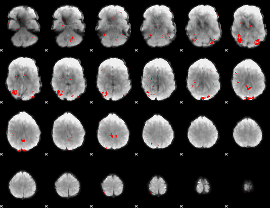 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
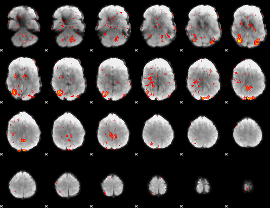 |
GLM Z-score result thresholded by 2
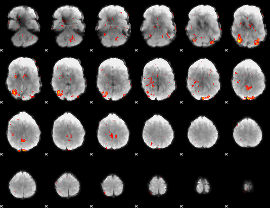 |
GLM Z-score result thresholded by 2.5
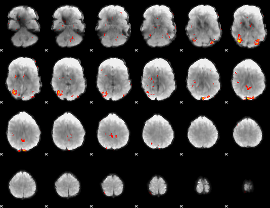 |
GLM Z-score result thresholded by 3
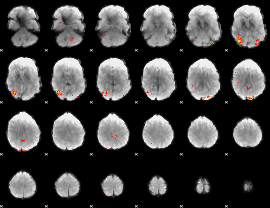 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:365; Jaccard:0.1524
 |
Component:365; Jaccard:0.16343
 |
Component:365; Jaccard:0.16095
 |
Component:365; Jaccard:0.13724
 |
Component:365; Jaccard:0.11004
 |
Component:365; Jaccard:0.083454
 |
Component:365; Jaccard:0.058792
 |
Correlation:0.54408
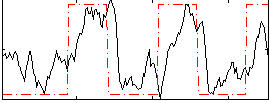 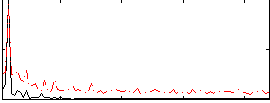 |
Correlation:0.54408
  |
Correlation:0.54408
  |
Correlation:0.54408
  |
Correlation:0.54408
  |
Correlation:0.54408
  |
Correlation:0.54408
  |
Component:228; Jaccard:0.114
 |
Component:228; Jaccard:0.12037
 |
Component:224; Jaccard:0.10778
 |
Component:224; Jaccard:0.092165
 |
Component:224; Jaccard:0.069789
 |
Component:224; Jaccard:0.047491
 |
Component:224; Jaccard:0.032578
 |
Correlation:0.050002
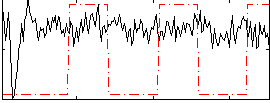 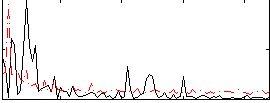 |
Correlation:0.050002
  |
Correlation:0.12503
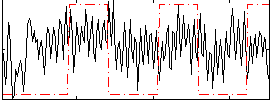 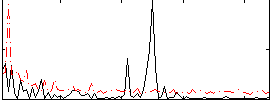 |
Correlation:0.12503
  |
Correlation:0.12503
  |
Correlation:0.12503
  |
Correlation:0.12503
  |
Component:224; Jaccard:0.11056
 |
Component:224; Jaccard:0.11677
 |
Component:228; Jaccard:0.10724
 |
Component:163; Jaccard:0.086051
 |
Component:163; Jaccard:0.067235
 |
Component:163; Jaccard:0.047037
 |
Component:163; Jaccard:0.030635
 |
Correlation:0.12503
  |
Correlation:0.12503
  |
Correlation:0.050002
  |
Correlation:0.31172
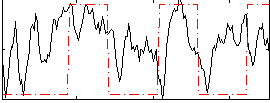  |
Correlation:0.31172
  |
Correlation:0.31172
  |
Correlation:0.31172
  |
Component:046; Jaccard:0.098772
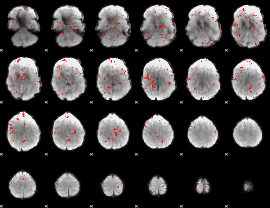 |
Component:163; Jaccard:0.099877
 |
Component:163; Jaccard:0.098247
 |
Component:228; Jaccard:0.084879
 |
Component:228; Jaccard:0.061533
 |
Component:228; Jaccard:0.039208
 |
Component:228; Jaccard:0.024785
 |
Correlation:0.2359
  |
Correlation:0.31172
  |
Correlation:0.31172
  |
Correlation:0.050002
  |
Correlation:0.050002
  |
Correlation:0.050002
  |
Correlation:0.050002
  |
Component:163; Jaccard:0.093628
 |
Component:046; Jaccard:0.091647
 |
Component:046; Jaccard:0.072237
 |
Component:393; Jaccard:0.05197
 |
Component:393; Jaccard:0.040489
 |
Component:393; Jaccard:0.030437
 |
Component:393; Jaccard:0.021853
 |
Correlation:0.31172
  |
Correlation:0.2359
  |
Correlation:0.2359
  |
Correlation:0.28648
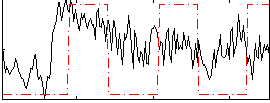 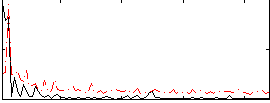 |
Correlation:0.28648
  |
Correlation:0.28648
  |
Correlation:0.28648
  |
Component:141; Jaccard:0.081345
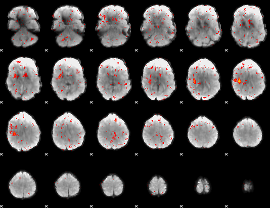 |
Component:141; Jaccard:0.072802
 |
Component:029; Jaccard:0.057895
 |
Component:046; Jaccard:0.049392
 |
Component:046; Jaccard:0.03513
 |
Component:046; Jaccard:0.023516
 |
Component:258; Jaccard:0.016767
 |
Correlation:-0.093249
  |
Correlation:-0.093249
  |
Correlation:-0.12392
  |
Correlation:0.2359
  |
Correlation:0.2359
  |
Correlation:0.2359
  |
Correlation:0.1273
  |
Component:343; Jaccard:0.073108
 |
Component:343; Jaccard:0.069682
 |
Component:393; Jaccard:0.057807
 |
Component:058; Jaccard:0.039905
 |
Component:058; Jaccard:0.031679
 |
Component:258; Jaccard:0.022373
 |
Component:046; Jaccard:0.015682
 |
Correlation:-0.078417
  |
Correlation:-0.078417
  |
Correlation:0.28648
  |
Correlation:0.069275
  |
Correlation:0.069275
  |
Correlation:0.1273
  |
Correlation:0.2359
  |
Component:258; Jaccard:0.06942
 |
Component:029; Jaccard:0.066838
 |
Component:141; Jaccard:0.055325
 |
Component:126; Jaccard:0.039753
 |
Component:258; Jaccard:0.028971
 |
Component:058; Jaccard:0.020197
 |
Component:126; Jaccard:0.013195
 |
Correlation:0.1273
  |
Correlation:-0.12392
  |
Correlation:-0.093249
  |
Correlation:-0.11397
  |
Correlation:0.1273
  |
Correlation:0.069275
  |
Correlation:-0.11397
  |
Component:301; Jaccard:0.068194
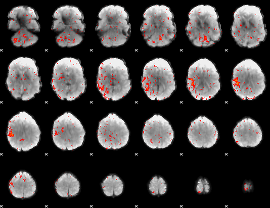 |
Component:258; Jaccard:0.06607
 |
Component:073; Jaccard:0.053859
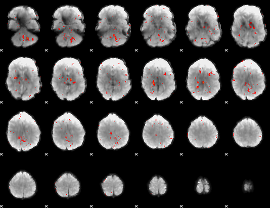 |
Component:029; Jaccard:0.039721
 |
Component:126; Jaccard:0.028007
 |
Component:126; Jaccard:0.019725
 |
Component:058; Jaccard:0.0122
 |
Correlation:0.06067
  |
Correlation:0.1273
  |
Correlation:-0.086496
  |
Correlation:-0.12392
  |
Correlation:-0.11397
  |
Correlation:-0.11397
  |
Correlation:0.069275
  |
Component:029; Jaccard:0.066871
 |
Component:126; Jaccard:0.063156
 |
Component:343; Jaccard:0.052607
 |
Component:343; Jaccard:0.038753
 |
Component:343; Jaccard:0.025547
 |
Component:079; Jaccard:0.01562
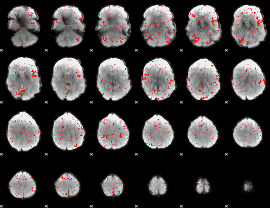 |
Component:079; Jaccard:0.012184
 |
Correlation:-0.12392
  |
Correlation:-0.11397
  |
Correlation:-0.078417
  |
Correlation:-0.078417
  |
Correlation:-0.078417
  |
Correlation:0.23221
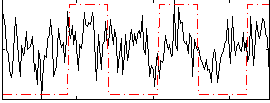 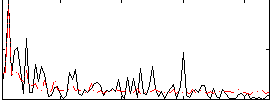 |
Correlation:0.23221
  |
Cope: 02:"SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
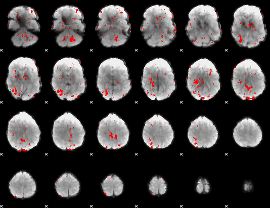 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
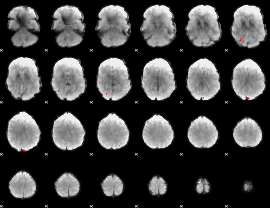 |
GLM Z-score result thresholded by 1
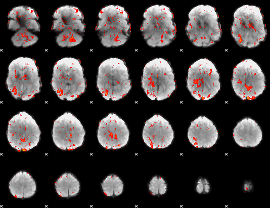 |
GLM Z-score result thresholded by 1.5
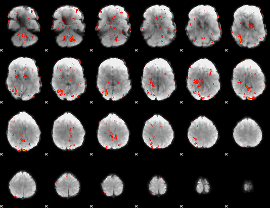 |
GLM Z-score result thresholded by 2
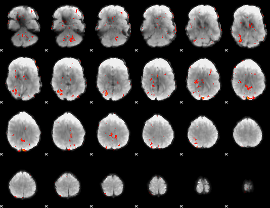 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:224; Jaccard:0.099268
 |
Component:224; Jaccard:0.10057
 |
Component:224; Jaccard:0.089607
 |
Component:073; Jaccard:0.071924
 |
Component:224; Jaccard:0.041009
 |
Component:163; Jaccard:0.022731
 |
Component:163; Jaccard:0.01443
 |
Correlation:-0.027095
  |
Correlation:-0.027095
  |
Correlation:-0.027095
  |
Correlation:0.28173
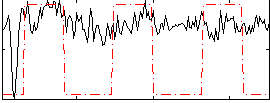 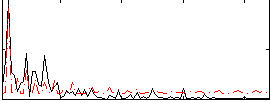 |
Correlation:-0.027095
  |
Correlation:-0.1588
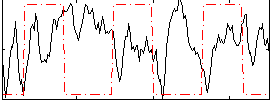 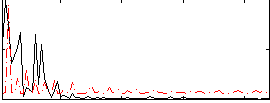 |
Correlation:-0.1588
  |
Component:228; Jaccard:0.097766
 |
Component:228; Jaccard:0.10055
 |
Component:228; Jaccard:0.087622
 |
Component:228; Jaccard:0.067232
 |
Component:343; Jaccard:0.039992
 |
Component:224; Jaccard:0.021288
 |
Component:393; Jaccard:0.010862
 |
Correlation:0.093554
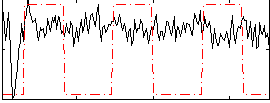 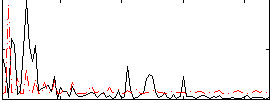 |
Correlation:0.093554
  |
Correlation:0.093554
  |
Correlation:0.093554
  |
Correlation:0.18497
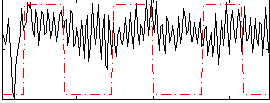 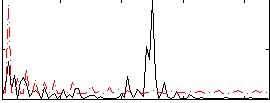 |
Correlation:-0.027095
  |
Correlation:-0.23932
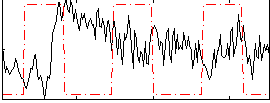 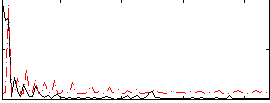 |
Component:343; Jaccard:0.094994
 |
Component:343; Jaccard:0.096931
 |
Component:073; Jaccard:0.087568
 |
Component:343; Jaccard:0.066667
 |
Component:228; Jaccard:0.039584
 |
Component:343; Jaccard:0.017724
 |
Component:224; Jaccard:0.009239
 |
Correlation:0.18497
  |
Correlation:0.18497
  |
Correlation:0.28173
  |
Correlation:0.18497
  |
Correlation:0.093554
  |
Correlation:0.18497
  |
Correlation:-0.027095
  |
Component:029; Jaccard:0.084046
 |
Component:029; Jaccard:0.090206
 |
Component:343; Jaccard:0.086107
 |
Component:224; Jaccard:0.066212
 |
Component:073; Jaccard:0.038333
 |
Component:228; Jaccard:0.016403
 |
Component:343; Jaccard:0.009033
 |
Correlation:0.33838
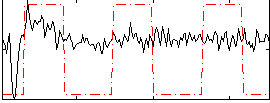 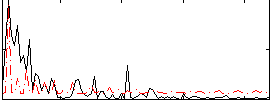 |
Correlation:0.33838
  |
Correlation:0.18497
  |
Correlation:-0.027095
  |
Correlation:0.28173
  |
Correlation:0.093554
  |
Correlation:0.18497
  |
Component:163; Jaccard:0.080509
 |
Component:073; Jaccard:0.087152
 |
Component:029; Jaccard:0.081759
 |
Component:029; Jaccard:0.062731
 |
Component:163; Jaccard:0.035426
 |
Component:073; Jaccard:0.016118
 |
Component:228; Jaccard:0.007616
 |
Correlation:-0.1588
  |
Correlation:0.28173
  |
Correlation:0.33838
  |
Correlation:0.33838
  |
Correlation:-0.1588
  |
Correlation:0.28173
  |
Correlation:0.093554
  |
Component:073; Jaccard:0.079539
 |
Component:163; Jaccard:0.080577
 |
Component:163; Jaccard:0.071132
 |
Component:163; Jaccard:0.054751
 |
Component:029; Jaccard:0.033677
 |
Component:393; Jaccard:0.015956
 |
Component:365; Jaccard:0.007169
 |
Correlation:0.28173
  |
Correlation:-0.1588
  |
Correlation:-0.1588
  |
Correlation:-0.1588
  |
Correlation:0.33838
  |
Correlation:-0.23932
  |
Correlation:-0.5592
 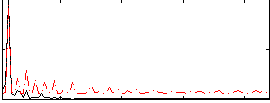 |
Component:141; Jaccard:0.072146
 |
Component:258; Jaccard:0.066416
 |
Component:126; Jaccard:0.054149
 |
Component:126; Jaccard:0.037719
 |
Component:393; Jaccard:0.025313
 |
Component:365; Jaccard:0.011716
 |
Component:073; Jaccard:0.005558
 |
Correlation:0.094264
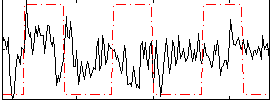 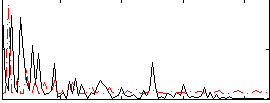 |
Correlation:0.0093243
 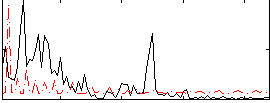 |
Correlation:0.22557
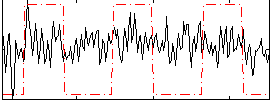 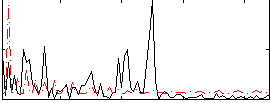 |
Correlation:0.22557
  |
Correlation:-0.23932
  |
Correlation:-0.5592
  |
Correlation:0.28173
  |
Component:258; Jaccard:0.071962
 |
Component:126; Jaccard:0.064805
 |
Component:258; Jaccard:0.052834
 |
Component:393; Jaccard:0.037364
 |
Component:396; Jaccard:0.020677
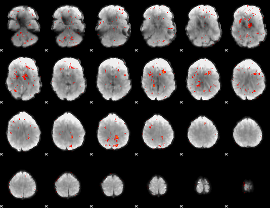 |
Component:029; Jaccard:0.011364
 |
Component:058; Jaccard:0.004988
 |
Correlation:0.0093243
  |
Correlation:0.22557
  |
Correlation:0.0093243
  |
Correlation:-0.23932
  |
Correlation:0.2199
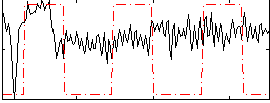  |
Correlation:0.33838
  |
Correlation:-0.045878
  |
Component:126; Jaccard:0.067584
 |
Component:141; Jaccard:0.061767
 |
Component:012; Jaccard:0.048621
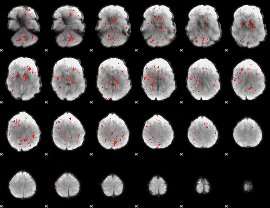 |
Component:258; Jaccard:0.036671
 |
Component:126; Jaccard:0.020322
 |
Component:058; Jaccard:0.010447
 |
Component:029; Jaccard:0.004851
 |
Correlation:0.22557
  |
Correlation:0.094264
  |
Correlation:0.27396
  |
Correlation:0.0093243
  |
Correlation:0.22557
  |
Correlation:-0.045878
  |
Correlation:0.33838
  |
Component:012; Jaccard:0.067319
 |
Component:012; Jaccard:0.059347
 |
Component:393; Jaccard:0.047619
 |
Component:396; Jaccard:0.033677
 |
Component:258; Jaccard:0.019719
 |
Component:126; Jaccard:0.007678
 |
Component:258; Jaccard:0.003983
 |
Correlation:0.27396
  |
Correlation:0.27396
  |
Correlation:-0.23932
  |
Correlation:0.2199
  |
Correlation:0.0093243
  |
Correlation:0.22557
  |
Correlation:0.0093243
  |
Cope: 03:"FACES-SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
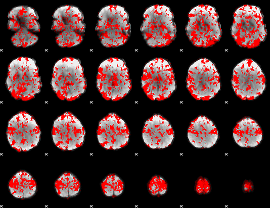 |
GLM binary result thresholded by 1.5
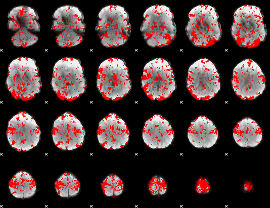 |
GLM binary result thresholded by 2
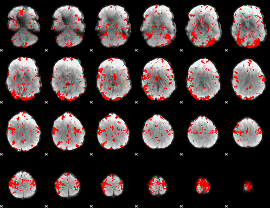 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
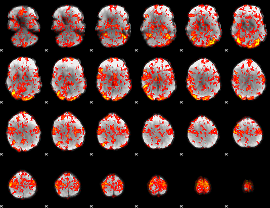 |
GLM Z-score result thresholded by 1.5
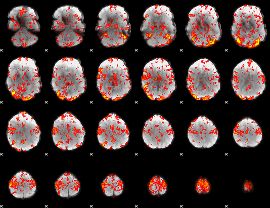 |
GLM Z-score result thresholded by 2
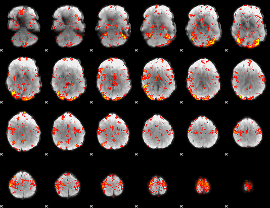 |
GLM Z-score result thresholded by 2.5
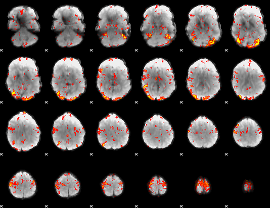 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:054; Jaccard:0.11995
 |
Component:365; Jaccard:0.15219
 |
Component:365; Jaccard:0.21188
 |
Component:365; Jaccard:0.27064
 |
Component:365; Jaccard:0.30499
 |
Component:365; Jaccard:0.29699
 |
Component:365; Jaccard:0.26819
 |
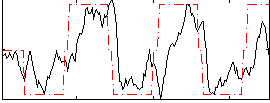
Correlation:0.32425
  |
Correlation:0.59843
 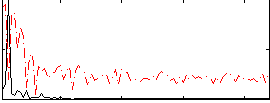 |
Correlation:0.59843
  |
Correlation:0.59843
  |
Correlation:0.59843
  |
Correlation:0.59843
  |
Correlation:0.59843
  |
Component:365; Jaccard:0.10777
 |
Component:054; Jaccard:0.13868
 |
Component:054; Jaccard:0.14846
 |
Component:054; Jaccard:0.14317
 |
Component:251; Jaccard:0.13371
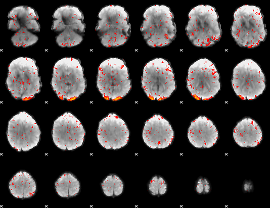 |
Component:251; Jaccard:0.12788
 |
Component:251; Jaccard:0.11277
 |
Correlation:0.59843
  |
Correlation:0.32425
  |
Correlation:0.32425
  |
Correlation:0.32425
  |
Correlation:0.36473
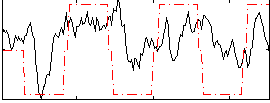  |
Correlation:0.36473
  |
Correlation:0.36473
  |
Component:118; Jaccard:0.097073
 |
Component:118; Jaccard:0.11406
 |
Component:118; Jaccard:0.12868
 |
Component:118; Jaccard:0.13416
 |
Component:140; Jaccard:0.12641
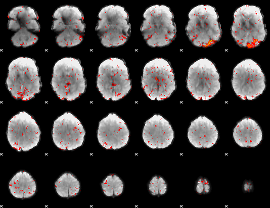 |
Component:140; Jaccard:0.11648
 |
Component:140; Jaccard:0.10204
 |
Correlation:0.5206
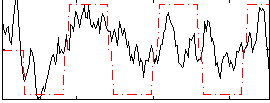  |
Correlation:0.5206
  |
Correlation:0.5206
  |
Correlation:0.5206
  |
Correlation:0.342
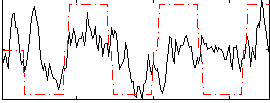 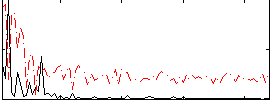 |
Correlation:0.342
  |
Correlation:0.342
  |
Component:166; Jaccard:0.096417
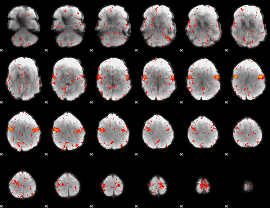 |
Component:166; Jaccard:0.10795
 |
Component:364; Jaccard:0.12152
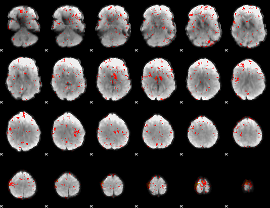 |
Component:251; Jaccard:0.12933
 |
Component:118; Jaccard:0.12432
 |
Component:118; Jaccard:0.10548
 |
Component:118; Jaccard:0.081087
 |
Correlation:0.33983
  |
Correlation:0.33983
  |
Correlation:0.26152
  |
Correlation:0.36473
  |
Correlation:0.5206
  |
Correlation:0.5206
  |
Correlation:0.5206
  |
Component:039; Jaccard:0.092047
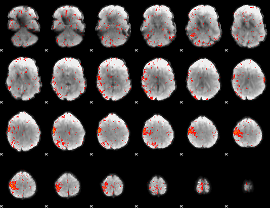 |
Component:364; Jaccard:0.10725
 |
Component:140; Jaccard:0.11642
 |
Component:364; Jaccard:0.12697
 |
Component:054; Jaccard:0.12333
 |
Component:364; Jaccard:0.10022
 |
Component:364; Jaccard:0.07349
 |
Correlation:0.42028
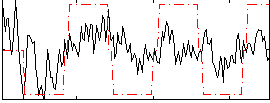 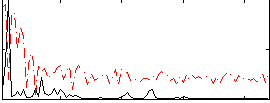 |
Correlation:0.26152
  |
Correlation:0.342
  |
Correlation:0.26152
  |
Correlation:0.32425
  |
Correlation:0.26152
  |
Correlation:0.26152
  |
Component:364; Jaccard:0.09084
 |
Component:039; Jaccard:0.10567
 |
Component:251; Jaccard:0.11603
 |
Component:140; Jaccard:0.12537
 |
Component:364; Jaccard:0.11824
 |
Component:054; Jaccard:0.096485
 |
Component:054; Jaccard:0.068891
 |
Correlation:0.26152
  |
Correlation:0.42028
  |
Correlation:0.36473
  |
Correlation:0.342
  |
Correlation:0.26152
  |
Correlation:0.32425
  |
Correlation:0.32425
  |
Component:247; Jaccard:0.086952
 |
Component:140; Jaccard:0.10107
 |
Component:039; Jaccard:0.11256
 |
Component:039; Jaccard:0.11155
 |
Component:039; Jaccard:0.10147
 |
Component:039; Jaccard:0.084466
 |
Component:039; Jaccard:0.061869
 |
Correlation:0.21175
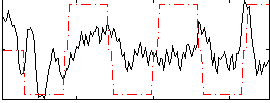 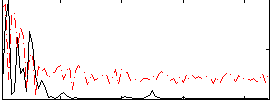 |
Correlation:0.342
  |
Correlation:0.42028
  |
Correlation:0.42028
  |
Correlation:0.42028
  |
Correlation:0.42028
  |
Correlation:0.42028
  |
Component:345; Jaccard:0.086344
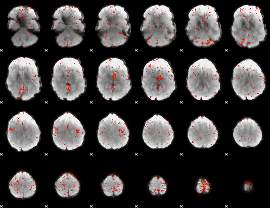 |
Component:251; Jaccard:0.10041
 |
Component:166; Jaccard:0.1119
 |
Component:166; Jaccard:0.10589
 |
Component:166; Jaccard:0.087848
 |
Component:369; Jaccard:0.070418
 |
Component:219; Jaccard:0.057485
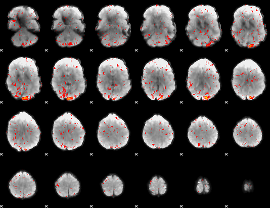 |
Correlation:0.12041
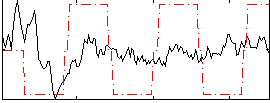 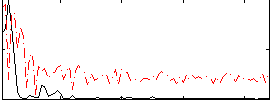 |
Correlation:0.36473
  |
Correlation:0.33983
  |
Correlation:0.33983
  |
Correlation:0.33983
  |
Correlation:0.34739
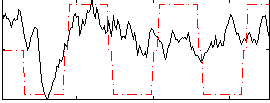 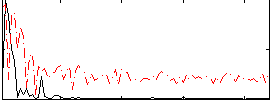 |
Correlation:0.44091
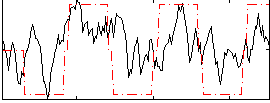 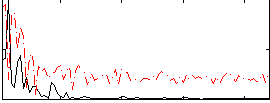 |
Component:053; Jaccard:0.084863
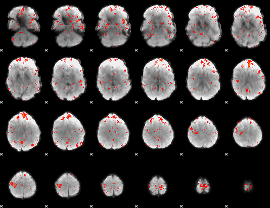 |
Component:345; Jaccard:0.095569
 |
Component:345; Jaccard:0.10023
 |
Component:345; Jaccard:0.095757
 |
Component:345; Jaccard:0.085012
 |
Component:345; Jaccard:0.06922
 |
Component:369; Jaccard:0.054457
 |
Correlation:0.27031
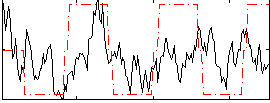 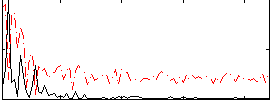 |
Correlation:0.12041
  |
Correlation:0.12041
  |
Correlation:0.12041
  |
Correlation:0.12041
  |
Correlation:0.12041
  |
Correlation:0.34739
  |
Component:082; Jaccard:0.084695
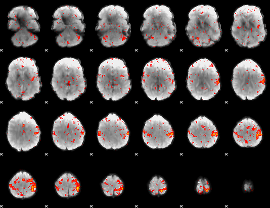 |
Component:053; Jaccard:0.094968
 |
Component:053; Jaccard:0.099182
 |
Component:369; Jaccard:0.093894
 |
Component:369; Jaccard:0.084578
 |
Component:219; Jaccard:0.068233
 |
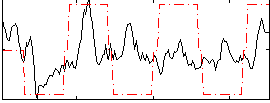
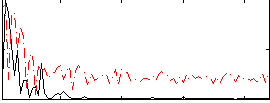
Component:034; Jaccard:0.052159
 |
Correlation:0.24236
  |
Correlation:0.27031
  |
Correlation:0.27031
  |
Correlation:0.34739
  |
Correlation:0.34739
  |
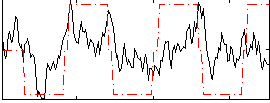
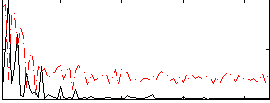
Correlation:0.44091
  |
Correlation:0.39948
  |
Cope: 04:"neg_FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
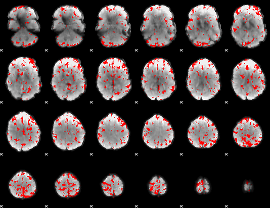 |
GLM binary result thresholded by 4
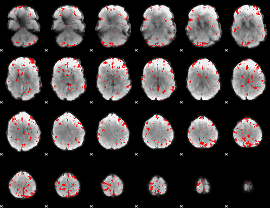 |
GLM Z-score result thresholded by 1
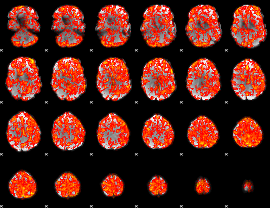 |
GLM Z-score result thresholded by 1.5
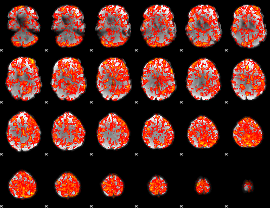 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:311; Jaccard:0.080165
 |
Component:311; Jaccard:0.094529
 |
Component:311; Jaccard:0.11202
 |
Component:311; Jaccard:0.13189
 |
Component:311; Jaccard:0.15184
 |
Component:311; Jaccard:0.16065
 |
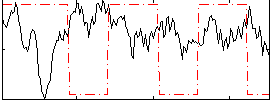
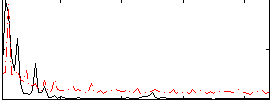
Component:311; Jaccard:0.1452
 |
Correlation:-0.25991
  |
Correlation:-0.25991
  |
Correlation:-0.25991
  |
Correlation:-0.25991
  |
Correlation:-0.25991
  |
Correlation:-0.25991
  |
Correlation:-0.25991
  |
Component:210; Jaccard:0.079359
 |
Component:358; Jaccard:0.089027
 |
Component:358; Jaccard:0.10454
 |
Component:358; Jaccard:0.12362
 |
Component:358; Jaccard:0.14146
 |
Component:358; Jaccard:0.15168
 |
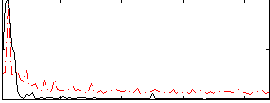
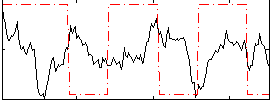
Component:358; Jaccard:0.14081
 |
Correlation:-0.27998
  |
Correlation:-0.098156
  |
Correlation:-0.098156
  |
Correlation:-0.098156
  |
Correlation:-0.098156
  |
Correlation:-0.098156
  |
Correlation:-0.098156
  |
Component:378; Jaccard:0.077102
 |
Component:378; Jaccard:0.088944
 |
Component:378; Jaccard:0.10268
 |
Component:378; Jaccard:0.1188
 |
Component:378; Jaccard:0.13125
 |
Component:378; Jaccard:0.13167
 |
Component:317; Jaccard:0.11886
 |
Correlation:-0.19137
  |
Correlation:-0.19137
  |
Correlation:-0.19137
  |
Correlation:-0.19137
  |
Correlation:-0.19137
  |
Correlation:-0.19137
  |
Correlation:0.24395
  |
Component:358; Jaccard:0.076526
 |
Component:210; Jaccard:0.08842
 |
Component:338; Jaccard:0.10001
 |
Component:338; Jaccard:0.11596
 |
Component:338; Jaccard:0.12741
 |
Component:338; Jaccard:0.12867
 |
Component:338; Jaccard:0.1172
 |
Correlation:-0.098156
  |
Correlation:-0.27998
  |
Correlation:-0.12606
  |
Correlation:-0.12606
  |
Correlation:-0.12606
  |
Correlation:-0.12606
  |
Correlation:-0.12606
  |
Component:338; Jaccard:0.075104
 |
Component:338; Jaccard:0.086736
 |
Component:210; Jaccard:0.098778
 |
Component:210; Jaccard:0.10774
 |
Component:317; Jaccard:0.11774
 |
Component:317; Jaccard:0.12442
 |
Component:378; Jaccard:0.11535
 |
Correlation:-0.12606
  |
Correlation:-0.12606
  |
Correlation:-0.27998
  |
Correlation:-0.27998
  |
Correlation:0.24395
  |
Correlation:0.24395
  |
Correlation:-0.19137
  |
Component:054; Jaccard:0.072912
 |
Component:284; Jaccard:0.083823
 |
Component:284; Jaccard:0.096608
 |
Component:317; Jaccard:0.10629
 |
Component:135; Jaccard:0.1141
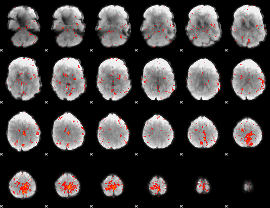 |
Component:135; Jaccard:0.11279
 |
Component:208; Jaccard:0.10379
 |
Correlation:-0.23167
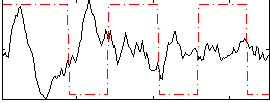 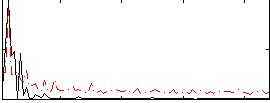 |
Correlation:-0.26494
 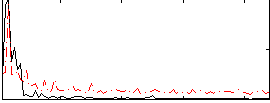 |
Correlation:-0.26494
  |
Correlation:0.24395
  |
Correlation:-0.14695
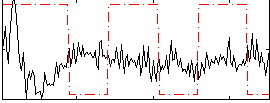  |
Correlation:-0.14695
  |
Correlation:-0.04345
  |
Component:284; Jaccard:0.0723
 |
Component:054; Jaccard:0.080478
 |
Component:317; Jaccard:0.092158
 |
Component:135; Jaccard:0.10515
 |
Component:070; Jaccard:0.1119
 |
Component:070; Jaccard:0.11261
 |
Component:070; Jaccard:0.098976
 |
Correlation:-0.26494
  |
Correlation:-0.23167
  |
Correlation:0.24395
  |
Correlation:-0.14695
  |
Correlation:-0.16673
  |
Correlation:-0.16673
  |
Correlation:-0.16673
  |
Component:082; Jaccard:0.070241
 |
Component:317; Jaccard:0.080042
 |
Component:135; Jaccard:0.091248
 |
Component:284; Jaccard:0.10397
 |
Component:210; Jaccard:0.10917
 |
Component:208; Jaccard:0.11059
 |
Component:390; Jaccard:0.098387
 |
Correlation:-0.16812
  |
Correlation:0.24395
  |
Correlation:-0.14695
  |
Correlation:-0.26494
  |
Correlation:-0.27998
  |
Correlation:-0.04345
  |
Correlation:-0.093505
  |
Component:317; Jaccard:0.069982
 |
Component:070; Jaccard:0.079231
 |
Component:070; Jaccard:0.090126
 |
Component:070; Jaccard:0.10199
 |
Component:208; Jaccard:0.10646
 |
Component:066; Jaccard:0.1045
 |
Component:066; Jaccard:0.096869
 |
Correlation:0.24395
  |
Correlation:-0.16673
  |
Correlation:-0.16673
  |
Correlation:-0.16673
  |
Correlation:-0.04345
  |
Correlation:0.13698
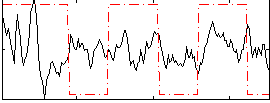 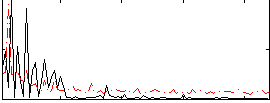 |
Correlation:0.13698
  |
Component:260; Jaccard:0.06907
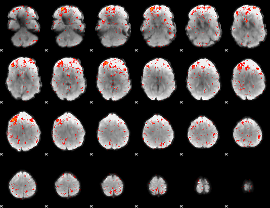 |
Component:260; Jaccard:0.078709
 |
Component:260; Jaccard:0.088672
 |
Component:260; Jaccard:0.098462
 |
Component:284; Jaccard:0.10645
 |
Component:390; Jaccard:0.1035
 |
Component:135; Jaccard:0.095869
 |
Correlation:0.02222
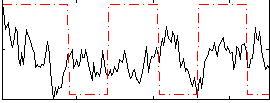  |
Correlation:0.02222
  |
Correlation:0.02222
  |
Correlation:0.02222
  |
Correlation:-0.26494
  |
Correlation:-0.093505
  |
Correlation:-0.14695
  |
Cope: 05:"neg_SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
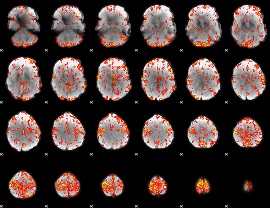 |
GLM Z-score result thresholded by 4
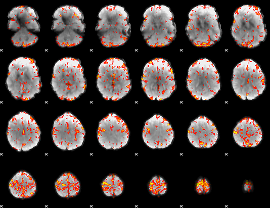 |
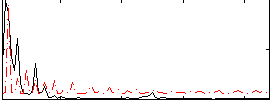
Component:210; Jaccard:0.077537
 |
Component:311; Jaccard:0.087432
 |
Component:311; Jaccard:0.10282
 |
Component:311; Jaccard:0.12176
 |
Component:311; Jaccard:0.14423
 |
Component:311; Jaccard:0.16787
 |
Component:311; Jaccard:0.18692
 |
Correlation:0.42183
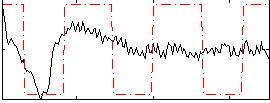  |
Correlation:0.40265
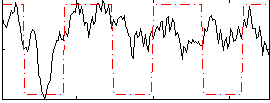  |
Correlation:0.40265
  |
Correlation:0.40265
  |
Correlation:0.40265
  |
Correlation:0.40265
  |
Correlation:0.40265
  |
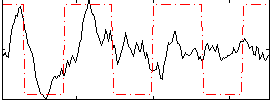
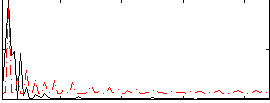
Component:311; Jaccard:0.075619
 |
Component:210; Jaccard:0.086766
 |
Component:054; Jaccard:0.10114
 |
Component:054; Jaccard:0.1189
 |
Component:054; Jaccard:0.13758
 |
Component:054; Jaccard:0.1561
 |
Component:054; Jaccard:0.16998
 |
Correlation:0.40265
  |
Correlation:0.42183
  |
Correlation:0.36613
  |
Correlation:0.36613
  |
Correlation:0.36613
  |
Correlation:0.36613
  |
Correlation:0.36613
  |
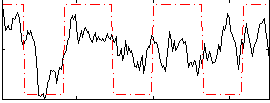
Component:054; Jaccard:0.075412
 |
Component:054; Jaccard:0.086625
 |
Component:378; Jaccard:0.098222
 |
Component:378; Jaccard:0.1141
 |
Component:378; Jaccard:0.13191
 |
Component:378; Jaccard:0.14974
 |
Component:378; Jaccard:0.16206
 |
Correlation:0.36613
  |
Correlation:0.36613
  |
Correlation:0.4167
  |
Correlation:0.4167
  |
Correlation:0.4167
  |
Correlation:0.4167
  |
Correlation:0.4167
  |
Component:378; Jaccard:0.073865
 |
Component:378; Jaccard:0.084759
 |
Component:210; Jaccard:0.097093
 |
Component:284; Jaccard:0.11181
 |
Component:284; Jaccard:0.12792
 |
Component:284; Jaccard:0.13994
 |
Component:369; Jaccard:0.14954
 |
Correlation:0.4167
  |
Correlation:0.4167
  |
Correlation:0.42183
  |
Correlation:0.45023
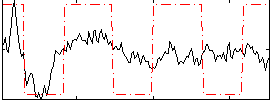  |
Correlation:0.45023
  |
Correlation:0.45023
  |
Correlation:0.39146
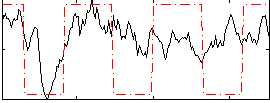 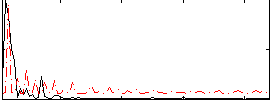 |
Component:082; Jaccard:0.070838
 |
Component:284; Jaccard:0.08097
 |
Component:284; Jaccard:0.0955
 |
Component:210; Jaccard:0.1089
 |
Component:210; Jaccard:0.11937
 |
Component:369; Jaccard:0.13519
 |
Component:284; Jaccard:0.14476
 |
Correlation:0.27869
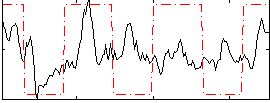 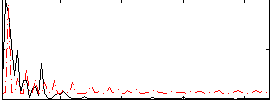 |
Correlation:0.45023
  |
Correlation:0.45023
  |
Correlation:0.42183
  |
Correlation:0.42183
  |
Correlation:0.39146
  |
Correlation:0.45023
  |
Component:338; Jaccard:0.070575
 |
Component:338; Jaccard:0.079937
 |
Component:338; Jaccard:0.091523
 |
Component:338; Jaccard:0.10399
 |
Component:369; Jaccard:0.11743
 |
Component:118; Jaccard:0.12799
 |
Component:118; Jaccard:0.14045
 |
Correlation:0.35803
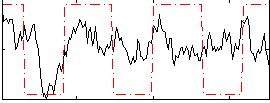 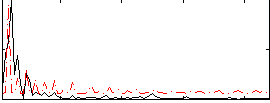 |
Correlation:0.35803
  |
Correlation:0.35803
  |
Correlation:0.35803
  |
Correlation:0.39146
  |
Correlation:0.50918
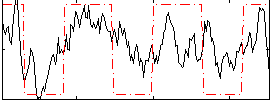  |
Correlation:0.50918
  |
Component:284; Jaccard:0.069896
 |
Component:082; Jaccard:0.079749
 |
Component:082; Jaccard:0.09026
 |
Component:082; Jaccard:0.10137
 |
Component:338; Jaccard:0.11681
 |
Component:338; Jaccard:0.12758
 |
Component:338; Jaccard:0.13276
 |
Correlation:0.45023
  |
Correlation:0.27869
  |
Correlation:0.27869
  |
Correlation:0.27869
  |
Correlation:0.35803
  |
Correlation:0.35803
  |
Correlation:0.35803
  |
Component:350; Jaccard:0.068815
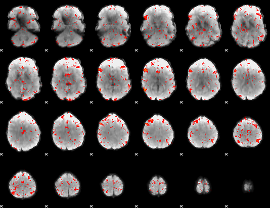 |
Component:350; Jaccard:0.077206
 |
Component:350; Jaccard:0.086566
 |
Component:369; Jaccard:0.10066
 |
Component:118; Jaccard:0.11282
 |
Component:210; Jaccard:0.12659
 |
Component:210; Jaccard:0.12642
 |
Correlation:0.34122
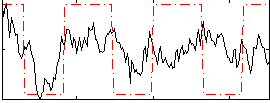  |
Correlation:0.34122
  |
Correlation:0.34122
  |
Correlation:0.39146
  |
Correlation:0.50918
  |
Correlation:0.42183
  |
Correlation:0.42183
  |
Component:358; Jaccard:0.066583
 |
Component:070; Jaccard:0.075129
 |
Component:369; Jaccard:0.086401
 |
Component:118; Jaccard:0.097762
 |
Component:082; Jaccard:0.11136
 |
Component:287; Jaccard:0.1182
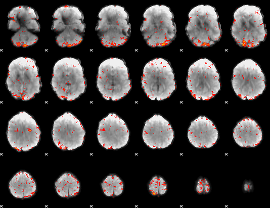 |
Component:287; Jaccard:0.12529
 |
Correlation:0.29296
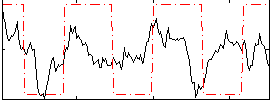  |
Correlation:0.29509
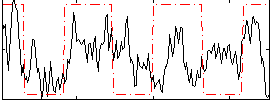 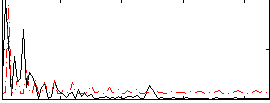 |
Correlation:0.39146
  |
Correlation:0.50918
  |
Correlation:0.27869
  |
Correlation:0.37933
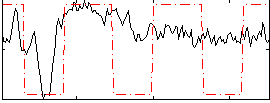 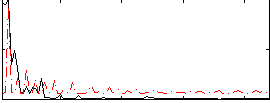 |
Correlation:0.37933
  |
Component:070; Jaccard:0.066538
 |
Component:369; Jaccard:0.07463
 |
Component:070; Jaccard:0.085016
 |
Component:350; Jaccard:0.097032
 |
Component:070; Jaccard:0.107
 |
Component:082; Jaccard:0.11729
 |
Component:070; Jaccard:0.11826
 |
Correlation:0.29509
  |
Correlation:0.39146
  |
Correlation:0.29509
  |
Correlation:0.34122
  |
Correlation:0.29509
  |
Correlation:0.27869
  |
Correlation:0.29509
  |
Cope: 06:"SHAPES-FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
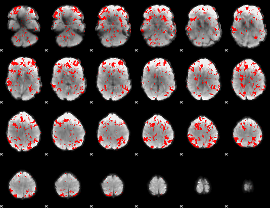 |
GLM binary result thresholded by 2
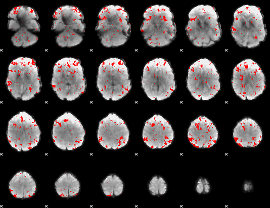 |
GLM binary result thresholded by 2.5
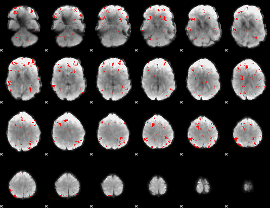 |
GLM binary result thresholded by 3
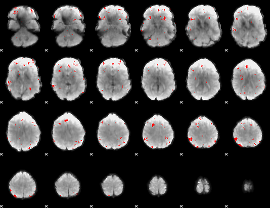 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
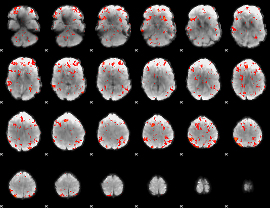 |
GLM Z-score result thresholded by 2.5
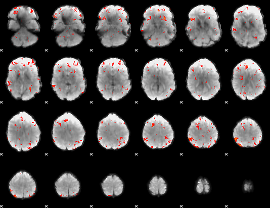 |
GLM Z-score result thresholded by 3
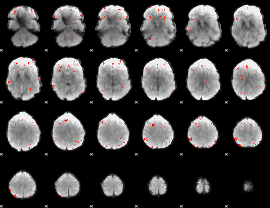 |
GLM Z-score result thresholded by 3.5
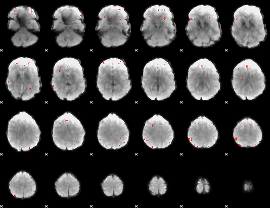 |
GLM Z-score result thresholded by 4
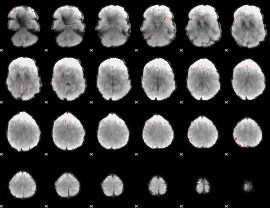 |
Component:309; Jaccard:0.10774
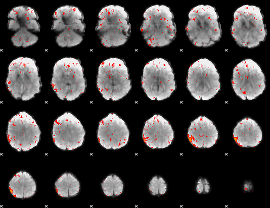 |
Component:309; Jaccard:0.13528
 |
Component:309; Jaccard:0.15618
 |
Component:309; Jaccard:0.15354
 |
Component:309; Jaccard:0.12154
 |
Component:309; Jaccard:0.070981
 |
Component:309; Jaccard:0.036797
 |
Correlation:0.26742
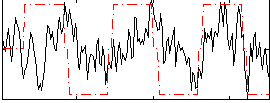 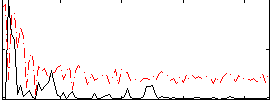 |
Correlation:0.26742
  |
Correlation:0.26742
  |
Correlation:0.26742
  |
Correlation:0.26742
  |
Correlation:0.26742
  |
Correlation:0.26742
  |
Component:358; Jaccard:0.099347
 |
Component:357; Jaccard:0.10626
 |
Component:357; Jaccard:0.10265
 |
Component:288; Jaccard:0.083101
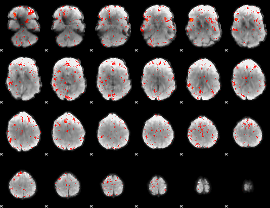 |
Component:288; Jaccard:0.055998
 |
Component:288; Jaccard:0.027997
 |
Component:288; Jaccard:0.013047
 |
Correlation:-0.21215
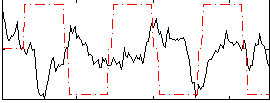 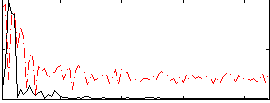 |
Correlation:-0.054858
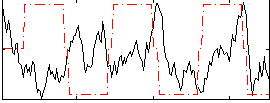  |
Correlation:-0.054858
  |
Correlation:-0.12291
  |
Correlation:-0.12291
  |
Correlation:-0.12291
  |
Correlation:-0.12291
  |
Component:357; Jaccard:0.097337
 |
Component:358; Jaccard:0.10356
 |
Component:288; Jaccard:0.097695
 |
Component:357; Jaccard:0.082813
 |
Component:357; Jaccard:0.052179
 |
Component:357; Jaccard:0.027763
 |
Component:330; Jaccard:0.011591
 |
Correlation:-0.054858
  |
Correlation:-0.21215
  |
Correlation:-0.12291
  |
Correlation:-0.054858
  |
Correlation:-0.054858
  |
Correlation:-0.054858
  |
Correlation:-0.06639
  |
Component:288; Jaccard:0.095452
 |
Component:288; Jaccard:0.10116
 |
Component:398; Jaccard:0.095839
 |
Component:358; Jaccard:0.075699
 |
Component:398; Jaccard:0.04887
 |
Component:398; Jaccard:0.025852
 |
Component:398; Jaccard:0.010171
 |
Correlation:-0.12291
  |
Correlation:-0.12291
  |
Correlation:0.31504
  |
Correlation:-0.21215
  |
Correlation:0.31504
  |
Correlation:0.31504
  |
Correlation:0.31504
  |
Component:398; Jaccard:0.092437
 |
Component:398; Jaccard:0.098842
 |
Component:358; Jaccard:0.095484
 |
Component:398; Jaccard:0.074036
 |
Component:358; Jaccard:0.044479
 |
Component:330; Jaccard:0.023146
 |
Component:357; Jaccard:0.009786
 |
Correlation:0.31504
  |
Correlation:0.31504
  |
Correlation:-0.21215
  |
Correlation:0.31504
  |
Correlation:-0.21215
  |
Correlation:-0.06639
  |
Correlation:-0.054858
  |
Component:011; Jaccard:0.085128
 |
Component:154; Jaccard:0.092212
 |
Component:154; Jaccard:0.08714
 |
Component:011; Jaccard:0.067189
 |
Component:011; Jaccard:0.044463
 |
Component:011; Jaccard:0.0231
 |
Component:011; Jaccard:0.009646
 |
Correlation:0.116
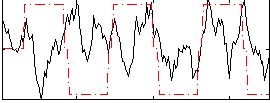 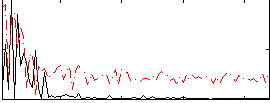 |
Correlation:0.22718
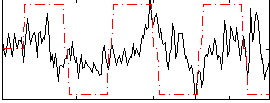 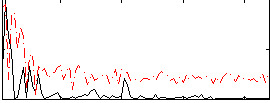 |
Correlation:0.22718
  |
Correlation:0.116
  |
Correlation:0.116
  |
Correlation:0.116
  |
Correlation:0.116
  |
Component:154; Jaccard:0.084094
 |
Component:011; Jaccard:0.090546
 |
Component:011; Jaccard:0.085778
 |
Component:205; Jaccard:0.066419
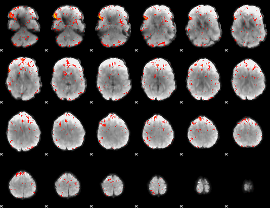 |
Component:205; Jaccard:0.042176
 |
Component:205; Jaccard:0.021821
 |
Component:081; Jaccard:0.009172
 |
Correlation:0.22718
  |
Correlation:0.116
  |
Correlation:0.116
  |
Correlation:-0.16265
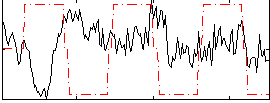  |
Correlation:-0.16265
  |
Correlation:-0.16265
  |
Correlation:-0.032909
  |
Component:317; Jaccard:0.083087
 |
Component:181; Jaccard:0.086182
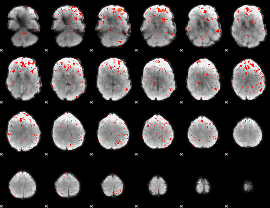 |
Component:181; Jaccard:0.081487
 |
Component:154; Jaccard:0.06585
 |
Component:154; Jaccard:0.039867
 |
Component:185; Jaccard:0.021806
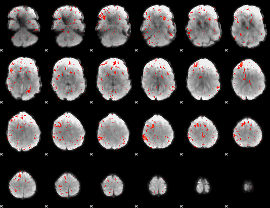 |
Component:205; Jaccard:0.009081
 |
Correlation:0.14219
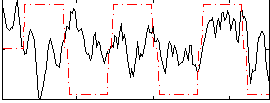 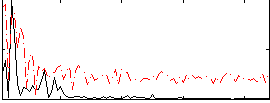 |
Correlation:0.2585
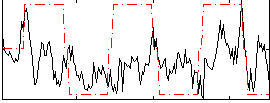  |
Correlation:0.2585
  |
Correlation:0.22718
  |
Correlation:0.22718
  |
Correlation:0.061668
  |
Correlation:-0.16265
  |
Component:081; Jaccard:0.082626
 |
Component:317; Jaccard:0.084851
 |
Component:205; Jaccard:0.078606
 |
Component:181; Jaccard:0.061428
 |
Component:330; Jaccard:0.039638
 |
Component:358; Jaccard:0.021107
 |
Component:181; Jaccard:0.00904
 |
Correlation:-0.032909
  |
Correlation:0.14219
  |
Correlation:-0.16265
  |
Correlation:0.2585
  |
Correlation:-0.06639
  |
Correlation:-0.21215
  |
Correlation:0.2585
  |
Component:181; Jaccard:0.082051
 |
Component:205; Jaccard:0.083376
 |
Component:317; Jaccard:0.077427
 |
Component:317; Jaccard:0.060278
 |
Component:181; Jaccard:0.039635
 |
Component:181; Jaccard:0.020187
 |
Component:185; Jaccard:0.008868
 |
Correlation:0.2585
  |
Correlation:-0.16265
  |
Correlation:0.14219
  |
Correlation:0.14219
  |
Correlation:0.2585
  |
Correlation:0.2585
  |
Correlation:0.061668
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































