Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
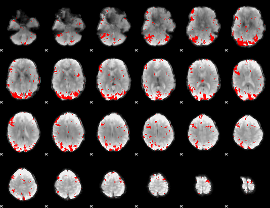 |
GLM binary result thresholded by 3
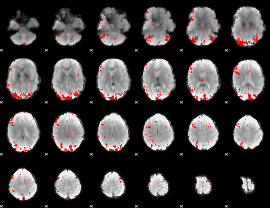 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
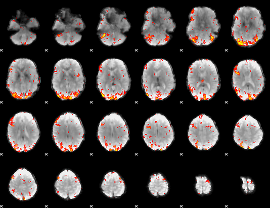 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:203; Jaccard:0.14594
 |
Component:203; Jaccard:0.19053
 |
Component:203; Jaccard:0.23819
 |
Component:203; Jaccard:0.27025
 |
Component:203; Jaccard:0.27651
 |
Component:203; Jaccard:0.25374
 |
Component:203; Jaccard:0.21583
 |
Correlation:0.25256
  |
Correlation:0.25256
  |
Correlation:0.25256
  |
Correlation:0.25256
  |
Correlation:0.25256
  |
Correlation:0.25256
  |
Correlation:0.25256
  |
Component:308; Jaccard:0.12075
 |
Component:308; Jaccard:0.14483
 |
Component:381; Jaccard:0.16926
 |
Component:381; Jaccard:0.18793
 |
Component:381; Jaccard:0.18981
 |
Component:381; Jaccard:0.1806
 |
Component:381; Jaccard:0.1571
 |
Correlation:0.5324
  |
Correlation:0.5324
  |
Correlation:0.55287
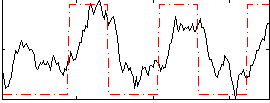  |
Correlation:0.55287
  |
Correlation:0.55287
  |
Correlation:0.55287
  |
Correlation:0.55287
  |
Component:381; Jaccard:0.11376
 |
Component:381; Jaccard:0.14185
 |
Component:308; Jaccard:0.16091
 |
Component:308; Jaccard:0.16686
 |
Component:308; Jaccard:0.15504
 |
Component:308; Jaccard:0.1371
 |
Component:308; Jaccard:0.11971
 |
Correlation:0.55287
  |
Correlation:0.55287
  |
Correlation:0.5324
  |
Correlation:0.5324
  |
Correlation:0.5324
  |
Correlation:0.5324
  |
Correlation:0.5324
  |
Component:137; Jaccard:0.11321
 |
Component:276; Jaccard:0.13176
 |
Component:276; Jaccard:0.14376
 |
Component:338; Jaccard:0.14958
 |
Component:338; Jaccard:0.14893
 |
Component:338; Jaccard:0.1367
 |
Component:338; Jaccard:0.11885
 |
Correlation:0.002245
  |
Correlation:0.13342
  |
Correlation:0.13342
  |
Correlation:0.12917
  |
Correlation:0.12917
  |
Correlation:0.12917
  |
Correlation:0.12917
  |
Component:276; Jaccard:0.1114
 |
Component:137; Jaccard:0.12313
 |
Component:338; Jaccard:0.13735
 |
Component:276; Jaccard:0.14426
 |
Component:276; Jaccard:0.12812
 |
Component:276; Jaccard:0.10213
 |
Component:276; Jaccard:0.078589
 |
Correlation:0.13342
  |
Correlation:0.002245
  |
Correlation:0.12917
  |
Correlation:0.13342
  |
Correlation:0.13342
  |
Correlation:0.13342
  |
Correlation:0.13342
  |
Component:338; Jaccard:0.089669
 |
Component:338; Jaccard:0.11294
 |
Component:137; Jaccard:0.12553
 |
Component:137; Jaccard:0.11714
 |
Component:137; Jaccard:0.10373
 |
Component:169; Jaccard:0.090425
 |
Component:169; Jaccard:0.075998
 |
Correlation:0.12917
  |
Correlation:0.12917
  |
Correlation:0.002245
  |
Correlation:0.002245
  |
Correlation:0.002245
  |
Correlation:-0.33666
  |
Correlation:-0.33666
  |
Component:293; Jaccard:0.088769
 |
Component:293; Jaccard:0.096646
 |
Component:169; Jaccard:0.10369
 |
Component:169; Jaccard:0.1093
 |
Component:169; Jaccard:0.10368
 |
Component:137; Jaccard:0.086978
 |
Component:028; Jaccard:0.075309
 |
Correlation:0.12326
  |
Correlation:0.12326
  |
Correlation:-0.33666
  |
Correlation:-0.33666
  |
Correlation:-0.33666
  |
Correlation:0.002245
  |
Correlation:0.010384
  |
Component:390; Jaccard:0.083533
 |
Component:169; Jaccard:0.094465
 |
Component:293; Jaccard:0.098906
 |
Component:293; Jaccard:0.092852
 |
Component:028; Jaccard:0.089536
 |
Component:028; Jaccard:0.084815
 |
Component:137; Jaccard:0.07009
 |
Correlation:0.13438
  |
Correlation:-0.33666
  |
Correlation:0.12326
  |
Correlation:0.12326
  |
Correlation:0.010384
  |
Correlation:0.010384
  |
Correlation:0.002245
  |
Component:106; Jaccard:0.081696
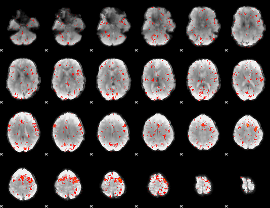 |
Component:390; Jaccard:0.089097
 |
Component:240; Jaccard:0.08987
 |
Component:028; Jaccard:0.089763
 |
Component:080; Jaccard:0.081781
 |
Component:080; Jaccard:0.073651
 |
Component:236; Jaccard:0.06317
 |
Correlation:0.067683
  |
Correlation:0.13438
  |
Correlation:0.078845
  |
Correlation:0.010384
  |
Correlation:0.054541
  |
Correlation:0.054541
  |
Correlation:0.17468
  |
Component:169; Jaccard:0.081646
 |
Component:240; Jaccard:0.087599
 |
Component:080; Jaccard:0.08965
 |
Component:080; Jaccard:0.088552
 |
Component:293; Jaccard:0.078819
 |
Component:236; Jaccard:0.072158
 |
Component:080; Jaccard:0.061631
 |
Correlation:-0.33666
  |
Correlation:0.078845
  |
Correlation:0.054541
  |
Correlation:0.054541
  |
Correlation:0.12326
  |
Correlation:0.17468
  |
Correlation:0.054541
  |
Cope: 02:"SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
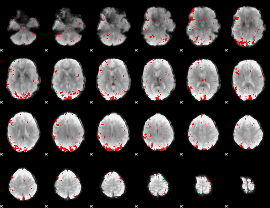 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
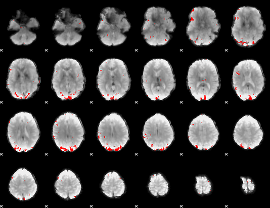 |
GLM binary result thresholded by 4
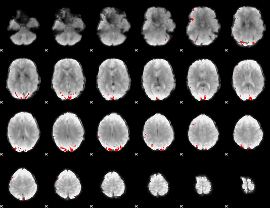 |
GLM Z-score result thresholded by 1
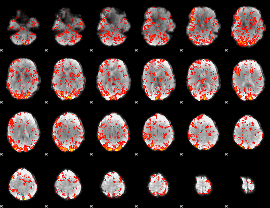 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
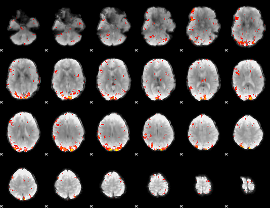 |
GLM Z-score result thresholded by 3
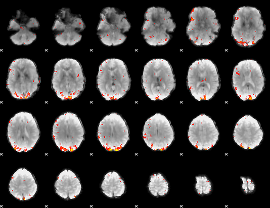 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:203; Jaccard:0.13325
 |
Component:169; Jaccard:0.16836
 |
Component:169; Jaccard:0.20043
 |
Component:169; Jaccard:0.21529
 |
Component:169; Jaccard:0.20539
 |
Component:169; Jaccard:0.17492
 |
Component:169; Jaccard:0.14053
 |
Correlation:-0.14818
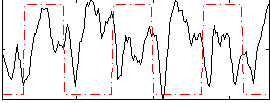  |
Correlation:0.43044
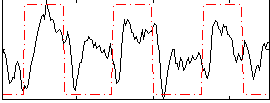 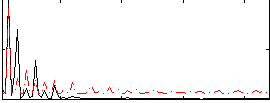 |
Correlation:0.43044
  |
Correlation:0.43044
  |
Correlation:0.43044
  |
Correlation:0.43044
  |
Correlation:0.43044
  |
Component:169; Jaccard:0.13171
 |
Component:203; Jaccard:0.16429
 |
Component:203; Jaccard:0.18583
 |
Component:203; Jaccard:0.18846
 |
Component:203; Jaccard:0.1624
 |
Component:203; Jaccard:0.12711
 |
Component:203; Jaccard:0.096177
 |
Correlation:0.43044
  |
Correlation:-0.14818
  |
Correlation:-0.14818
  |
Correlation:-0.14818
  |
Correlation:-0.14818
  |
Correlation:-0.14818
  |
Correlation:-0.14818
  |
Component:137; Jaccard:0.10348
 |
Component:137; Jaccard:0.10603
 |
Component:369; Jaccard:0.11198
 |
Component:369; Jaccard:0.10638
 |
Component:369; Jaccard:0.092933
 |
Component:369; Jaccard:0.076103
 |
Component:236; Jaccard:0.063025
 |
Correlation:0.05031
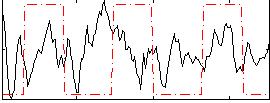  |
Correlation:0.05031
  |
Correlation:0.20267
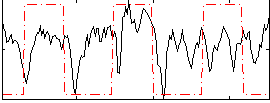  |
Correlation:0.20267
  |
Correlation:0.20267
  |
Correlation:0.20267
  |
Correlation:-0.053568
  |
Component:369; Jaccard:0.091532
 |
Component:369; Jaccard:0.10418
 |
Component:137; Jaccard:0.10216
 |
Component:236; Jaccard:0.091541
 |
Component:236; Jaccard:0.08384
 |
Component:236; Jaccard:0.074488
 |
Component:369; Jaccard:0.059748
 |
Correlation:0.20267
  |
Correlation:0.20267
  |
Correlation:0.05031
  |
Correlation:-0.053568
  |
Correlation:-0.053568
  |
Correlation:-0.053568
  |
Correlation:0.20267
  |
Component:291; Jaccard:0.090105
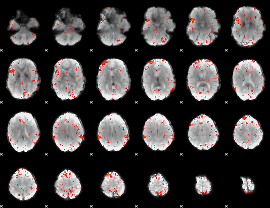 |
Component:276; Jaccard:0.095573
 |
Component:080; Jaccard:0.095494
 |
Component:080; Jaccard:0.086479
 |
Component:080; Jaccard:0.07468
 |
Component:080; Jaccard:0.056765
 |
Component:028; Jaccard:0.042536
 |
Correlation:0.21646
  |
Correlation:-0.0024445
  |
Correlation:-0.020935
  |
Correlation:-0.020935
  |
Correlation:-0.020935
  |
Correlation:-0.020935
  |
Correlation:0.064862
  |
Component:276; Jaccard:0.089098
 |
Component:291; Jaccard:0.094673
 |
Component:236; Jaccard:0.092467
 |
Component:137; Jaccard:0.086266
 |
Component:028; Jaccard:0.070968
 |
Component:028; Jaccard:0.055861
 |
Component:080; Jaccard:0.04207
 |
Correlation:-0.0024445
  |
Correlation:0.21646
  |
Correlation:-0.053568
  |
Correlation:0.05031
  |
Correlation:0.064862
  |
Correlation:0.064862
  |
Correlation:-0.020935
  |
Component:080; Jaccard:0.086437
 |
Component:080; Jaccard:0.094236
 |
Component:276; Jaccard:0.09171
 |
Component:276; Jaccard:0.081216
 |
Component:137; Jaccard:0.06893
 |
Component:137; Jaccard:0.05217
 |
Component:137; Jaccard:0.038871
 |
Correlation:-0.020935
  |
Correlation:-0.020935
  |
Correlation:-0.0024445
  |
Correlation:-0.0024445
  |
Correlation:0.05031
  |
Correlation:0.05031
  |
Correlation:0.05031
  |
Component:240; Jaccard:0.078464
 |
Component:240; Jaccard:0.086083
 |
Component:291; Jaccard:0.09061
 |
Component:028; Jaccard:0.080566
 |
Component:338; Jaccard:0.062643
 |
Component:338; Jaccard:0.048213
 |
Component:074; Jaccard:0.033023
 |
Correlation:-0.082427
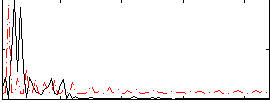  |
Correlation:-0.082427
  |
Correlation:0.21646
  |
Correlation:0.064862
  |
Correlation:-0.015896
  |
Correlation:-0.015896
  |
Correlation:0.34663
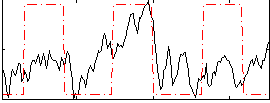  |
Component:293; Jaccard:0.075581
 |
Component:236; Jaccard:0.083844
 |
Component:240; Jaccard:0.087648
 |
Component:240; Jaccard:0.079269
 |
Component:276; Jaccard:0.060951
 |
Component:240; Jaccard:0.041878
 |
Component:348; Jaccard:0.031227
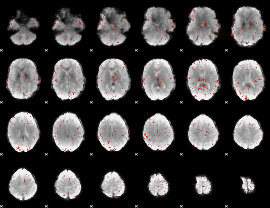 |
Correlation:-0.15063
  |
Correlation:-0.053568
  |
Correlation:-0.082427
  |
Correlation:-0.082427
  |
Correlation:-0.0024445
  |
Correlation:-0.082427
  |
Correlation:0.21117
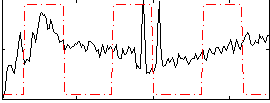  |
Component:390; Jaccard:0.073317
 |
Component:042; Jaccard:0.08126
 |
Component:338; Jaccard:0.085442
 |
Component:338; Jaccard:0.079076
 |
Component:240; Jaccard:0.059599
 |
Component:074; Jaccard:0.041469
 |
Component:338; Jaccard:0.031196
 |
Correlation:0.13794
  |
Correlation:0.15894
 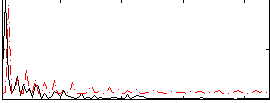 |
Correlation:-0.015896
  |
Correlation:-0.015896
  |
Correlation:-0.082427
  |
Correlation:0.34663
  |
Correlation:-0.015896
  |
Cope: 03:"FACES-SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
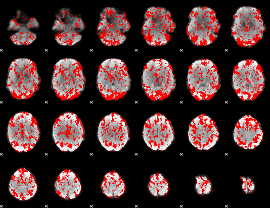 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
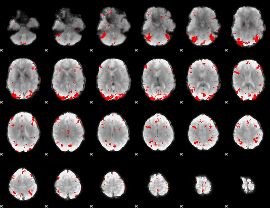 |
GLM binary result thresholded by 3.5
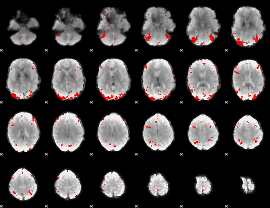 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
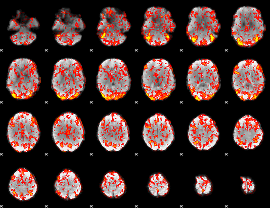 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
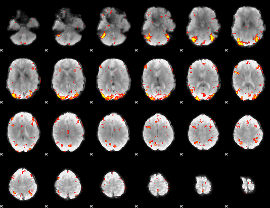 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:308; Jaccard:0.12873
 |
Component:381; Jaccard:0.18096
 |
Component:381; Jaccard:0.25428
 |
Component:381; Jaccard:0.32187
 |
Component:381; Jaccard:0.35941
 |
Component:381; Jaccard:0.36845
 |
Component:381; Jaccard:0.33894
 |
Correlation:0.55948
  |
Correlation:0.54973
  |
Correlation:0.54973
  |
Correlation:0.54973
  |
Correlation:0.54973
  |
Correlation:0.54973
  |
Correlation:0.54973
  |
Component:381; Jaccard:0.12471
 |
Component:308; Jaccard:0.17316
 |
Component:308; Jaccard:0.21646
 |
Component:308; Jaccard:0.23633
 |
Component:308; Jaccard:0.23086
 |
Component:308; Jaccard:0.2081
 |
Component:308; Jaccard:0.18026
 |
Correlation:0.54973
  |
Correlation:0.55948
  |
Correlation:0.55948
  |
Correlation:0.55948
  |
Correlation:0.55948
  |
Correlation:0.55948
  |
Correlation:0.55948
  |
Component:277; Jaccard:0.097015
 |
Component:277; Jaccard:0.11671
 |
Component:203; Jaccard:0.13467
 |
Component:203; Jaccard:0.14914
 |
Component:203; Jaccard:0.15062
 |
Component:203; Jaccard:0.14551
 |
Component:203; Jaccard:0.127
 |
Correlation:0.14723
  |
Correlation:0.14723
  |
Correlation:0.21736
  |
Correlation:0.21736
  |
Correlation:0.21736
  |
Correlation:0.21736
  |
Correlation:0.21736
  |
Component:203; Jaccard:0.095176
 |
Component:203; Jaccard:0.1157
 |
Component:277; Jaccard:0.13378
 |
Component:277; Jaccard:0.14067
 |
Component:250; Jaccard:0.13451
 |
Component:338; Jaccard:0.13247
 |
Component:338; Jaccard:0.12319
 |
Correlation:0.21736
  |
Correlation:0.21736
  |
Correlation:0.14723
  |
Correlation:0.14723
  |
Correlation:0.26948
  |
Correlation:0.078684
  |
Correlation:0.078684
  |
Component:183; Jaccard:0.091185
 |
Component:183; Jaccard:0.10563
 |
Component:183; Jaccard:0.11644
 |
Component:250; Jaccard:0.13027
 |
Component:277; Jaccard:0.13375
 |
Component:250; Jaccard:0.12325
 |
Component:250; Jaccard:0.10291
 |
Correlation:0.3326
  |
Correlation:0.3326
  |
Correlation:0.3326
  |
Correlation:0.26948
  |
Correlation:0.14723
  |
Correlation:0.26948
  |
Correlation:0.26948
  |
Component:276; Jaccard:0.084643
 |
Component:276; Jaccard:0.098698
 |
Component:250; Jaccard:0.1149
 |
Component:338; Jaccard:0.12183
 |
Component:338; Jaccard:0.12992
 |
Component:277; Jaccard:0.11791
 |
Component:277; Jaccard:0.10037
 |
Correlation:0.073692
  |
Correlation:0.073692
  |
Correlation:0.26948
  |
Correlation:0.078684
  |
Correlation:0.078684
  |
Correlation:0.14723
  |
Correlation:0.14723
  |
Component:334; Jaccard:0.08316
 |
Component:250; Jaccard:0.093377
 |
Component:276; Jaccard:0.10887
 |
Component:183; Jaccard:0.11849
 |
Component:183; Jaccard:0.10982
 |
Component:205; Jaccard:0.098621
 |
Component:205; Jaccard:0.08687
 |
Correlation:0.089988
  |
Correlation:0.26948
  |
Correlation:0.073692
  |
Correlation:0.3326
  |
Correlation:0.3326
  |
Correlation:0.39925
  |
Correlation:0.39925
  |
Component:386; Jaccard:0.080906
 |
Component:205; Jaccard:0.092427
 |
Component:205; Jaccard:0.10624
 |
Component:205; Jaccard:0.11072
 |
Component:205; Jaccard:0.1063
 |
Component:183; Jaccard:0.091887
 |
Component:183; Jaccard:0.075674
 |
Correlation:0.42839
  |
Correlation:0.39925
  |
Correlation:0.39925
  |
Correlation:0.39925
  |
Correlation:0.39925
  |
Correlation:0.3326
  |
Correlation:0.3326
  |
Component:215; Jaccard:0.078799
 |
Component:386; Jaccard:0.08964
 |
Component:338; Jaccard:0.10402
 |
Component:276; Jaccard:0.10897
 |
Component:276; Jaccard:0.097407
 |
Component:276; Jaccard:0.083181
 |
Component:190; Jaccard:0.071228
 |
Correlation:0.19514
  |
Correlation:0.42839
  |
Correlation:0.078684
  |
Correlation:0.073692
  |
Correlation:0.073692
  |
Correlation:0.073692
  |
Correlation:0.33799
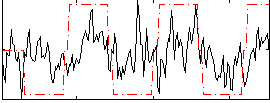  |
Component:137; Jaccard:0.077525
 |
Component:215; Jaccard:0.089006
 |
Component:215; Jaccard:0.096991
 |
Component:215; Jaccard:0.094623
 |
Component:215; Jaccard:0.084694
 |
Component:190; Jaccard:0.078636
 |
Component:276; Jaccard:0.065392
 |
Correlation:-0.026071
 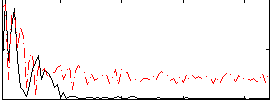 |
Correlation:0.19514
  |
Correlation:0.19514
  |
Correlation:0.19514
  |
Correlation:0.19514
  |
Correlation:0.33799
  |
Correlation:0.073692
  |
Cope: 04:"neg_FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
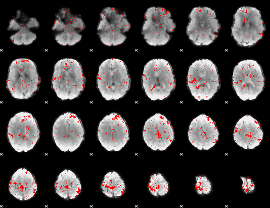 |
GLM binary result thresholded by 3
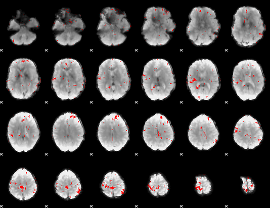 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
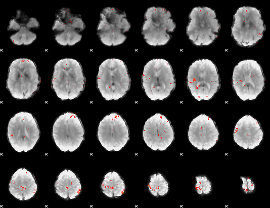 |
GLM Z-score result thresholded by 4
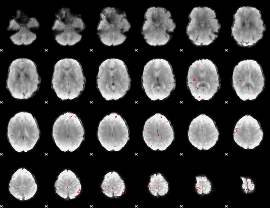 |
Component:178; Jaccard:0.10538
 |
Component:178; Jaccard:0.12153
 |
Component:178; Jaccard:0.12769
 |
Component:178; Jaccard:0.11835
 |
Component:178; Jaccard:0.08954
 |
Component:178; Jaccard:0.051236
 |
Component:178; Jaccard:0.025786
 |
Correlation:0.048103
  |
Correlation:0.048103
  |
Correlation:0.048103
  |
Correlation:0.048103
  |
Correlation:0.048103
  |
Correlation:0.048103
  |
Correlation:0.048103
  |
Component:053; Jaccard:0.098033
 |
Component:053; Jaccard:0.11545
 |
Component:053; Jaccard:0.1248
 |
Component:053; Jaccard:0.11167
 |
Component:053; Jaccard:0.082161
 |
Component:053; Jaccard:0.043157
 |
Component:133; Jaccard:0.021646
 |
Correlation:0.13224
  |
Correlation:0.13224
  |
Correlation:0.13224
  |
Correlation:0.13224
  |
Correlation:0.13224
  |
Correlation:0.13224
  |
Correlation:0.074358
  |
Component:165; Jaccard:0.090497
 |
Component:165; Jaccard:0.10137
 |
Component:350; Jaccard:0.10688
 |
Component:133; Jaccard:0.090068
 |
Component:133; Jaccard:0.06714
 |
Component:133; Jaccard:0.041769
 |
Component:033; Jaccard:0.020627
 |
Correlation:0.14154
  |
Correlation:0.14154
  |
Correlation:0.044439
  |
Correlation:0.074358
  |
Correlation:0.074358
  |
Correlation:0.074358
  |
Correlation:0.0071142
  |
Component:350; Jaccard:0.081828
 |
Component:350; Jaccard:0.098897
 |
Component:165; Jaccard:0.10387
 |
Component:350; Jaccard:0.089505
 |
Component:020; Jaccard:0.065005
 |
Component:020; Jaccard:0.036617
 |
Component:020; Jaccard:0.017612
 |
Correlation:0.044439
  |
Correlation:0.044439
  |
Correlation:0.14154
  |
Correlation:0.044439
  |
Correlation:-0.090381
  |
Correlation:-0.090381
  |
Correlation:-0.090381
  |
Component:020; Jaccard:0.081351
 |
Component:020; Jaccard:0.093716
 |
Component:020; Jaccard:0.099169
 |
Component:020; Jaccard:0.089438
 |
Component:191; Jaccard:0.058098
 |
Component:191; Jaccard:0.034491
 |
Component:191; Jaccard:0.017425
 |
Correlation:-0.090381
  |
Correlation:-0.090381
  |
Correlation:-0.090381
  |
Correlation:-0.090381
  |
Correlation:0.01996
  |
Correlation:0.01996
  |
Correlation:0.01996
  |
Component:380; Jaccard:0.077015
 |
Component:133; Jaccard:0.086117
 |
Component:133; Jaccard:0.093999
 |
Component:165; Jaccard:0.087767
 |
Component:165; Jaccard:0.057098
 |
Component:033; Jaccard:0.031736
 |
Component:053; Jaccard:0.017138
 |
Correlation:-0.094461
  |
Correlation:0.074358
  |
Correlation:0.074358
  |
Correlation:0.14154
  |
Correlation:0.14154
  |
Correlation:0.0071142
  |
Correlation:0.13224
  |
Component:133; Jaccard:0.076916
 |
Component:191; Jaccard:0.08209
 |
Component:191; Jaccard:0.084956
 |
Component:191; Jaccard:0.079768
 |
Component:350; Jaccard:0.056001
 |
Component:165; Jaccard:0.029715
 |
Component:165; Jaccard:0.014074
 |
Correlation:0.074358
  |
Correlation:0.01996
  |
Correlation:0.01996
  |
Correlation:0.01996
  |
Correlation:0.044439
  |
Correlation:0.14154
  |
Correlation:0.14154
  |
Component:383; Jaccard:0.075839
 |
Component:372; Jaccard:0.07872
 |
Component:372; Jaccard:0.078881
 |
Component:136; Jaccard:0.069914
 |
Component:136; Jaccard:0.049926
 |
Component:280; Jaccard:0.027775
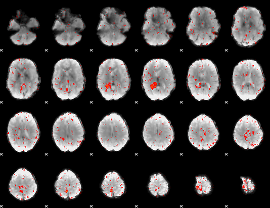 |
Component:136; Jaccard:0.012546
 |
Correlation:-0.015439
  |
Correlation:0.15193
  |
Correlation:0.15193
  |
Correlation:0.10176
 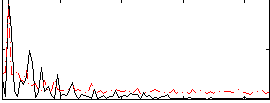 |
Correlation:0.10176
  |
Correlation:0.027955
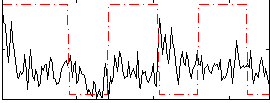  |
Correlation:0.10176
  |
Component:191; Jaccard:0.073628
 |
Component:280; Jaccard:0.078203
 |
Component:280; Jaccard:0.078607
 |
Component:280; Jaccard:0.069508
 |
Component:383; Jaccard:0.049902
 |
Component:136; Jaccard:0.027321
 |
Component:383; Jaccard:0.011948
 |
Correlation:0.01996
  |
Correlation:0.027955
  |
Correlation:0.027955
  |
Correlation:0.027955
  |
Correlation:-0.015439
  |
Correlation:0.10176
  |
Correlation:-0.015439
  |
Component:372; Jaccard:0.073274
 |
Component:383; Jaccard:0.07812
 |
Component:136; Jaccard:0.078216
 |
Component:383; Jaccard:0.065933
 |
Component:280; Jaccard:0.048872
 |
Component:383; Jaccard:0.027292
 |
Component:292; Jaccard:0.01186
 |
Correlation:0.15193
  |
Correlation:-0.015439
  |
Correlation:0.10176
  |
Correlation:-0.015439
  |
Correlation:0.027955
  |
Correlation:-0.015439
  |
Correlation:-0.19815
  |
Cope: 05:"neg_SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
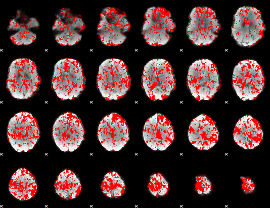 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:165; Jaccard:0.095614
 |
Component:165; Jaccard:0.11128
 |
Component:165; Jaccard:0.12149
 |
Component:165; Jaccard:0.11499
 |
Component:183; Jaccard:0.091511
 |
Component:183; Jaccard:0.065636
 |
Component:133; Jaccard:0.038606
 |
Correlation:0.11337
  |
Correlation:0.11337
  |
Correlation:0.11337
  |
Correlation:0.11337
  |
Correlation:0.29246
  |
Correlation:0.29246
  |
Correlation:0.23247
  |
Component:178; Jaccard:0.092175
 |
Component:178; Jaccard:0.10266
 |
Component:020; Jaccard:0.11071
 |
Component:020; Jaccard:0.11263
 |
Component:020; Jaccard:0.091038
 |
Component:133; Jaccard:0.06557
 |
Component:183; Jaccard:0.034842
 |
Correlation:-0.0070125
  |
Correlation:-0.0070125
  |
Correlation:0.21366
  |
Correlation:0.21366
  |
Correlation:0.21366
  |
Correlation:0.23247
  |
Correlation:0.29246
  |
Component:380; Jaccard:0.085796
 |
Component:020; Jaccard:0.099306
 |
Component:133; Jaccard:0.10562
 |
Component:133; Jaccard:0.10768
 |
Component:133; Jaccard:0.090493
 |
Component:020; Jaccard:0.058464
 |
Component:165; Jaccard:0.030187
 |
Correlation:0.21072
  |
Correlation:0.21366
  |
Correlation:0.23247
  |
Correlation:0.23247
  |
Correlation:0.23247
  |
Correlation:0.21366
  |
Correlation:0.11337
  |
Component:051; Jaccard:0.084552
 |
Component:380; Jaccard:0.094313
 |
Component:178; Jaccard:0.10504
 |
Component:183; Jaccard:0.10476
 |
Component:165; Jaccard:0.08914
 |
Component:165; Jaccard:0.056625
 |
Component:191; Jaccard:0.029766
 |
Correlation:0.18166
  |
Correlation:0.21072
  |
Correlation:-0.0070125
  |
Correlation:0.29246
  |
Correlation:0.11337
  |
Correlation:0.11337
  |
Correlation:0.027731
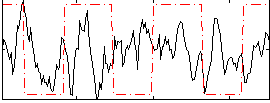 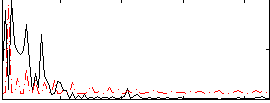 |
Component:020; Jaccard:0.082779
 |
Component:183; Jaccard:0.093022
 |
Component:183; Jaccard:0.10224
 |
Component:178; Jaccard:0.093133
 |
Component:191; Jaccard:0.076417
 |
Component:191; Jaccard:0.052283
 |
Component:020; Jaccard:0.028907
 |
Correlation:0.21366
  |
Correlation:0.29246
  |
Correlation:0.29246
  |
Correlation:-0.0070125
  |
Correlation:0.027731
  |
Correlation:0.027731
  |
Correlation:0.21366
  |
Component:183; Jaccard:0.080768
 |
Component:133; Jaccard:0.092708
 |
Component:380; Jaccard:0.098536
 |
Component:191; Jaccard:0.092173
 |
Component:380; Jaccard:0.072044
 |
Component:215; Jaccard:0.046701
 |
Component:378; Jaccard:0.026578
 |
Correlation:0.29246
  |
Correlation:0.23247
  |
Correlation:0.21072
  |
Correlation:0.027731
  |
Correlation:0.21072
  |
Correlation:0.2405
  |
Correlation:-0.052377
  |
Component:133; Jaccard:0.079122
 |
Component:191; Jaccard:0.085497
 |
Component:191; Jaccard:0.09287
 |
Component:380; Jaccard:0.091976
 |
Component:178; Jaccard:0.07158
 |
Component:178; Jaccard:0.0463
 |
Component:215; Jaccard:0.025753
 |
Correlation:0.23247
  |
Correlation:0.027731
  |
Correlation:0.027731
  |
Correlation:0.21072
  |
Correlation:-0.0070125
  |
Correlation:-0.0070125
  |
Correlation:0.2405
  |
Component:053; Jaccard:0.076928
 |
Component:051; Jaccard:0.084979
 |
Component:053; Jaccard:0.084073
 |
Component:215; Jaccard:0.07935
 |
Component:215; Jaccard:0.065657
 |
Component:380; Jaccard:0.045066
 |
Component:178; Jaccard:0.02461
 |
Correlation:0.063399
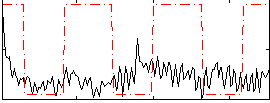 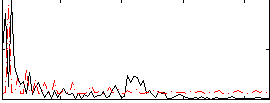 |
Correlation:0.18166
  |
Correlation:0.063399
  |
Correlation:0.2405
  |
Correlation:0.2405
  |
Correlation:0.21072
  |
Correlation:-0.0070125
  |
Component:191; Jaccard:0.074122
 |
Component:053; Jaccard:0.084469
 |
Component:215; Jaccard:0.081173
 |
Component:053; Jaccard:0.072412
 |
Component:205; Jaccard:0.059725
 |
Component:378; Jaccard:0.042743
 |
Component:205; Jaccard:0.024403
 |
Correlation:0.027731
  |
Correlation:0.063399
  |
Correlation:0.2405
  |
Correlation:0.063399
  |
Correlation:0.32391
  |
Correlation:-0.052377
  |
Correlation:0.32391
  |
Component:292; Jaccard:0.073583
 |
Component:292; Jaccard:0.077009
 |
Component:051; Jaccard:0.080005
 |
Component:205; Jaccard:0.072346
 |
Component:378; Jaccard:0.059303
 |
Component:205; Jaccard:0.041662
 |
Component:033; Jaccard:0.023256
 |
Correlation:0.11488
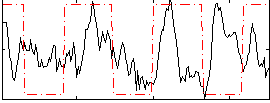  |
Correlation:0.11488
  |
Correlation:0.18166
  |
Correlation:0.32391
  |
Correlation:-0.052377
  |
Correlation:0.32391
  |
Correlation:0.001792
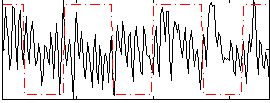  |
Cope: 06:"SHAPES-FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
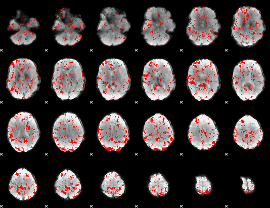 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
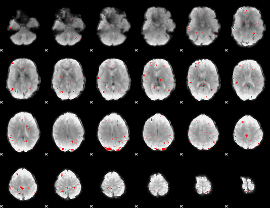 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:021; Jaccard:0.095714
 |
Component:227; Jaccard:0.10327
 |
Component:227; Jaccard:0.098588
 |
Component:169; Jaccard:0.083533
 |
Component:169; Jaccard:0.066193
 |
Component:169; Jaccard:0.047142
 |
Component:169; Jaccard:0.028031
 |
Correlation:0.15992
  |
Correlation:0.16402
  |
Correlation:0.16402
  |
Correlation:0.41609
  |
Correlation:0.41609
  |
Correlation:0.41609
  |
Correlation:0.41609
  |
Component:227; Jaccard:0.094042
 |
Component:021; Jaccard:0.10288
 |
Component:021; Jaccard:0.094462
 |
Component:227; Jaccard:0.07177
 |
Component:074; Jaccard:0.04557
 |
Component:074; Jaccard:0.027531
 |
Component:074; Jaccard:0.012549
 |
Correlation:0.16402
  |
Correlation:0.15992
  |
Correlation:0.15992
  |
Correlation:0.16402
  |
Correlation:0.3186
  |
Correlation:0.3186
  |
Correlation:0.3186
  |
Component:136; Jaccard:0.092307
 |
Component:239; Jaccard:0.097568
 |
Component:169; Jaccard:0.090427
 |
Component:021; Jaccard:0.069063
 |
Component:227; Jaccard:0.044722
 |
Component:021; Jaccard:0.022158
 |
Component:021; Jaccard:0.010251
 |
Correlation:0.031225
  |
Correlation:0.095218
  |
Correlation:0.41609
  |
Correlation:0.15992
  |
Correlation:0.16402
  |
Correlation:0.15992
  |
Correlation:0.15992
  |
Component:239; Jaccard:0.089352
 |
Component:136; Jaccard:0.097483
 |
Component:239; Jaccard:0.089985
 |
Component:074; Jaccard:0.066438
 |
Component:021; Jaccard:0.042154
 |
Component:227; Jaccard:0.021704
 |
Component:107; Jaccard:0.009845
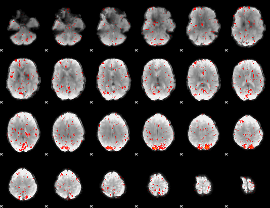 |
Correlation:0.095218
  |
Correlation:0.031225
  |
Correlation:0.095218
  |
Correlation:0.3186
  |
Correlation:0.15992
  |
Correlation:0.16402
  |
Correlation:0.14936
  |
Component:383; Jaccard:0.084513
 |
Component:169; Jaccard:0.087126
 |
Component:136; Jaccard:0.088046
 |
Component:239; Jaccard:0.064283
 |
Component:136; Jaccard:0.037033
 |
Component:107; Jaccard:0.019247
 |
Component:227; Jaccard:0.009675
 |
Correlation:0.074415
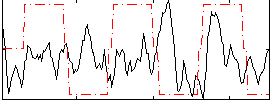 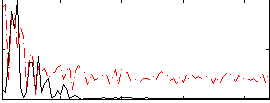 |
Correlation:0.41609
  |
Correlation:0.031225
  |
Correlation:0.095218
  |
Correlation:0.031225
  |
Correlation:0.14936
  |
Correlation:0.16402
  |
Component:169; Jaccard:0.076493
 |
Component:383; Jaccard:0.083311
 |
Component:074; Jaccard:0.075743
 |
Component:136; Jaccard:0.062032
 |
Component:239; Jaccard:0.036867
 |
Component:354; Jaccard:0.017897
 |
Component:369; Jaccard:0.007769
 |
Correlation:0.41609
  |
Correlation:0.074415
  |
Correlation:0.3186
  |
Correlation:0.031225
  |
Correlation:0.095218
  |
Correlation:0.13898
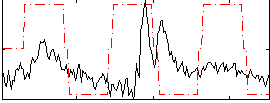  |
Correlation:0.26152
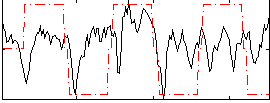  |
Component:107; Jaccard:0.074646
 |
Component:074; Jaccard:0.076615
 |
Component:383; Jaccard:0.071534
 |
Component:171; Jaccard:0.053454
 |
Component:354; Jaccard:0.035848
 |
Component:136; Jaccard:0.016564
 |
Component:136; Jaccard:0.007045
 |
Correlation:0.14936
  |
Correlation:0.3186
  |
Correlation:0.074415
  |
Correlation:0.3217
  |
Correlation:0.13898
  |
Correlation:0.031225
  |
Correlation:0.031225
  |
Component:368; Jaccard:0.074339
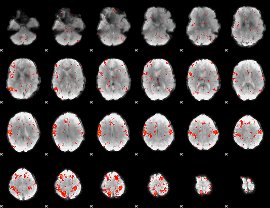 |
Component:171; Jaccard:0.075518
 |
Component:171; Jaccard:0.070336
 |
Component:354; Jaccard:0.053169
 |
Component:107; Jaccard:0.034045
 |
Component:383; Jaccard:0.016296
 |
Component:171; Jaccard:0.007004
 |
Correlation:0.082101
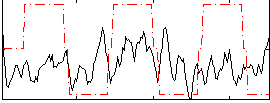 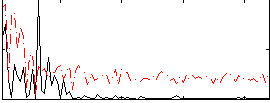 |
Correlation:0.3217
  |
Correlation:0.3217
  |
Correlation:0.13898
  |
Correlation:0.14936
  |
Correlation:0.074415
  |
Correlation:0.3217
  |
Component:053; Jaccard:0.073921
 |
Component:107; Jaccard:0.073353
 |
Component:107; Jaccard:0.065366
 |
Component:383; Jaccard:0.050694
 |
Component:383; Jaccard:0.033018
 |
Component:369; Jaccard:0.015688
 |
Component:383; Jaccard:0.00673
 |
Correlation:0.03734
  |
Correlation:0.14936
  |
Correlation:0.14936
  |
Correlation:0.074415
  |
Correlation:0.074415
  |
Correlation:0.26152
  |
Correlation:0.074415
  |
Component:171; Jaccard:0.07111
 |
Component:368; Jaccard:0.071488
 |
Component:354; Jaccard:0.064915
 |
Component:107; Jaccard:0.050068
 |
Component:171; Jaccard:0.03189
 |
Component:171; Jaccard:0.015309
 |
Component:070; Jaccard:0.005621
 |
Correlation:0.3217
  |
Correlation:0.082101
  |
Correlation:0.13898
  |
Correlation:0.14936
  |
Correlation:0.3217
  |
Correlation:0.3217
  |
Correlation:0.31585
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































