Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
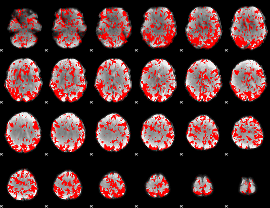 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
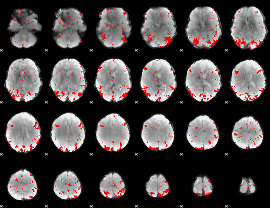 |
GLM binary result thresholded by 3.5
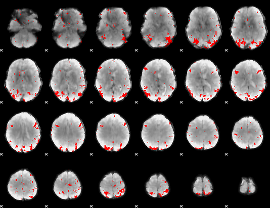 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
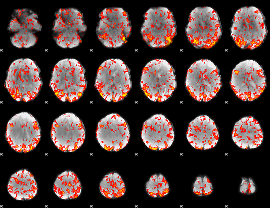 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
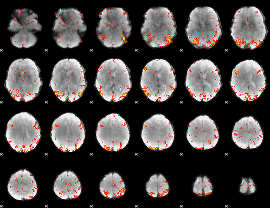 |
GLM Z-score result thresholded by 3.5
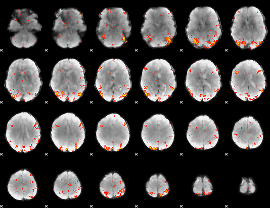 |
GLM Z-score result thresholded by 4
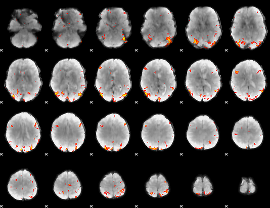 |
Component:228; Jaccard:0.12608
 |
Component:228; Jaccard:0.15614
 |
Component:228; Jaccard:0.1846
 |
Component:228; Jaccard:0.20442
 |
Component:329; Jaccard:0.22046
 |
Component:329; Jaccard:0.21727
 |
Component:329; Jaccard:0.19852
 |
Correlation:0.55497
  |
Correlation:0.55497
  |
Correlation:0.55497
  |
Correlation:0.55497
  |
Correlation:0.32479
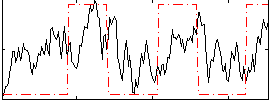 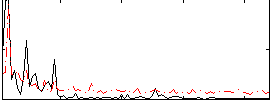 |
Correlation:0.32479
  |
Correlation:0.32479
  |
Component:329; Jaccard:0.11446
 |
Component:329; Jaccard:0.14206
 |
Component:329; Jaccard:0.17092
 |
Component:329; Jaccard:0.20167
 |
Component:228; Jaccard:0.20632
 |
Component:228; Jaccard:0.19326
 |
Component:228; Jaccard:0.16596
 |
Correlation:0.32479
  |
Correlation:0.32479
  |
Correlation:0.32479
  |
Correlation:0.32479
  |
Correlation:0.55497
  |
Correlation:0.55497
  |
Correlation:0.55497
  |
Component:297; Jaccard:0.11438
 |
Component:297; Jaccard:0.13925
 |
Component:297; Jaccard:0.16266
 |
Component:297; Jaccard:0.17775
 |
Component:297; Jaccard:0.18158
 |
Component:297; Jaccard:0.16848
 |
Component:111; Jaccard:0.14458
 |
Correlation:0.50037
  |
Correlation:0.50037
  |
Correlation:0.50037
  |
Correlation:0.50037
  |
Correlation:0.50037
  |
Correlation:0.50037
  |
Correlation:0.57447
  |
Component:040; Jaccard:0.10878
 |
Component:068; Jaccard:0.1329
 |
Component:068; Jaccard:0.15861
 |
Component:068; Jaccard:0.17655
 |
Component:068; Jaccard:0.17799
 |
Component:068; Jaccard:0.16438
 |
Component:297; Jaccard:0.14085
 |
Correlation:0.41985
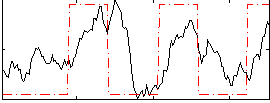  |
Correlation:0.23464
  |
Correlation:0.23464
  |
Correlation:0.23464
  |
Correlation:0.23464
  |
Correlation:0.23464
  |
Correlation:0.50037
  |
Component:068; Jaccard:0.10759
 |
Component:040; Jaccard:0.12688
 |
Component:040; Jaccard:0.14444
 |
Component:040; Jaccard:0.15604
 |
Component:111; Jaccard:0.15785
 |
Component:111; Jaccard:0.1582
 |
Component:068; Jaccard:0.13399
 |
Correlation:0.23464
  |
Correlation:0.41985
  |
Correlation:0.41985
  |
Correlation:0.41985
  |
Correlation:0.57447
  |
Correlation:0.57447
  |
Correlation:0.23464
  |
Component:292; Jaccard:0.10129
 |
Component:198; Jaccard:0.10807
 |
Component:111; Jaccard:0.12336
 |
Component:111; Jaccard:0.14399
 |
Component:040; Jaccard:0.15593
 |
Component:040; Jaccard:0.14465
 |
Component:040; Jaccard:0.12425
 |
Correlation:0.069799
 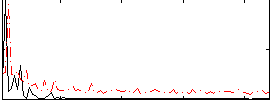 |
Correlation:0.23854
  |
Correlation:0.57447
  |
Correlation:0.57447
  |
Correlation:0.41985
  |
Correlation:0.41985
  |
Correlation:0.41985
  |
Component:198; Jaccard:0.091342
 |
Component:292; Jaccard:0.10785
 |
Component:198; Jaccard:0.12078
 |
Component:198; Jaccard:0.13139
 |
Component:198; Jaccard:0.13309
 |
Component:198; Jaccard:0.12297
 |
Component:198; Jaccard:0.10483
 |
Correlation:0.23854
  |
Correlation:0.069799
  |
Correlation:0.23854
  |
Correlation:0.23854
  |
Correlation:0.23854
  |
Correlation:0.23854
  |
Correlation:0.23854
  |
Component:300; Jaccard:0.09131
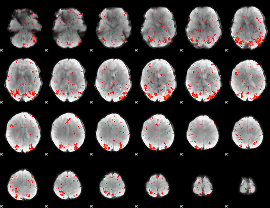 |
Component:300; Jaccard:0.1039
 |
Component:300; Jaccard:0.11603
 |
Component:300; Jaccard:0.1249
 |
Component:300; Jaccard:0.1246
 |
Component:300; Jaccard:0.11689
 |
Component:300; Jaccard:0.10034
 |
Correlation:0.24502
  |
Correlation:0.24502
  |
Correlation:0.24502
  |
Correlation:0.24502
  |
Correlation:0.24502
  |
Correlation:0.24502
  |
Correlation:0.24502
  |
Component:113; Jaccard:0.090936
 |
Component:113; Jaccard:0.10238
 |
Component:113; Jaccard:0.11153
 |
Component:113; Jaccard:0.11734
 |
Component:113; Jaccard:0.11776
 |
Component:113; Jaccard:0.10784
 |
Component:113; Jaccard:0.091225
 |
Correlation:0.3461
  |
Correlation:0.3461
  |
Correlation:0.3461
  |
Correlation:0.3461
  |
Correlation:0.3461
  |
Correlation:0.3461
  |
Correlation:0.3461
  |
Component:319; Jaccard:0.090746
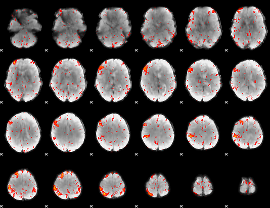 |
Component:111; Jaccard:0.10214
 |
Component:292; Jaccard:0.10816
 |
Component:292; Jaccard:0.10371
 |
Component:372; Jaccard:0.097954
 |
Component:372; Jaccard:0.094325
 |
Component:372; Jaccard:0.081668
 |
Correlation:0.3027
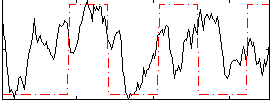  |
Correlation:0.57447
  |
Correlation:0.069799
  |
Correlation:0.069799
  |
Correlation:0.30212
  |
Correlation:0.30212
  |
Correlation:0.30212
  |
Cope: 02:"SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
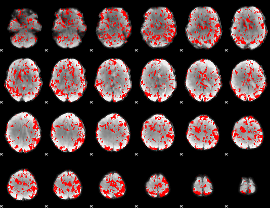 |
GLM binary result thresholded by 1.5
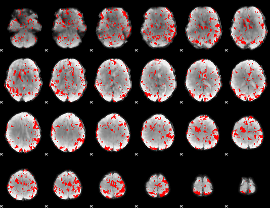 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:068; Jaccard:0.12127
 |
Component:068; Jaccard:0.14012
 |
Component:068; Jaccard:0.14812
 |
Component:068; Jaccard:0.13999
 |
Component:068; Jaccard:0.10808
 |
Component:068; Jaccard:0.067585
 |
Component:366; Jaccard:0.040638
 |
Correlation:0.056465
  |
Correlation:0.056465
  |
Correlation:0.056465
  |
Correlation:0.056465
  |
Correlation:0.056465
  |
Correlation:0.056465
  |
Correlation:0.27244
  |
Component:311; Jaccard:0.11213
 |
Component:311; Jaccard:0.125
 |
Component:311; Jaccard:0.12608
 |
Component:311; Jaccard:0.11223
 |
Component:366; Jaccard:0.08675
 |
Component:366; Jaccard:0.065239
 |
Component:068; Jaccard:0.032769
 |
Correlation:0.052585
  |
Correlation:0.052585
  |
Correlation:0.052585
  |
Correlation:0.052585
  |
Correlation:0.27244
  |
Correlation:0.27244
  |
Correlation:0.056465
  |
Component:292; Jaccard:0.097777
 |
Component:257; Jaccard:0.10369
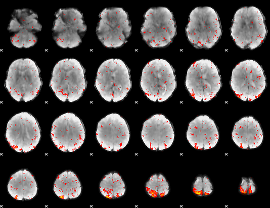 |
Component:257; Jaccard:0.10805
 |
Component:257; Jaccard:0.0991
 |
Component:311; Jaccard:0.082621
 |
Component:257; Jaccard:0.054344
 |
Component:257; Jaccard:0.030182
 |
Correlation:0.15403
  |
Correlation:0.26631
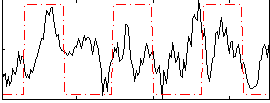 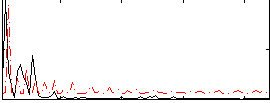 |
Correlation:0.26631
  |
Correlation:0.26631
  |
Correlation:0.052585
  |
Correlation:0.26631
  |
Correlation:0.26631
  |
Component:257; Jaccard:0.093478
 |
Component:292; Jaccard:0.096207
 |
Component:366; Jaccard:0.1009
 |
Component:366; Jaccard:0.09821
 |
Component:257; Jaccard:0.081463
 |
Component:311; Jaccard:0.048993
 |
Component:329; Jaccard:0.02359
 |
Correlation:0.26631
  |
Correlation:0.15403
  |
Correlation:0.27244
  |
Correlation:0.27244
  |
Correlation:0.26631
  |
Correlation:0.052585
  |
Correlation:-0.20025
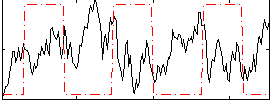  |
Component:043; Jaccard:0.086779
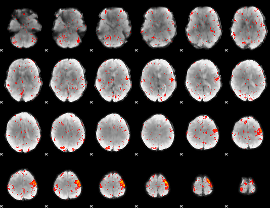 |
Component:366; Jaccard:0.094161
 |
Component:337; Jaccard:0.092889
 |
Component:329; Jaccard:0.085869
 |
Component:329; Jaccard:0.068764
 |
Component:329; Jaccard:0.044563
 |
Component:311; Jaccard:0.022765
 |
Correlation:0.27066
 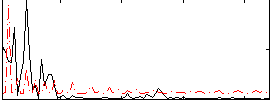 |
Correlation:0.27244
  |
Correlation:-0.068801
  |
Correlation:-0.20025
  |
Correlation:-0.20025
  |
Correlation:-0.20025
  |
Correlation:0.052585
  |
Component:329; Jaccard:0.086431
 |
Component:329; Jaccard:0.091844
 |
Component:329; Jaccard:0.091736
 |
Component:337; Jaccard:0.079213
 |
Component:337; Jaccard:0.056078
 |
Component:043; Jaccard:0.035578
 |
Component:043; Jaccard:0.021532
 |
Correlation:-0.20025
  |
Correlation:-0.20025
  |
Correlation:-0.20025
  |
Correlation:-0.068801
  |
Correlation:-0.068801
  |
Correlation:0.27066
  |
Correlation:0.27066
  |
Component:366; Jaccard:0.082001
 |
Component:043; Jaccard:0.091561
 |
Component:043; Jaccard:0.08794
 |
Component:265; Jaccard:0.075968
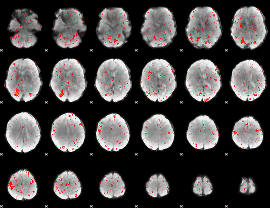 |
Component:265; Jaccard:0.055597
 |
Component:265; Jaccard:0.031976
 |
Component:018; Jaccard:0.018055
 |
Correlation:0.27244
  |
Correlation:0.27066
  |
Correlation:0.27066
  |
Correlation:-0.021003
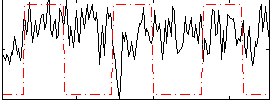  |
Correlation:-0.021003
  |
Correlation:-0.021003
  |
Correlation:0.11578
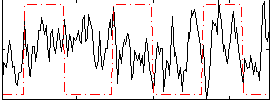  |
Component:082; Jaccard:0.078799
 |
Component:337; Jaccard:0.08632
 |
Component:265; Jaccard:0.084436
 |
Component:043; Jaccard:0.075147
 |
Component:043; Jaccard:0.054573
 |
Component:337; Jaccard:0.030801
 |
Component:198; Jaccard:0.016379
 |
Correlation:0.13765
  |
Correlation:-0.068801
  |
Correlation:-0.021003
  |
Correlation:0.27066
  |
Correlation:0.27066
  |
Correlation:-0.068801
  |
Correlation:-0.12303
  |
Component:228; Jaccard:0.076525
 |
Component:265; Jaccard:0.082181
 |
Component:292; Jaccard:0.083468
 |
Component:082; Jaccard:0.064268
 |
Component:018; Jaccard:0.048772
 |
Component:018; Jaccard:0.030068
 |
Component:082; Jaccard:0.014634
 |
Correlation:-0.52605
  |
Correlation:-0.021003
  |
Correlation:0.15403
  |
Correlation:0.13765
  |
Correlation:0.11578
  |
Correlation:0.11578
  |
Correlation:0.13765
  |
Component:265; Jaccard:0.075264
 |
Component:082; Jaccard:0.081863
 |
Component:082; Jaccard:0.076664
 |
Component:292; Jaccard:0.063189
 |
Component:082; Jaccard:0.045654
 |
Component:198; Jaccard:0.028617
 |
Component:265; Jaccard:0.013924
 |
Correlation:-0.021003
  |
Correlation:0.13765
  |
Correlation:0.13765
  |
Correlation:0.15403
  |
Correlation:0.13765
  |
Correlation:-0.12303
  |
Correlation:-0.021003
  |
Cope: 03:"FACES-SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
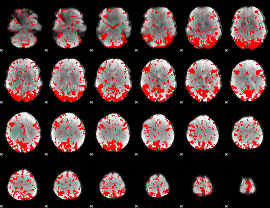 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
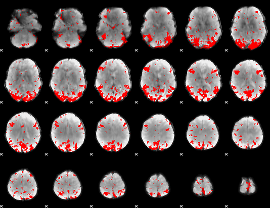 |
GLM binary result thresholded by 3.5
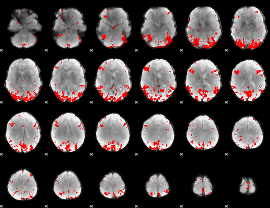 |
GLM binary result thresholded by 4
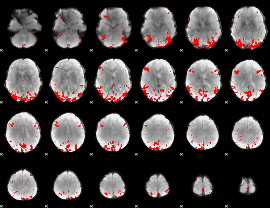 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
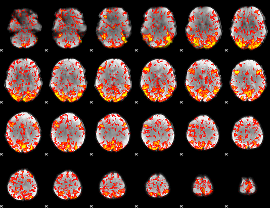 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
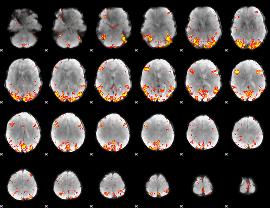 |
GLM Z-score result thresholded by 4
 |
Component:040; Jaccard:0.11654
 |
Component:040; Jaccard:0.15552
 |
Component:040; Jaccard:0.20283
 |
Component:040; Jaccard:0.25053
 |
Component:040; Jaccard:0.28148
 |
Component:040; Jaccard:0.29245
 |
Component:040; Jaccard:0.2798
 |
Correlation:0.35985
  |
Correlation:0.35985
  |
Correlation:0.35985
  |
Correlation:0.35985
  |
Correlation:0.35985
  |
Correlation:0.35985
  |
Correlation:0.35985
  |
Component:005; Jaccard:0.10626
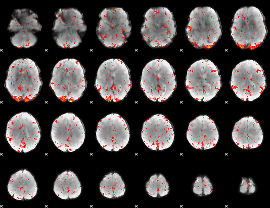 |
Component:228; Jaccard:0.13752
 |
Component:228; Jaccard:0.17403
 |
Component:228; Jaccard:0.20624
 |
Component:228; Jaccard:0.22678
 |
Component:228; Jaccard:0.23221
 |
Component:228; Jaccard:0.22879
 |
Correlation:0.51024
  |
Correlation:0.58636
  |
Correlation:0.58636
  |
Correlation:0.58636
  |
Correlation:0.58636
  |
Correlation:0.58636
  |
Correlation:0.58636
  |
Component:228; Jaccard:0.10538
 |
Component:005; Jaccard:0.12992
 |
Component:297; Jaccard:0.1579
 |
Component:297; Jaccard:0.18214
 |
Component:297; Jaccard:0.19552
 |
Component:393; Jaccard:0.20478
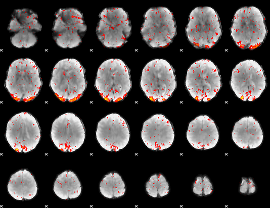 |
Component:393; Jaccard:0.20519
 |
Correlation:0.58636
  |
Correlation:0.51024
  |
Correlation:0.438
  |
Correlation:0.438
  |
Correlation:0.438
  |
Correlation:0.56626
  |
Correlation:0.56626
  |
Component:297; Jaccard:0.099922
 |
Component:297; Jaccard:0.12801
 |
Component:005; Jaccard:0.15179
 |
Component:393; Jaccard:0.17239
 |
Component:393; Jaccard:0.19338
 |
Component:297; Jaccard:0.19439
 |
Component:111; Jaccard:0.19322
 |
Correlation:0.438
  |
Correlation:0.438
  |
Correlation:0.51024
  |
Correlation:0.56626
  |
Correlation:0.56626
  |
Correlation:0.438
  |
Correlation:0.53804
  |
Component:393; Jaccard:0.093771
 |
Component:393; Jaccard:0.11868
 |
Component:393; Jaccard:0.14613
 |
Component:329; Jaccard:0.1678
 |
Component:329; Jaccard:0.18339
 |
Component:111; Jaccard:0.18982
 |
Component:329; Jaccard:0.18792
 |
Correlation:0.56626
  |
Correlation:0.56626
  |
Correlation:0.56626
  |
Correlation:0.28479
  |
Correlation:0.28479
  |
Correlation:0.53804
  |
Correlation:0.28479
  |
Component:329; Jaccard:0.092732
 |
Component:329; Jaccard:0.11735
 |
Component:329; Jaccard:0.14387
 |
Component:005; Jaccard:0.16643
 |
Component:111; Jaccard:0.17644
 |
Component:329; Jaccard:0.1882
 |
Component:297; Jaccard:0.1856
 |
Correlation:0.28479
  |
Correlation:0.28479
  |
Correlation:0.28479
  |
Correlation:0.51024
  |
Correlation:0.53804
  |
Correlation:0.28479
  |
Correlation:0.438
  |
Component:363; Jaccard:0.091909
 |
Component:381; Jaccard:0.1086
 |
Component:113; Jaccard:0.1297
 |
Component:111; Jaccard:0.15316
 |
Component:005; Jaccard:0.17309
 |
Component:005; Jaccard:0.17118
 |
Component:005; Jaccard:0.15822
 |
Correlation:0.0088294
  |
Correlation:0.31021
  |
Correlation:0.29743
  |
Correlation:0.53804
  |
Correlation:0.51024
  |
Correlation:0.51024
  |
Correlation:0.51024
  |
Component:319; Jaccard:0.089986
 |
Component:113; Jaccard:0.10749
 |
Component:381; Jaccard:0.12813
 |
Component:113; Jaccard:0.14962
 |
Component:113; Jaccard:0.15891
 |
Component:113; Jaccard:0.1579
 |
Component:113; Jaccard:0.15234
 |
Correlation:0.27284
  |
Correlation:0.29743
  |
Correlation:0.31021
  |
Correlation:0.29743
  |
Correlation:0.29743
  |
Correlation:0.29743
  |
Correlation:0.29743
  |
Component:381; Jaccard:0.089065
 |
Component:300; Jaccard:0.10285
 |
Component:111; Jaccard:0.12423
 |
Component:381; Jaccard:0.14311
 |
Component:381; Jaccard:0.14839
 |
Component:162; Jaccard:0.14708
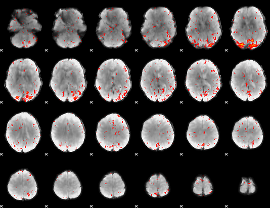 |
Component:162; Jaccard:0.14774
 |
Correlation:0.31021
  |
Correlation:0.17093
  |
Correlation:0.53804
  |
Correlation:0.31021
  |
Correlation:0.31021
  |
Correlation:0.46171
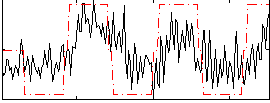 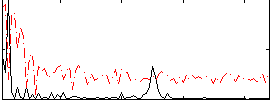 |
Correlation:0.46171
  |
Component:113; Jaccard:0.08698
 |
Component:319; Jaccard:0.10075
 |
Component:300; Jaccard:0.11923
 |
Component:300; Jaccard:0.13063
 |
Component:162; Jaccard:0.13867
 |
Component:381; Jaccard:0.14533
 |
Component:098; Jaccard:0.14135
 |
Correlation:0.29743
  |
Correlation:0.27284
  |
Correlation:0.17093
  |
Correlation:0.17093
  |
Correlation:0.46171
  |
Correlation:0.31021
  |
Correlation:0.39757
 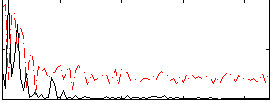 |
Cope: 04:"neg_FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
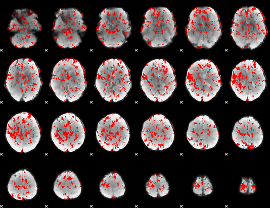 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:205; Jaccard:0.089679
 |
Component:103; Jaccard:0.092002
 |
Component:103; Jaccard:0.10634
 |
Component:103; Jaccard:0.1073
 |
Component:103; Jaccard:0.088244
 |
Component:103; Jaccard:0.048859
 |
Component:272; Jaccard:0.027304
 |
Correlation:-0.083035
  |
Correlation:0.27494
  |
Correlation:0.27494
  |
Correlation:0.27494
  |
Correlation:0.27494
  |
Correlation:0.27494
  |
Correlation:0.19746
  |
Component:126; Jaccard:0.079556
 |
Component:205; Jaccard:0.090938
 |
Component:370; Jaccard:0.097952
 |
Component:370; Jaccard:0.095213
 |
Component:224; Jaccard:0.074668
 |
Component:272; Jaccard:0.048167
 |
Component:103; Jaccard:0.019272
 |
Correlation:0.036116
  |
Correlation:-0.083035
  |
Correlation:0.13669
  |
Correlation:0.13669
  |
Correlation:0.19457
  |
Correlation:0.19746
  |
Correlation:0.27494
  |
Component:324; Jaccard:0.079121
 |
Component:370; Jaccard:0.088128
 |
Component:224; Jaccard:0.094153
 |
Component:224; Jaccard:0.093124
 |
Component:370; Jaccard:0.071986
 |
Component:224; Jaccard:0.042292
 |
Component:127; Jaccard:0.018032
 |
Correlation:-0.085286
  |
Correlation:0.13669
  |
Correlation:0.19457
  |
Correlation:0.19457
  |
Correlation:0.13669
  |
Correlation:0.19457
  |
Correlation:0.2868
  |
Component:321; Jaccard:0.078392
 |
Component:224; Jaccard:0.086313
 |
Component:321; Jaccard:0.085017
 |
Component:104; Jaccard:0.07838
 |
Component:272; Jaccard:0.063377
 |
Component:370; Jaccard:0.038791
 |
Component:104; Jaccard:0.016807
 |
Correlation:0.063006
  |
Correlation:0.19457
  |
Correlation:0.063006
  |
Correlation:0.22287
  |
Correlation:0.19746
  |
Correlation:0.13669
  |
Correlation:0.22287
  |
Component:209; Jaccard:0.075454
 |
Component:321; Jaccard:0.084677
 |
Component:205; Jaccard:0.084697
 |
Component:321; Jaccard:0.074396
 |
Component:104; Jaccard:0.061089
 |
Component:104; Jaccard:0.035653
 |
Component:209; Jaccard:0.016472
 |
Correlation:-0.14319
  |
Correlation:0.063006
  |
Correlation:-0.083035
  |
Correlation:0.063006
  |
Correlation:0.22287
  |
Correlation:0.22287
  |
Correlation:-0.14319
  |
Component:239; Jaccard:0.075303
 |
Component:209; Jaccard:0.081889
 |
Component:104; Jaccard:0.082784
 |
Component:272; Jaccard:0.073872
 |
Component:321; Jaccard:0.054625
 |
Component:127; Jaccard:0.033592
 |
Component:224; Jaccard:0.016295
 |
Correlation:0.22043
  |
Correlation:-0.14319
  |
Correlation:0.22287
  |
Correlation:0.19746
  |
Correlation:0.063006
  |
Correlation:0.2868
  |
Correlation:0.19457
  |
Component:103; Jaccard:0.075216
 |
Component:126; Jaccard:0.079323
 |
Component:209; Jaccard:0.081643
 |
Component:209; Jaccard:0.072884
 |
Component:209; Jaccard:0.052877
 |
Component:209; Jaccard:0.032815
 |
Component:123; Jaccard:0.015968
 |
Correlation:0.27494
  |
Correlation:0.036116
  |
Correlation:-0.14319
  |
Correlation:-0.14319
  |
Correlation:-0.14319
  |
Correlation:-0.14319
  |
Correlation:-0.012098
  |
Component:370; Jaccard:0.075204
 |
Component:104; Jaccard:0.079289
 |
Component:140; Jaccard:0.073939
 |
Component:263; Jaccard:0.067875
 |
Component:263; Jaccard:0.050729
 |
Component:263; Jaccard:0.030897
 |
Component:370; Jaccard:0.014507
 |
Correlation:0.13669
  |
Correlation:0.22287
  |
Correlation:0.024857
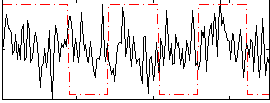  |
Correlation:0.065262
 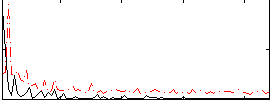 |
Correlation:0.065262
  |
Correlation:0.065262
  |
Correlation:0.13669
  |
Component:224; Jaccard:0.071865
 |
Component:324; Jaccard:0.079117
 |
Component:126; Jaccard:0.073527
 |
Component:205; Jaccard:0.067869
 |
Component:127; Jaccard:0.04629
 |
Component:321; Jaccard:0.030064
 |
Component:321; Jaccard:0.014422
 |
Correlation:0.19457
  |
Correlation:-0.085286
  |
Correlation:0.036116
  |
Correlation:-0.083035
  |
Correlation:0.2868
  |
Correlation:0.063006
  |
Correlation:0.063006
  |
Component:104; Jaccard:0.0715
 |
Component:239; Jaccard:0.074592
 |
Component:263; Jaccard:0.073097
 |
Component:140; Jaccard:0.06454
 |
Component:205; Jaccard:0.045059
 |
Component:123; Jaccard:0.026973
 |
Component:167; Jaccard:0.012678
 |
Correlation:0.22287
  |
Correlation:0.22043
  |
Correlation:0.065262
  |
Correlation:0.024857
  |
Correlation:-0.083035
  |
Correlation:-0.012098
  |
Correlation:-0.051203
  |
Cope: 05:"neg_SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
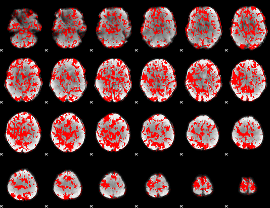 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
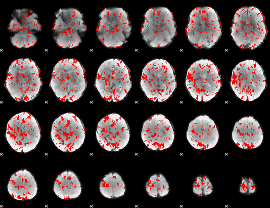 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
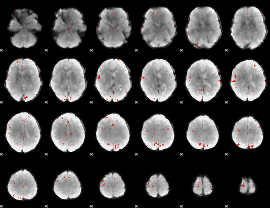 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:205; Jaccard:0.10755
 |
Component:205; Jaccard:0.12239
 |
Component:205; Jaccard:0.13279
 |
Component:205; Jaccard:0.13108
 |
Component:205; Jaccard:0.11288
 |
Component:205; Jaccard:0.083784
 |
Component:233; Jaccard:0.053739
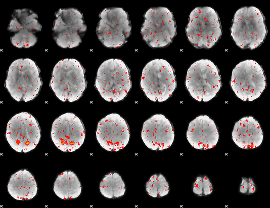 |
Correlation:0.13612
  |
Correlation:0.13612
  |
Correlation:0.13612
  |
Correlation:0.13612
  |
Correlation:0.13612
  |
Correlation:0.13612
  |
Correlation:0.20436
  |
Component:324; Jaccard:0.088633
 |
Component:381; Jaccard:0.097411
 |
Component:381; Jaccard:0.10466
 |
Component:224; Jaccard:0.10475
 |
Component:224; Jaccard:0.10101
 |
Component:209; Jaccard:0.077839
 |
Component:209; Jaccard:0.053158
 |
Correlation:-0.073963
 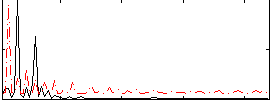 |
Correlation:0.20684
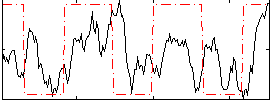 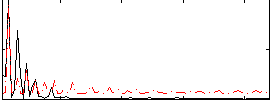 |
Correlation:0.20684
  |
Correlation:-0.0030287
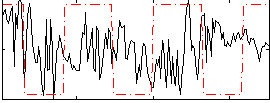  |
Correlation:-0.0030287
  |
Correlation:0.19656
 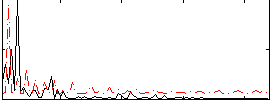 |
Correlation:0.19656
  |
Component:381; Jaccard:0.086114
 |
Component:324; Jaccard:0.097326
 |
Component:324; Jaccard:0.10307
 |
Component:381; Jaccard:0.10462
 |
Component:381; Jaccard:0.094927
 |
Component:381; Jaccard:0.076888
 |
Component:205; Jaccard:0.051105
 |
Correlation:0.20684
  |
Correlation:-0.073963
  |
Correlation:-0.073963
  |
Correlation:0.20684
  |
Correlation:0.20684
  |
Correlation:0.20684
  |
Correlation:0.13612
  |
Component:270; Jaccard:0.080969
 |
Component:209; Jaccard:0.092535
 |
Component:209; Jaccard:0.10216
 |
Component:209; Jaccard:0.10423
 |
Component:209; Jaccard:0.094253
 |
Component:233; Jaccard:0.076864
 |
Component:381; Jaccard:0.051102
 |
Correlation:0.016259
  |
Correlation:0.19656
  |
Correlation:0.19656
  |
Correlation:0.19656
  |
Correlation:0.19656
  |
Correlation:0.20436
  |
Correlation:0.20684
  |
Component:209; Jaccard:0.080684
 |
Component:270; Jaccard:0.084655
 |
Component:370; Jaccard:0.095229
 |
Component:370; Jaccard:0.10061
 |
Component:233; Jaccard:0.092335
 |
Component:224; Jaccard:0.072108
 |
Component:224; Jaccard:0.043813
 |
Correlation:0.19656
  |
Correlation:0.016259
  |
Correlation:0.055698
 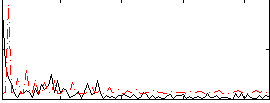 |
Correlation:0.055698
  |
Correlation:0.20436
  |
Correlation:-0.0030287
  |
Correlation:-0.0030287
  |
Component:243; Jaccard:0.074507
 |
Component:233; Jaccard:0.083343
 |
Component:224; Jaccard:0.095033
 |
Component:233; Jaccard:0.10044
 |
Component:370; Jaccard:0.091339
 |
Component:370; Jaccard:0.064507
 |
Component:324; Jaccard:0.040339
 |
Correlation:0.11655
  |
Correlation:0.20436
  |
Correlation:-0.0030287
  |
Correlation:0.20436
  |
Correlation:0.055698
  |
Correlation:0.055698
  |
Correlation:-0.073963
  |
Component:233; Jaccard:0.071616
 |
Component:224; Jaccard:0.081221
 |
Component:233; Jaccard:0.094728
 |
Component:324; Jaccard:0.098518
 |
Component:103; Jaccard:0.088196
 |
Component:324; Jaccard:0.064203
 |
Component:370; Jaccard:0.037245
 |
Correlation:0.20436
  |
Correlation:-0.0030287
  |
Correlation:0.20436
  |
Correlation:-0.073963
  |
Correlation:-0.069028
  |
Correlation:-0.073963
  |
Correlation:0.055698
  |
Component:126; Jaccard:0.069334
 |
Component:370; Jaccard:0.080804
 |
Component:103; Jaccard:0.088488
 |
Component:103; Jaccard:0.095518
 |
Component:324; Jaccard:0.085203
 |
Component:103; Jaccard:0.062022
 |
Component:103; Jaccard:0.03528
 |
Correlation:-0.04402
  |
Correlation:0.055698
  |
Correlation:-0.069028
  |
Correlation:-0.069028
  |
Correlation:-0.073963
  |
Correlation:-0.069028
  |
Correlation:-0.069028
  |
Component:097; Jaccard:0.06847
 |
Component:243; Jaccard:0.078489
 |
Component:270; Jaccard:0.08442
 |
Component:270; Jaccard:0.07898
 |
Component:104; Jaccard:0.067732
 |
Component:243; Jaccard:0.052777
 |
Component:167; Jaccard:0.03346
 |
Correlation:0.063146
  |
Correlation:0.11655
  |
Correlation:0.016259
  |
Correlation:0.016259
  |
Correlation:-0.059497
  |
Correlation:0.11655
  |
Correlation:0.05842
  |
Component:370; Jaccard:0.067517
 |
Component:103; Jaccard:0.07396
 |
Component:243; Jaccard:0.080193
 |
Component:243; Jaccard:0.077526
 |
Component:243; Jaccard:0.067293
 |
Component:104; Jaccard:0.04972
 |
Component:243; Jaccard:0.032295
 |
Correlation:0.055698
  |
Correlation:-0.069028
  |
Correlation:0.11655
  |
Correlation:0.11655
  |
Correlation:0.11655
  |
Correlation:-0.059497
  |
Correlation:0.11655
  |
Cope: 06:"SHAPES-FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
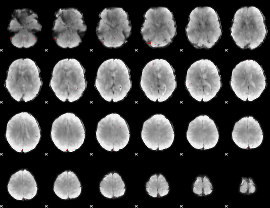 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:010; Jaccard:0.098801
 |
Component:010; Jaccard:0.094191
 |
Component:272; Jaccard:0.10191
 |
Component:272; Jaccard:0.097444
 |
Component:272; Jaccard:0.076263
 |
Component:272; Jaccard:0.057497
 |
Component:272; Jaccard:0.038009
 |
Correlation:0.29788
  |
Correlation:0.29788
  |
Correlation:0.2363
  |
Correlation:0.2363
  |
Correlation:0.2363
  |
Correlation:0.2363
  |
Correlation:0.2363
  |
Component:204; Jaccard:0.076901
 |
Component:272; Jaccard:0.092575
 |
Component:010; Jaccard:0.073467
 |
Component:185; Jaccard:0.069391
 |
Component:185; Jaccard:0.053946
 |
Component:185; Jaccard:0.038185
 |
Component:185; Jaccard:0.02627
 |
Correlation:0.23007
  |
Correlation:0.2363
  |
Correlation:0.29788
  |
Correlation:0.20981
  |
Correlation:0.20981
  |
Correlation:0.20981
  |
Correlation:0.20981
  |
Component:272; Jaccard:0.075519
 |
Component:204; Jaccard:0.071624
 |
Component:185; Jaccard:0.072022
 |
Component:127; Jaccard:0.047315
 |
Component:127; Jaccard:0.036547
 |
Component:127; Jaccard:0.027039
 |
Component:078; Jaccard:0.019362
 |
Correlation:0.2363
  |
Correlation:0.23007
  |
Correlation:0.20981
  |
Correlation:0.22162
  |
Correlation:0.22162
  |
Correlation:0.22162
  |
Correlation:0.15089
  |
Component:172; Jaccard:0.071877
 |
Component:343; Jaccard:0.070496
 |
Component:343; Jaccard:0.063395
 |
Component:343; Jaccard:0.045721
 |
Component:078; Jaccard:0.032437
 |
Component:078; Jaccard:0.022993
 |
Component:127; Jaccard:0.017148
 |
Correlation:0.39591
  |
Correlation:0.28524
  |
Correlation:0.28524
  |
Correlation:0.28524
  |
Correlation:0.15089
  |
Correlation:0.15089
  |
Correlation:0.22162
  |
Component:343; Jaccard:0.070089
 |
Component:185; Jaccard:0.067751
 |
Component:204; Jaccard:0.052311
 |
Component:010; Jaccard:0.042657
 |
Component:343; Jaccard:0.027243
 |
Component:325; Jaccard:0.018502
 |
Component:325; Jaccard:0.012095
 |
Correlation:0.28524
  |
Correlation:0.20981
  |
Correlation:0.23007
  |
Correlation:0.29788
  |
Correlation:0.28524
  |
Correlation:0.047991
  |
Correlation:0.047991
  |
Component:282; Jaccard:0.069264
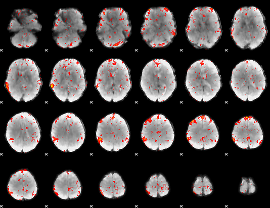 |
Component:172; Jaccard:0.066196
 |
Component:172; Jaccard:0.051012
 |
Component:078; Jaccard:0.040422
 |
Component:325; Jaccard:0.027243
 |
Component:343; Jaccard:0.013767
 |
Component:096; Jaccard:0.009484
 |
Correlation:0.15429
  |
Correlation:0.39591
  |
Correlation:0.39591
  |
Correlation:0.15089
  |
Correlation:0.047991
  |
Correlation:0.28524
  |
Correlation:0.063158
  |
Component:050; Jaccard:0.065324
 |
Component:050; Jaccard:0.062069
 |
Component:050; Jaccard:0.050741
 |
Component:050; Jaccard:0.037507
 |
Component:050; Jaccard:0.023442
 |
Component:262; Jaccard:0.013245
 |
Component:036; Jaccard:0.009241
 |
Correlation:0.35978
  |
Correlation:0.35978
  |
Correlation:0.35978
  |
Correlation:0.35978
  |
Correlation:0.35978
  |
Correlation:0.15343
  |
Correlation:0.042113
  |
Component:330; Jaccard:0.064156
 |
Component:282; Jaccard:0.059421
 |
Component:127; Jaccard:0.050363
 |
Component:262; Jaccard:0.034235
 |
Component:262; Jaccard:0.021154
 |
Component:096; Jaccard:0.012472
 |
Component:343; Jaccard:0.008389
 |
Correlation:0.12498
  |
Correlation:0.15429
  |
Correlation:0.22162
  |
Correlation:0.15343
  |
Correlation:0.15343
  |
Correlation:0.063158
  |
Correlation:0.28524
  |
Component:251; Jaccard:0.062065
 |
Component:330; Jaccard:0.055278
 |
Component:099; Jaccard:0.047795
 |
Component:325; Jaccard:0.031777
 |
Component:010; Jaccard:0.02056
 |
Component:036; Jaccard:0.01229
 |
Component:262; Jaccard:0.008262
 |
Correlation:0.15105
  |
Correlation:0.12498
  |
Correlation:0.15374
  |
Correlation:0.047991
  |
Correlation:0.29788
  |
Correlation:0.042113
  |
Correlation:0.15343
  |
Component:262; Jaccard:0.059298
 |
Component:099; Jaccard:0.055146
 |
Component:262; Jaccard:0.043941
 |
Component:099; Jaccard:0.030167
 |
Component:115; Jaccard:0.018778
 |
Component:050; Jaccard:0.011809
 |
Component:107; Jaccard:0.007737
 |
Correlation:0.15343
  |
Correlation:0.15374
  |
Correlation:0.15343
  |
Correlation:0.15374
  |
Correlation:0.14288
  |
Correlation:0.35978
  |
Correlation:0.08033
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































