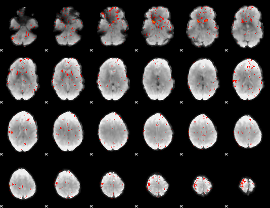
Task name: RELATIONAL, TR=0.72, totally 232 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"MATCH" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
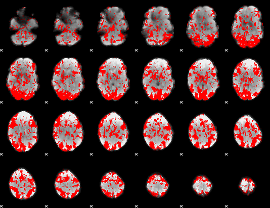 |
GLM binary result thresholded by 1.5
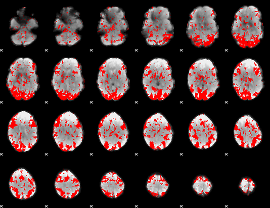 |
GLM binary result thresholded by 2
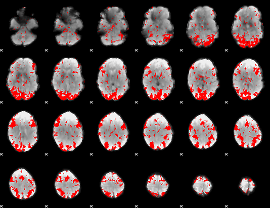 |
GLM binary result thresholded by 2.5
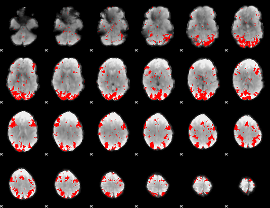 |
GLM binary result thresholded by 3
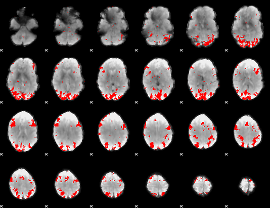 |
GLM binary result thresholded by 3.5
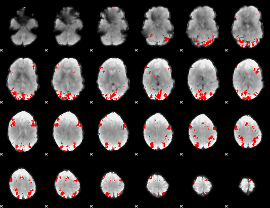 |
GLM binary result thresholded by 4
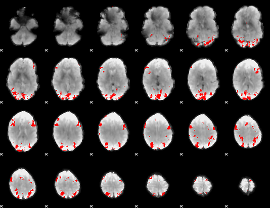 |
GLM Z-score result thresholded by 1
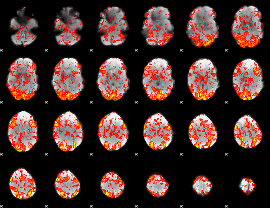 |
GLM Z-score result thresholded by 1.5
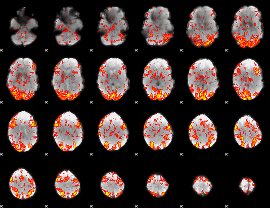 |
GLM Z-score result thresholded by 2
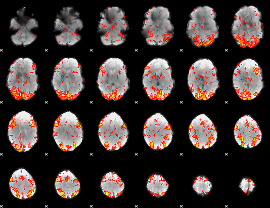 |
GLM Z-score result thresholded by 2.5
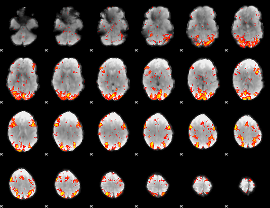 |
GLM Z-score result thresholded by 3
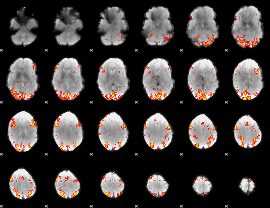 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:268; Jaccard:0.15473
 |
Component:268; Jaccard:0.20545
 |
Component:268; Jaccard:0.26463
 |
Component:268; Jaccard:0.31504
 |
Component:268; Jaccard:0.34248
 |
Component:268; Jaccard:0.34094
 |
Component:268; Jaccard:0.31651
 |
Correlation:0.26572
  |
Correlation:0.26572
  |
Correlation:0.26572
  |
Correlation:0.26572
  |
Correlation:0.26572
  |
Correlation:0.26572
  |
Correlation:0.26572
  |
Component:130; Jaccard:0.13485
 |
Component:130; Jaccard:0.1736
 |
Component:130; Jaccard:0.21456
 |
Component:130; Jaccard:0.24436
 |
Component:130; Jaccard:0.25461
 |
Component:130; Jaccard:0.24797
 |
Component:130; Jaccard:0.23017
 |
Correlation:0.1604
  |
Correlation:0.1604
  |
Correlation:0.1604
  |
Correlation:0.1604
  |
Correlation:0.1604
  |
Correlation:0.1604
  |
Correlation:0.1604
  |
Component:310; Jaccard:0.13207
 |
Component:310; Jaccard:0.16173
 |
Component:161; Jaccard:0.19311
 |
Component:161; Jaccard:0.21726
 |
Component:161; Jaccard:0.22355
 |
Component:161; Jaccard:0.21229
 |
Component:161; Jaccard:0.18899
 |
Correlation:0.15945
  |
Correlation:0.15945
  |
Correlation:0.148
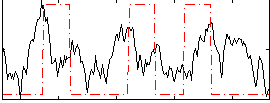 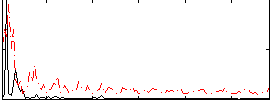 |
Correlation:0.148
  |
Correlation:0.148
  |
Correlation:0.148
  |
Correlation:0.148
  |
Component:161; Jaccard:0.1285
 |
Component:161; Jaccard:0.16028
 |
Component:310; Jaccard:0.18723
 |
Component:310; Jaccard:0.20156
 |
Component:310; Jaccard:0.20001
 |
Component:310; Jaccard:0.18541
 |
Component:310; Jaccard:0.16544
 |
Correlation:0.148
  |
Correlation:0.148
  |
Correlation:0.15945
  |
Correlation:0.15945
  |
Correlation:0.15945
  |
Correlation:0.15945
  |
Correlation:0.15945
  |
Component:181; Jaccard:0.12134
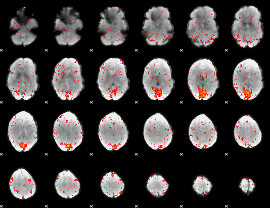 |
Component:399; Jaccard:0.14684
 |
Component:399; Jaccard:0.17098
 |
Component:332; Jaccard:0.18555
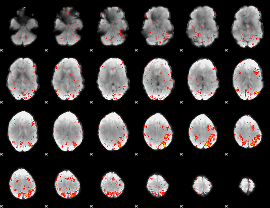 |
Component:332; Jaccard:0.18831
 |
Component:332; Jaccard:0.17648
 |
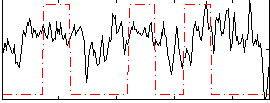
Component:071; Jaccard:0.15551
 |
Correlation:0.21874
 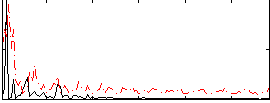 |
Correlation:0.17868
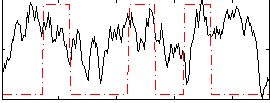 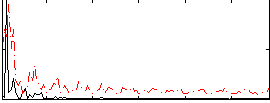 |
Correlation:0.17868
  |
Correlation:0.20799
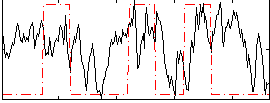 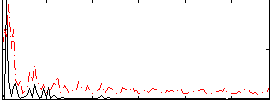 |
Correlation:0.20799
  |
Correlation:0.20799
  |
Correlation:-0.0050556
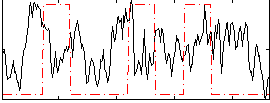 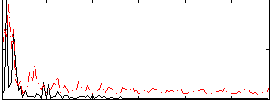 |
Component:231; Jaccard:0.12003
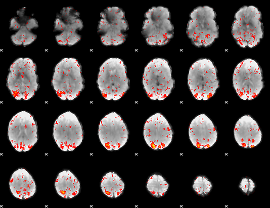 |
Component:071; Jaccard:0.14499
 |
Component:071; Jaccard:0.17068
 |
Component:399; Jaccard:0.18481
 |
Component:071; Jaccard:0.18562
 |
Component:071; Jaccard:0.17497
 |
Component:332; Jaccard:0.15437
 |
Correlation:0.18769
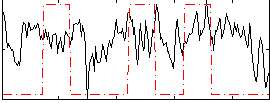 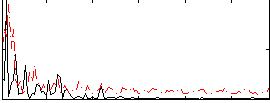 |
Correlation:-0.0050556
  |
Correlation:-0.0050556
  |
Correlation:0.17868
  |
Correlation:-0.0050556
  |
Correlation:-0.0050556
  |
Correlation:0.20799
  |
Component:399; Jaccard:0.11958
 |
Component:181; Jaccard:0.14208
 |
Component:332; Jaccard:0.16676
 |
Component:071; Jaccard:0.18382
 |
Component:399; Jaccard:0.18538
 |
Component:399; Jaccard:0.17202
 |
Component:399; Jaccard:0.1485
 |
Correlation:0.17868
  |
Correlation:0.21874
  |
Correlation:0.20799
  |
Correlation:-0.0050556
  |
Correlation:0.17868
  |
Correlation:0.17868
  |
Correlation:0.17868
  |
Component:071; Jaccard:0.11874
 |
Component:332; Jaccard:0.14068
 |
Component:088; Jaccard:0.158
 |
Component:088; Jaccard:0.17302
 |
Component:088; Jaccard:0.17188
 |
Component:182; Jaccard:0.15824
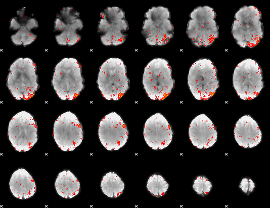 |
Component:182; Jaccard:0.1426
 |
Correlation:-0.0050556
  |
Correlation:0.20799
  |
Correlation:0.1466
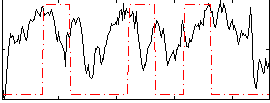 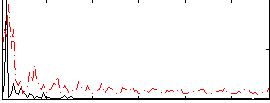 |
Correlation:0.1466
  |
Correlation:0.1466
  |
Correlation:0.20297
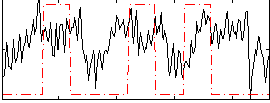 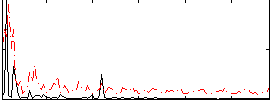 |
Correlation:0.20297
  |
Component:332; Jaccard:0.11467
 |
Component:231; Jaccard:0.13838
 |
Component:181; Jaccard:0.15645
 |
Component:284; Jaccard:0.16154
 |
Component:182; Jaccard:0.16376
 |
Component:088; Jaccard:0.15445
 |
Component:284; Jaccard:0.1337
 |
Correlation:0.20799
  |
Correlation:0.18769
  |
Correlation:0.21874
  |
Correlation:0.094368
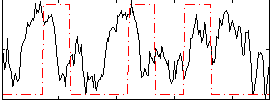 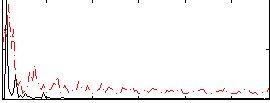 |
Correlation:0.20297
  |
Correlation:0.1466
  |
Correlation:0.094368
  |
Component:199; Jaccard:0.10977
 |
Component:088; Jaccard:0.13275
 |
Component:231; Jaccard:0.15409
 |
Component:181; Jaccard:0.15844
 |
Component:284; Jaccard:0.16242
 |
Component:284; Jaccard:0.15305
 |
Component:088; Jaccard:0.13082
 |
Correlation:0.22457
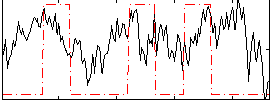 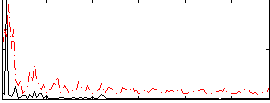 |
Correlation:0.1466
  |
Correlation:0.18769
  |
Correlation:0.21874
  |
Correlation:0.094368
  |
Correlation:0.094368
  |
Correlation:0.1466
  |
Cope: 02:"REL" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
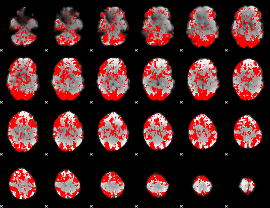 |
GLM binary result thresholded by 1.5
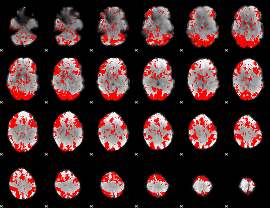 |
GLM binary result thresholded by 2
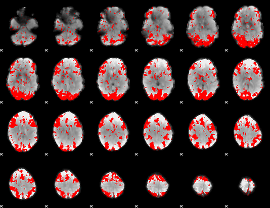 |
GLM binary result thresholded by 2.5
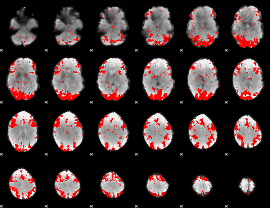 |
GLM binary result thresholded by 3
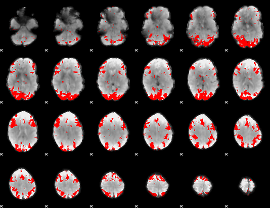 |
GLM binary result thresholded by 3.5
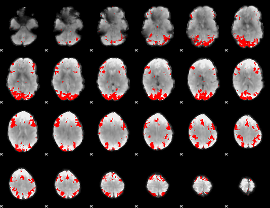 |
GLM binary result thresholded by 4
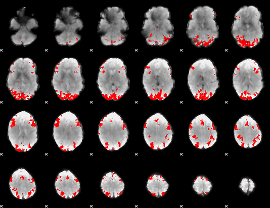 |
GLM Z-score result thresholded by 1
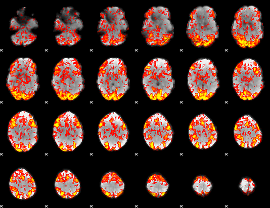 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
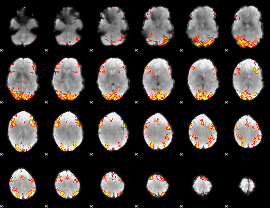 |
GLM Z-score result thresholded by 4
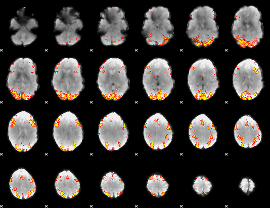 |
Component:268; Jaccard:0.12665
 |
Component:268; Jaccard:0.15982
 |
Component:268; Jaccard:0.19867
 |
Component:268; Jaccard:0.23593
 |
Component:268; Jaccard:0.26586
 |
Component:268; Jaccard:0.28258
 |
Component:268; Jaccard:0.29132
 |
Correlation:0.24544
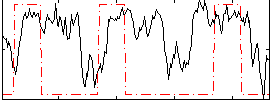 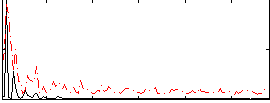 |
Correlation:0.24544
  |
Correlation:0.24544
  |
Correlation:0.24544
  |
Correlation:0.24544
  |
Correlation:0.24544
  |
Correlation:0.24544
  |
Component:310; Jaccard:0.1247
 |
Component:310; Jaccard:0.15148
 |
Component:161; Jaccard:0.18259
 |
Component:161; Jaccard:0.21587
 |
Component:161; Jaccard:0.24423
 |
Component:161; Jaccard:0.26252
 |
Component:161; Jaccard:0.27332
 |
Correlation:0.18228
  |
Correlation:0.18228
  |
Correlation:0.13423
  |
Correlation:0.13423
  |
Correlation:0.13423
  |
Correlation:0.13423
  |
Correlation:0.13423
  |
Component:161; Jaccard:0.12063
 |
Component:161; Jaccard:0.14937
 |
Component:310; Jaccard:0.18105
 |
Component:310; Jaccard:0.20778
 |
Component:130; Jaccard:0.23044
 |
Component:130; Jaccard:0.24384
 |
Component:130; Jaccard:0.24727
 |
Correlation:0.13423
  |
Correlation:0.13423
  |
Correlation:0.18228
  |
Correlation:0.18228
  |
Correlation:0.13809
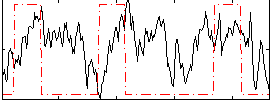 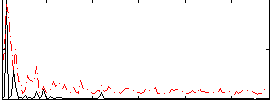 |
Correlation:0.13809
  |
Correlation:0.13809
  |
Component:199; Jaccard:0.11619
 |
Component:130; Jaccard:0.14407
 |
Component:130; Jaccard:0.17614
 |
Component:130; Jaccard:0.20632
 |
Component:310; Jaccard:0.22669
 |
Component:310; Jaccard:0.23558
 |
Component:310; Jaccard:0.23654
 |
Correlation:0.28246
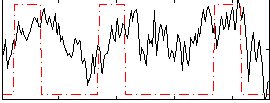 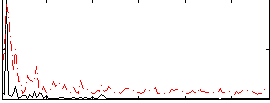 |
Correlation:0.13809
  |
Correlation:0.13809
  |
Correlation:0.13809
  |
Correlation:0.18228
  |
Correlation:0.18228
  |
Correlation:0.18228
  |
Component:130; Jaccard:0.11482
 |
Component:399; Jaccard:0.1437
 |
Component:399; Jaccard:0.17529
 |
Component:399; Jaccard:0.20255
 |
Component:399; Jaccard:0.21853
 |
Component:284; Jaccard:0.22623
 |
Component:284; Jaccard:0.23448
 |
Correlation:0.13809
  |
Correlation:0.30754
  |
Correlation:0.30754
  |
Correlation:0.30754
  |
Correlation:0.30754
  |
Correlation:0.38927
  |
Correlation:0.38927
  |
Component:399; Jaccard:0.11454
 |
Component:199; Jaccard:0.13795
 |
Component:071; Jaccard:0.16334
 |
Component:071; Jaccard:0.18642
 |
Component:284; Jaccard:0.20965
 |
Component:399; Jaccard:0.225
 |
Component:399; Jaccard:0.21925
 |
Correlation:0.30754
  |
Correlation:0.28246
  |
Correlation:0.22993
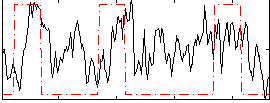 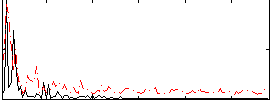 |
Correlation:0.22993
  |
Correlation:0.38927
  |
Correlation:0.30754
  |
Correlation:0.30754
  |
Component:071; Jaccard:0.11176
 |
Component:071; Jaccard:0.13625
 |
Component:284; Jaccard:0.15918
 |
Component:284; Jaccard:0.18636
 |
Component:071; Jaccard:0.20335
 |
Component:071; Jaccard:0.21121
 |
Component:071; Jaccard:0.2147
 |
Correlation:0.22993
  |
Correlation:0.22993
  |
Correlation:0.38927
  |
Correlation:0.38927
  |
Correlation:0.22993
  |
Correlation:0.22993
  |
Correlation:0.22993
  |
Component:326; Jaccard:0.11025
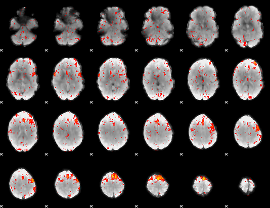 |
Component:284; Jaccard:0.13043
 |
Component:199; Jaccard:0.15797
 |
Component:199; Jaccard:0.17447
 |
Component:088; Jaccard:0.18521
 |
Component:088; Jaccard:0.19418
 |
Component:088; Jaccard:0.19476
 |
Correlation:0.47996
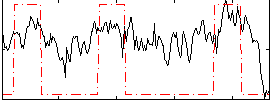  |
Correlation:0.38927
  |
Correlation:0.28246
  |
Correlation:0.28246
  |
Correlation:0.27201
  |
Correlation:0.27201
  |
Correlation:0.27201
  |
Component:231; Jaccard:0.10914
 |
Component:231; Jaccard:0.12506
 |
Component:088; Jaccard:0.14501
 |
Component:088; Jaccard:0.16767
 |
Component:199; Jaccard:0.1852
 |
Component:199; Jaccard:0.1893
 |
Component:199; Jaccard:0.18585
 |
Correlation:0.12882
  |
Correlation:0.12882
  |
Correlation:0.27201
  |
Correlation:0.27201
  |
Correlation:0.28246
  |
Correlation:0.28246
  |
Correlation:0.28246
  |
Component:284; Jaccard:0.10565
 |
Component:326; Jaccard:0.12404
 |
Component:231; Jaccard:0.13936
 |
Component:231; Jaccard:0.14981
 |
Component:231; Jaccard:0.15538
 |
Component:182; Jaccard:0.16083
 |
Component:182; Jaccard:0.16299
 |
Correlation:0.38927
  |
Correlation:0.47996
  |
Correlation:0.12882
  |
Correlation:0.12882
  |
Correlation:0.12882
  |
Correlation:0.21061
  |
Correlation:0.21061
  |
Cope: 03:"MATCH-REL" BACK
|
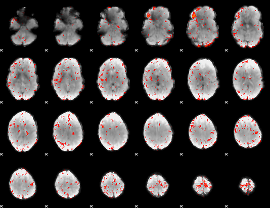
GLM result group-wise
 |
GLM binary result thresholded by 1
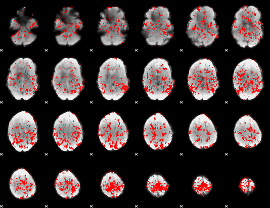 |
GLM binary result thresholded by 1.5
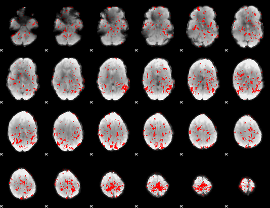 |
GLM binary result thresholded by 2
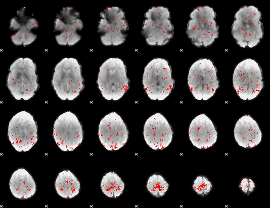 |
GLM binary result thresholded by 2.5
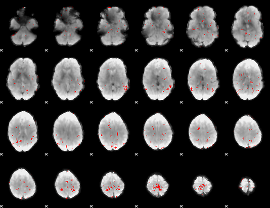 |
GLM binary result thresholded by 3
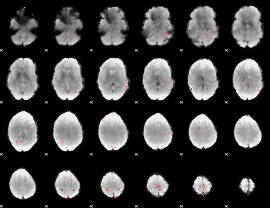 |
GLM binary result thresholded by 3.5
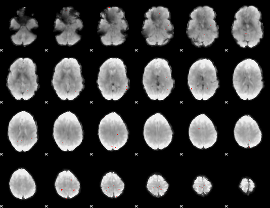 |
GLM binary result thresholded by 4
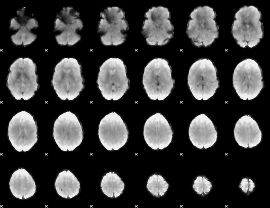 |
GLM Z-score result thresholded by 1
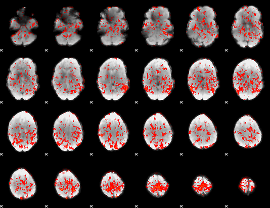 |
GLM Z-score result thresholded by 1.5
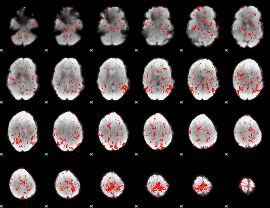 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
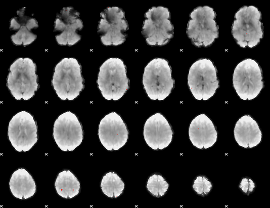 |
Component:355; Jaccard:0.092859
 |
Component:355; Jaccard:0.088075
 |
Component:147; Jaccard:0.072916
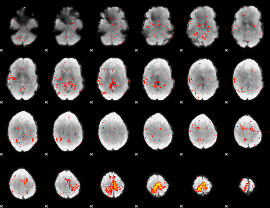 |
Component:170; Jaccard:0.043764
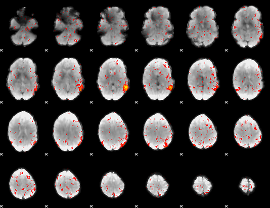 |
Component:170; Jaccard:0.0222
 |
Component:170; Jaccard:0.007955
 |
Component:108; Jaccard:0.003044
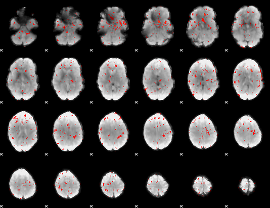 |
Correlation:0.21541
  |
Correlation:0.21541
  |
Correlation:0.25309
  |
Correlation:0.2227
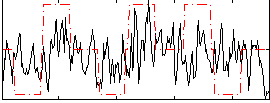 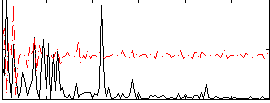 |
Correlation:0.2227
  |
Correlation:0.2227
  |
Correlation:0.080465
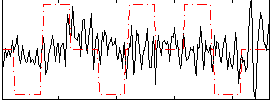 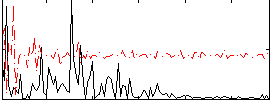 |
Component:147; Jaccard:0.085091
 |
Component:147; Jaccard:0.08762
 |
Component:087; Jaccard:0.068246
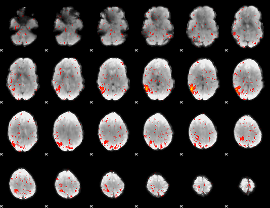 |
Component:147; Jaccard:0.043306
 |
Component:087; Jaccard:0.018942
 |
Component:131; Jaccard:0.007899
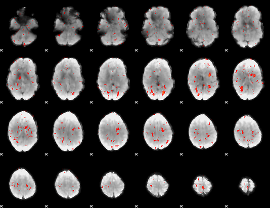 |
Component:170; Jaccard:0.002651
 |
Correlation:0.25309
  |
Correlation:0.25309
  |
Correlation:0.23429
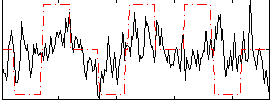 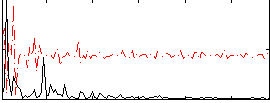 |
Correlation:0.25309
  |
Correlation:0.23429
  |
Correlation:-0.028677
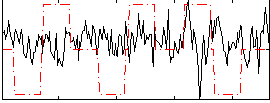 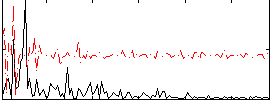 |
Correlation:0.2227
  |
Component:087; Jaccard:0.083499
 |
Component:087; Jaccard:0.084984
 |
Component:355; Jaccard:0.068188
 |
Component:355; Jaccard:0.041202
 |
Component:131; Jaccard:0.018456
 |
Component:087; Jaccard:0.006126
 |
Component:003; Jaccard:0.00182
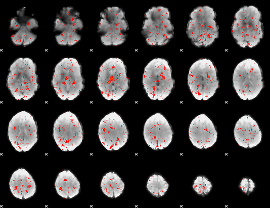 |
Correlation:0.23429
  |
Correlation:0.23429
  |
Correlation:0.21541
  |
Correlation:0.21541
  |
Correlation:-0.028677
  |
Correlation:0.23429
  |
Correlation:0.060312
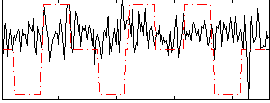  |
Component:137; Jaccard:0.08027
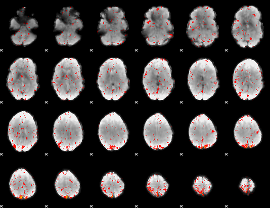 |
Component:137; Jaccard:0.076429
 |
Component:170; Jaccard:0.063462
 |
Component:087; Jaccard:0.041201
 |
Component:147; Jaccard:0.017286
 |
Component:108; Jaccard:0.00558
 |
Component:087; Jaccard:0.001779
 |
Correlation:0.19166
  |
Correlation:0.19166
  |
Correlation:0.2227
  |
Correlation:0.23429
  |
Correlation:0.25309
  |
Correlation:0.080465
  |
Correlation:0.23429
  |
Component:170; Jaccard:0.071482
 |
Component:170; Jaccard:0.073254
 |
Component:137; Jaccard:0.058639
 |
Component:007; Jaccard:0.035096
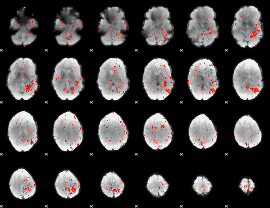 |
Component:355; Jaccard:0.0164
 |
Component:338; Jaccard:0.005187
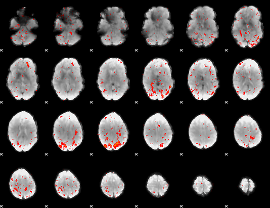 |
Component:338; Jaccard:0.001765
 |
Correlation:0.2227
  |
Correlation:0.2227
  |
Correlation:0.19166
  |
Correlation:0.19275
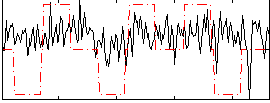 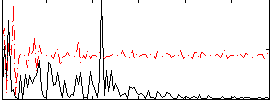 |
Correlation:0.21541
  |
Correlation:-0.049506
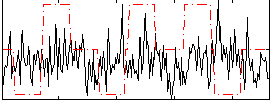 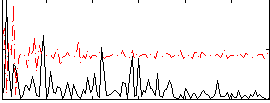 |
Correlation:-0.049506
  |
Component:271; Jaccard:0.068009
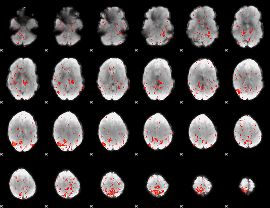 |
Component:271; Jaccard:0.065452
 |
Component:007; Jaccard:0.053107
 |
Component:137; Jaccard:0.033781
 |
Component:007; Jaccard:0.015199
 |
Component:355; Jaccard:0.004854
 |
Component:137; Jaccard:0.001735
 |
Correlation:0.09164
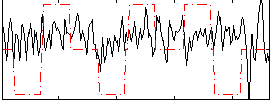  |
Correlation:0.09164
  |
Correlation:0.19275
  |
Correlation:0.19166
  |
Correlation:0.19275
  |
Correlation:0.21541
  |
Correlation:0.19166
  |
Component:280; Jaccard:0.066578
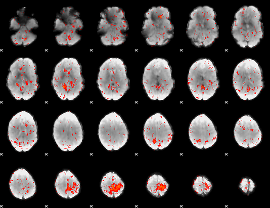 |
Component:007; Jaccard:0.062494
 |
Component:271; Jaccard:0.052441
 |
Component:131; Jaccard:0.031209
 |
Component:157; Jaccard:0.014678
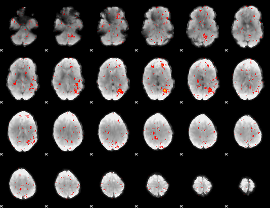 |
Component:147; Jaccard:0.004795
 |
Component:133; Jaccard:0.001357
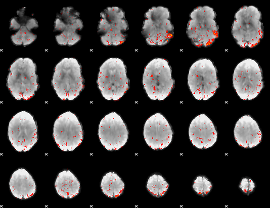 |
Correlation:0.18187
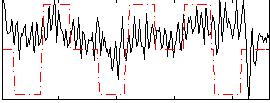 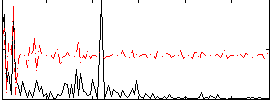 |
Correlation:0.19275
  |
Correlation:0.09164
  |
Correlation:-0.028677
  |
Correlation:0.062234
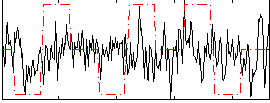 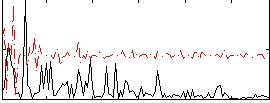 |
Correlation:0.25309
  |
Correlation:0.065802
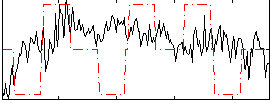 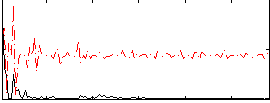 |
Component:392; Jaccard:0.065918
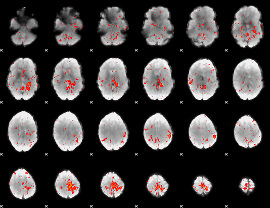 |
Component:392; Jaccard:0.061822
 |
Component:117; Jaccard:0.047112
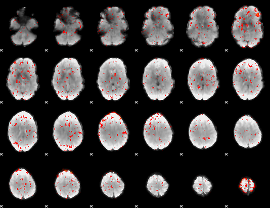 |
Component:271; Jaccard:0.029022
 |
Component:277; Jaccard:0.013262
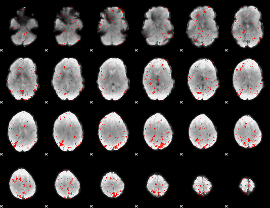 |
Component:007; Jaccard:0.00469
 |
Component:355; Jaccard:0.001343
 |
Correlation:0.19603
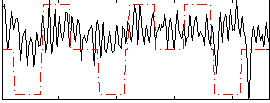 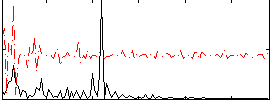 |
Correlation:0.19603
  |
Correlation:0.27736
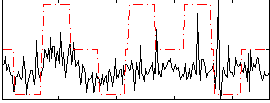  |
Correlation:0.09164
  |
Correlation:0.14113
  |
Correlation:0.19275
  |
Correlation:0.21541
  |
Component:117; Jaccard:0.064934
 |
Component:117; Jaccard:0.059691
 |
Component:277; Jaccard:0.046955
 |
Component:277; Jaccard:0.028346
 |
Component:137; Jaccard:0.013223
 |
Component:137; Jaccard:0.004273
 |
Component:131; Jaccard:0.001283
 |
Correlation:0.27736
  |
Correlation:0.27736
  |
Correlation:0.14113
  |
Correlation:0.14113
  |
Correlation:0.19166
  |
Correlation:0.19166
  |
Correlation:-0.028677
  |
Component:007; Jaccard:0.064808
 |
Component:277; Jaccard:0.059108
 |
Component:392; Jaccard:0.046103
 |
Component:338; Jaccard:0.028077
 |
Component:193; Jaccard:0.012242
 |
Component:117; Jaccard:0.00424
 |
Component:371; Jaccard:0.001056
 |
Correlation:0.19275
  |
Correlation:0.14113
  |
Correlation:0.19603
  |
Correlation:-0.049506
  |
Correlation:0.093297
  |
Correlation:0.27736
  |
Correlation:0.20263
  |
Cope: 04:"REL-MATCH" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:326; Jaccard:0.12541
 |
Component:326; Jaccard:0.15202
 |
Component:326; Jaccard:0.17504
 |
Component:326; Jaccard:0.18046
 |
Component:284; Jaccard:0.18006
 |
Component:284; Jaccard:0.16555
 |
Component:284; Jaccard:0.13222
 |
Correlation:0.3715
  |
Correlation:0.3715
  |
Correlation:0.3715
  |
Correlation:0.3715
  |
Correlation:0.17479
  |
Correlation:0.17479
  |
Correlation:0.17479
  |
Component:213; Jaccard:0.11843
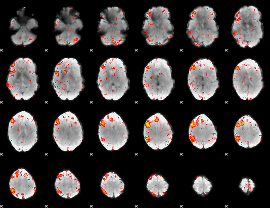 |
Component:213; Jaccard:0.14028
 |
Component:213; Jaccard:0.15809
 |
Component:284; Jaccard:0.17318
 |
Component:326; Jaccard:0.15493
 |
Component:161; Jaccard:0.1333
 |
Component:161; Jaccard:0.10437
 |
Correlation:0.033706
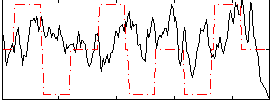 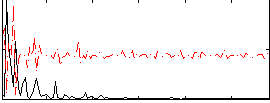 |
Correlation:0.033706
  |
Correlation:0.033706
  |
Correlation:0.17479
  |
Correlation:0.3715
  |
Correlation:-0.0081605
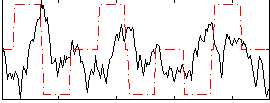 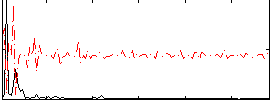 |
Correlation:-0.0081605
  |
Component:284; Jaccard:0.10327
 |
Component:284; Jaccard:0.12784
 |
Component:284; Jaccard:0.15233
 |
Component:213; Jaccard:0.15728
 |
Component:161; Jaccard:0.15154
 |
Component:326; Jaccard:0.11708
 |
Component:071; Jaccard:0.076951
 |
Correlation:0.17479
  |
Correlation:0.17479
  |
Correlation:0.17479
  |
Correlation:0.033706
  |
Correlation:-0.0081605
  |
Correlation:0.3715
  |
Correlation:0.13928
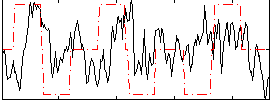 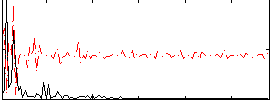 |
Component:199; Jaccard:0.10228
 |
Component:161; Jaccard:0.11827
 |
Component:161; Jaccard:0.13638
 |
Component:161; Jaccard:0.15039
 |
Component:213; Jaccard:0.13581
 |
Component:071; Jaccard:0.10292
 |
Component:310; Jaccard:0.074646
 |
Correlation:0.034306
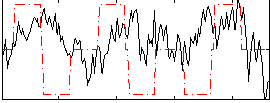 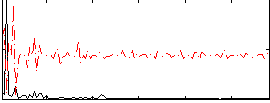 |
Correlation:-0.0081605
  |
Correlation:-0.0081605
  |
Correlation:-0.0081605
  |
Correlation:0.033706
  |
Correlation:0.13928
  |
Correlation:0.013531
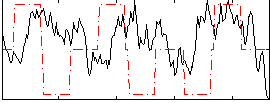 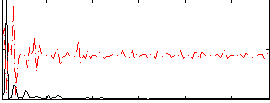 |
Component:161; Jaccard:0.10064
 |
Component:199; Jaccard:0.11587
 |
Component:199; Jaccard:0.12603
 |
Component:310; Jaccard:0.12882
 |
Component:310; Jaccard:0.12241
 |
Component:310; Jaccard:0.10005
 |
Component:326; Jaccard:0.074212
 |
Correlation:-0.0081605
  |
Correlation:0.034306
  |
Correlation:0.034306
  |
Correlation:0.013531
  |
Correlation:0.013531
  |
Correlation:0.013531
  |
Correlation:0.3715
  |
Component:310; Jaccard:0.1002
 |
Component:310; Jaccard:0.11362
 |
Component:310; Jaccard:0.12407
 |
Component:199; Jaccard:0.12477
 |
Component:071; Jaccard:0.1192
 |
Component:213; Jaccard:0.098826
 |
Component:199; Jaccard:0.069552
 |
Correlation:0.013531
  |
Correlation:0.013531
  |
Correlation:0.013531
  |
Correlation:0.034306
  |
Correlation:0.13928
  |
Correlation:0.033706
  |
Correlation:0.034306
  |
Component:399; Jaccard:0.092333
 |
Component:399; Jaccard:0.10592
 |
Component:071; Jaccard:0.11642
 |
Component:071; Jaccard:0.12264
 |
Component:199; Jaccard:0.11478
 |
Component:199; Jaccard:0.093593
 |
Component:185; Jaccard:0.068532
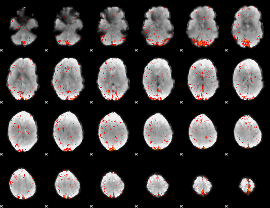 |
Correlation:0.076377
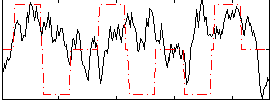  |
Correlation:0.076377
  |
Correlation:0.13928
  |
Correlation:0.13928
  |
Correlation:0.034306
  |
Correlation:0.034306
  |
Correlation:-0.074915
  |
Component:071; Jaccard:0.091092
 |
Component:071; Jaccard:0.1044
 |
Component:399; Jaccard:0.11545
 |
Component:399; Jaccard:0.11286
 |
Component:185; Jaccard:0.10446
 |
Component:185; Jaccard:0.089053
 |
Component:088; Jaccard:0.064973
 |
Correlation:0.13928
  |
Correlation:0.13928
  |
Correlation:0.076377
  |
Correlation:0.076377
  |
Correlation:-0.074915
  |
Correlation:-0.074915
  |
Correlation:0.07433
  |
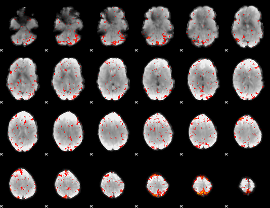
Component:316; Jaccard:0.08392
 |
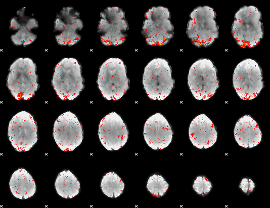
Component:008; Jaccard:0.092881
 |
Component:008; Jaccard:0.10259
 |
Component:185; Jaccard:0.10752
 |
Component:008; Jaccard:0.10121
 |
Component:088; Jaccard:0.087049
 |
Component:213; Jaccard:0.06208
 |
Correlation:-0.054088
  |
Correlation:0.096471
  |
Correlation:0.096471
  |
Correlation:-0.074915
  |
Correlation:0.096471
  |
Correlation:0.07433
  |
Correlation:0.033706
  |
Component:205; Jaccard:0.081217
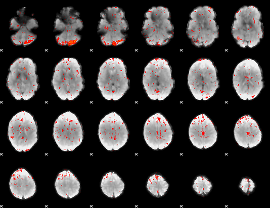 |
Component:185; Jaccard:0.091867
 |
Component:185; Jaccard:0.1012
 |
Component:008; Jaccard:0.10644
 |
Component:088; Jaccard:0.10076
 |
Component:008; Jaccard:0.083986
 |
Component:008; Jaccard:0.061869
 |
Correlation:0.24075
  |
Correlation:-0.074915
  |
Correlation:-0.074915
  |
Correlation:0.096471
  |
Correlation:0.07433
  |
Correlation:0.096471
  |
Correlation:0.096471
  |
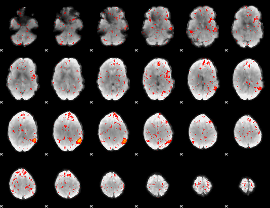
Cope: 05:"neg_MATCH" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
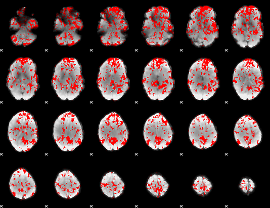 |
GLM binary result thresholded by 1.5
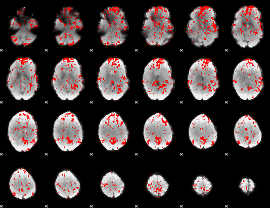 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
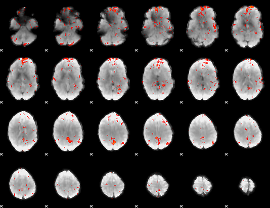 |
GLM Z-score result thresholded by 3
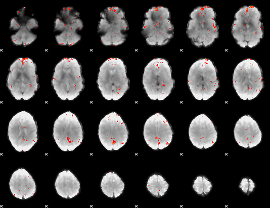 |
GLM Z-score result thresholded by 3.5
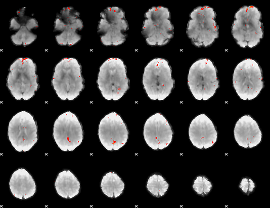 |
GLM Z-score result thresholded by 4
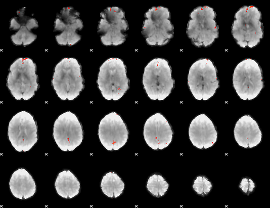 |
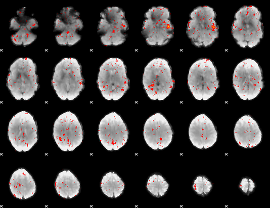
Component:251; Jaccard:0.11199
 |
Component:251; Jaccard:0.13807
 |
Component:251; Jaccard:0.15458
 |
Component:251; Jaccard:0.14727
 |
Component:251; Jaccard:0.1095
 |
Component:251; Jaccard:0.071189
 |
Component:251; Jaccard:0.039609
 |
Correlation:0.035239
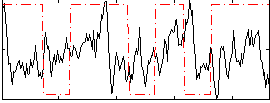 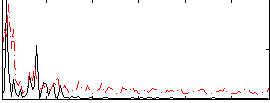 |
Correlation:0.035239
  |
Correlation:0.035239
  |
Correlation:0.035239
  |
Correlation:0.035239
  |
Correlation:0.035239
  |
Correlation:0.035239
  |
Component:234; Jaccard:0.1028
 |
Component:234; Jaccard:0.12339
 |
Component:234; Jaccard:0.13231
 |
Component:234; Jaccard:0.11537
 |
Component:110; Jaccard:0.080476
 |
Component:110; Jaccard:0.053462
 |
Component:110; Jaccard:0.031855
 |
Correlation:-0.13167
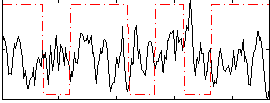 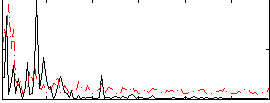 |
Correlation:-0.13167
  |
Correlation:-0.13167
  |
Correlation:-0.13167
  |
Correlation:0.17537
  |
Correlation:0.17537
  |
Correlation:0.17537
  |
Component:110; Jaccard:0.094018
 |
Component:110; Jaccard:0.10954
 |
Component:110; Jaccard:0.11547
 |
Component:110; Jaccard:0.10477
 |
Component:234; Jaccard:0.078408
 |
Component:068; Jaccard:0.04999
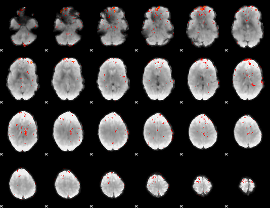 |
Component:068; Jaccard:0.029308
 |
Correlation:0.17537
  |
Correlation:0.17537
  |
Correlation:0.17537
  |
Correlation:0.17537
  |
Correlation:-0.13167
  |
Correlation:0.34622
  |
Correlation:0.34622
  |
Component:386; Jaccard:0.092796
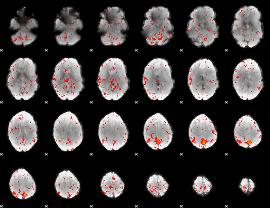 |
Component:203; Jaccard:0.098647
 |
Component:203; Jaccard:0.099922
 |
Component:203; Jaccard:0.091423
 |
Component:203; Jaccard:0.069914
 |
Component:203; Jaccard:0.044772
 |
Component:203; Jaccard:0.026308
 |
Correlation:-0.10589
  |
Correlation:0.12477
  |
Correlation:0.12477
  |
Correlation:0.12477
  |
Correlation:0.12477
  |
Correlation:0.12477
  |
Correlation:0.12477
  |
Component:203; Jaccard:0.086421
 |
Component:386; Jaccard:0.093026
 |
Component:386; Jaccard:0.083133
 |
Component:068; Jaccard:0.076115
 |
Component:068; Jaccard:0.067007
 |
Component:234; Jaccard:0.042822
 |
Component:278; Jaccard:0.022418
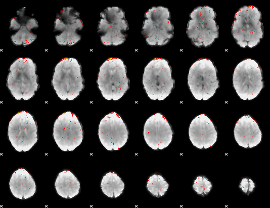 |
Correlation:0.12477
  |
Correlation:-0.10589
  |
Correlation:-0.10589
  |
Correlation:0.34622
  |
Correlation:0.34622
  |
Correlation:-0.13167
  |
Correlation:0.2421
  |
Component:252; Jaccard:0.080866
 |
Component:341; Jaccard:0.080084
 |
Component:232; Jaccard:0.07214
 |
Component:232; Jaccard:0.06609
 |
Component:278; Jaccard:0.055957
 |
Component:278; Jaccard:0.03793
 |
Component:234; Jaccard:0.020061
 |
Correlation:0.025143
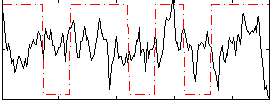 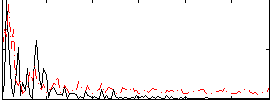 |
Correlation:0.25247
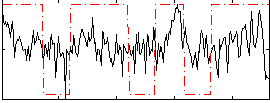 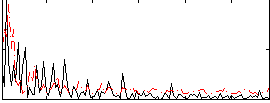 |
Correlation:0.1932
  |
Correlation:0.1932
  |
Correlation:0.2421
  |
Correlation:0.2421
  |
Correlation:-0.13167
  |
Component:341; Jaccard:0.080125
 |
Component:252; Jaccard:0.078958
 |
Component:252; Jaccard:0.071449
 |
Component:317; Jaccard:0.063114
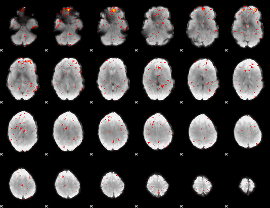 |
Component:232; Jaccard:0.051018
 |
Component:232; Jaccard:0.032721
 |
Component:285; Jaccard:0.01734
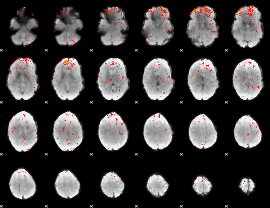 |
Correlation:0.25247
  |
Correlation:0.025143
  |
Correlation:0.025143
  |
Correlation:0.13441
  |
Correlation:0.1932
  |
Correlation:0.1932
  |
Correlation:0.27929
  |
Component:032; Jaccard:0.079483
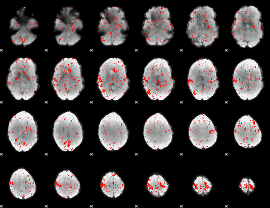 |
Component:002; Jaccard:0.075774
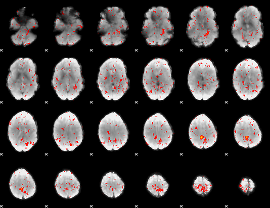 |
Component:068; Jaccard:0.07063
 |
Component:386; Jaccard:0.061099
 |
Component:317; Jaccard:0.048693
 |
Component:317; Jaccard:0.030077
 |
Component:232; Jaccard:0.017195
 |
Correlation:-0.013479
  |
Correlation:-0.018672
  |
Correlation:0.34622
  |
Correlation:-0.10589
  |
Correlation:0.13441
  |
Correlation:0.13441
  |
Correlation:0.1932
  |
Component:002; Jaccard:0.076015
 |
Component:032; Jaccard:0.074757
 |
Component:040; Jaccard:0.070124
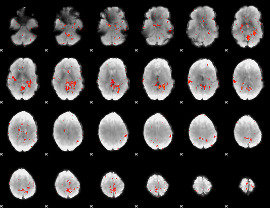 |
Component:278; Jaccard:0.060967
 |
Component:135; Jaccard:0.045148
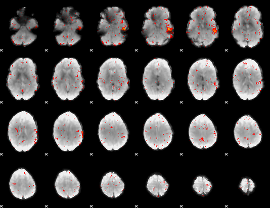 |
Component:285; Jaccard:0.028826
 |
Component:018; Jaccard:0.014479
 |
Correlation:-0.018672
  |
Correlation:-0.013479
  |
Correlation:-0.036647
  |
Correlation:0.2421
  |
Correlation:0.022351
  |
Correlation:0.27929
  |
Correlation:0.37506
  |
Component:250; Jaccard:0.073243
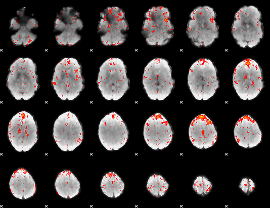 |
Component:053; Jaccard:0.071176
 |
Component:341; Jaccard:0.068536
 |
Component:135; Jaccard:0.060252
 |
Component:285; Jaccard:0.041534
 |
Component:135; Jaccard:0.025136
 |
Component:186; Jaccard:0.014352
 |
Correlation:0.069993
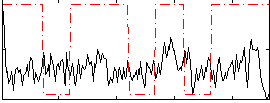 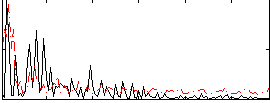 |
Correlation:-0.050831
  |
Correlation:0.25247
  |
Correlation:0.022351
  |
Correlation:0.27929
  |
Correlation:0.022351
  |
Correlation:-0.075208
  |
Cope: 06:"neg_REL" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
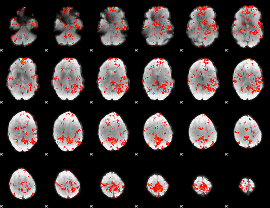 |
GLM Z-score result thresholded by 2
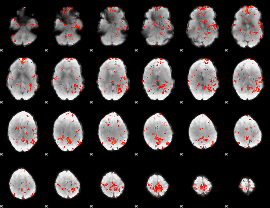 |
GLM Z-score result thresholded by 2.5
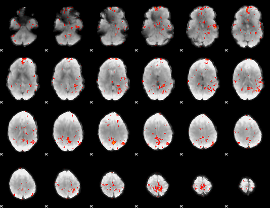 |
GLM Z-score result thresholded by 3
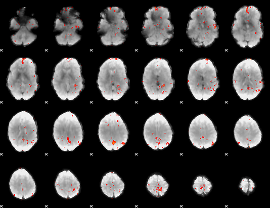 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:234; Jaccard:0.11236
 |
Component:234; Jaccard:0.13188
 |
Component:234; Jaccard:0.13951
 |
Component:234; Jaccard:0.13171
 |
Component:234; Jaccard:0.096983
 |
Component:234; Jaccard:0.055659
 |
Component:234; Jaccard:0.027251
 |
Correlation:0.19359
  |
Correlation:0.19359
  |
Correlation:0.19359
  |
Correlation:0.19359
  |
Correlation:0.19359
  |
Correlation:0.19359
  |
Correlation:0.19359
  |
Component:251; Jaccard:0.097992
 |
Component:251; Jaccard:0.10897
 |
Component:251; Jaccard:0.11131
 |
Component:251; Jaccard:0.10317
 |
Component:251; Jaccard:0.08
 |
Component:251; Jaccard:0.045482
 |
Component:251; Jaccard:0.023757
 |
Correlation:0.17942
  |
Correlation:0.17942
  |
Correlation:0.17942
  |
Correlation:0.17942
  |
Correlation:0.17942
  |
Correlation:0.17942
  |
Correlation:0.17942
  |
Component:110; Jaccard:0.094159
 |
Component:110; Jaccard:0.10552
 |
Component:110; Jaccard:0.10962
 |
Component:110; Jaccard:0.09861
 |
Component:110; Jaccard:0.074741
 |
Component:110; Jaccard:0.043812
 |
Component:110; Jaccard:0.023502
 |
Correlation:0.21299
  |
Correlation:0.21299
  |
Correlation:0.21299
  |
Correlation:0.21299
  |
Correlation:0.21299
  |
Correlation:0.21299
  |
Correlation:0.21299
  |
Component:355; Jaccard:0.092278
 |
Component:002; Jaccard:0.09469
 |
Component:002; Jaccard:0.088734
 |
Component:002; Jaccard:0.07099
 |
Component:002; Jaccard:0.048048
 |
Component:131; Jaccard:0.03561
 |
Component:131; Jaccard:0.021114
 |
Correlation:0.282
  |
Correlation:0.11554
  |
Correlation:0.11554
  |
Correlation:0.11554
  |
Correlation:0.11554
  |
Correlation:0.025513
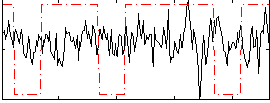 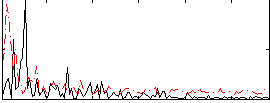 |
Correlation:0.025513
  |
Component:002; Jaccard:0.088545
 |
Component:355; Jaccard:0.091398
 |
Component:355; Jaccard:0.080836
 |
Component:040; Jaccard:0.070899
 |
Component:131; Jaccard:0.047922
 |
Component:186; Jaccard:0.028722
 |
Component:193; Jaccard:0.016492
 |
Correlation:0.11554
  |
Correlation:0.282
  |
Correlation:0.282
  |
Correlation:0.061272
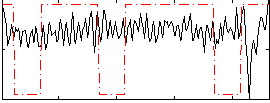 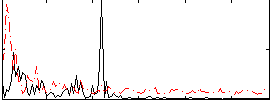 |
Correlation:0.025513
  |
Correlation:0.11176
  |
Correlation:0.20162
  |
Component:147; Jaccard:0.078239
 |
Component:147; Jaccard:0.079365
 |
Component:040; Jaccard:0.077595
 |
Component:355; Jaccard:0.061471
 |
Component:040; Jaccard:0.047587
 |
Component:178; Jaccard:0.027851
 |
Component:186; Jaccard:0.015618
 |
Correlation:0.21431
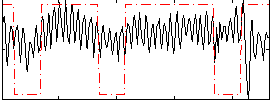 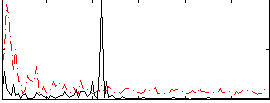 |
Correlation:0.21431
  |
Correlation:0.061272
  |
Correlation:0.282
  |
Correlation:0.061272
  |
Correlation:0.20748
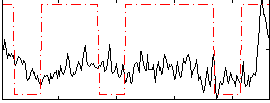  |
Correlation:0.11176
  |
Component:032; Jaccard:0.077725
 |
Component:040; Jaccard:0.076707
 |
Component:147; Jaccard:0.073633
 |
Component:203; Jaccard:0.058123
 |
Component:178; Jaccard:0.045704
 |
Component:203; Jaccard:0.026766
 |
Component:178; Jaccard:0.015318
 |
Correlation:-0.013554
  |
Correlation:0.061272
  |
Correlation:0.21431
  |
Correlation:0.24019
 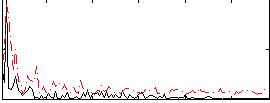 |
Correlation:0.20748
  |
Correlation:0.24019
  |
Correlation:0.20748
  |
Component:392; Jaccard:0.074503
 |
Component:203; Jaccard:0.074421
 |
Component:203; Jaccard:0.071059
 |
Component:157; Jaccard:0.057802
 |
Component:193; Jaccard:0.042764
 |
Component:193; Jaccard:0.026418
 |
Component:203; Jaccard:0.013951
 |
Correlation:0.16023
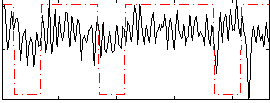 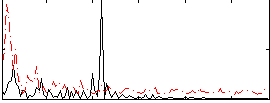 |
Correlation:0.24019
  |
Correlation:0.24019
  |
Correlation:0.073349
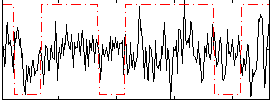  |
Correlation:0.20162
  |
Correlation:0.20162
  |
Correlation:0.24019
  |
Component:386; Jaccard:0.073592
 |
Component:032; Jaccard:0.074309
 |
Component:157; Jaccard:0.068237
 |
Component:007; Jaccard:0.055948
 |
Component:203; Jaccard:0.042698
 |
Component:040; Jaccard:0.024614
 |
Component:151; Jaccard:0.013619
 |
Correlation:0.012932
  |
Correlation:-0.013554
  |
Correlation:0.073349
  |
Correlation:0.16334
  |
Correlation:0.24019
  |
Correlation:0.061272
  |
Correlation:0.10619
  |
Component:203; Jaccard:0.073336
 |
Component:392; Jaccard:0.072856
 |
Component:007; Jaccard:0.068207
 |
Component:131; Jaccard:0.055779
 |
Component:186; Jaccard:0.042416
 |
Component:002; Jaccard:0.024215
 |
Component:278; Jaccard:0.013437
 |
Correlation:0.24019
  |
Correlation:0.16023
  |
Correlation:0.16334
  |
Correlation:0.025513
  |
Correlation:0.11176
  |
Correlation:0.11554
  |
Correlation:0.032238
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































