Task name: EMOTION, TR=0.72, totally 176 volumes, 10 waves, 6 copes BACK
|
Cope: 01:"FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
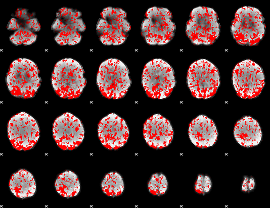 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
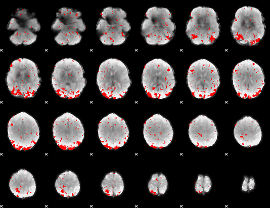 |
GLM binary result thresholded by 3
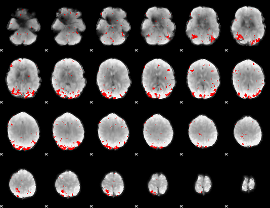 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
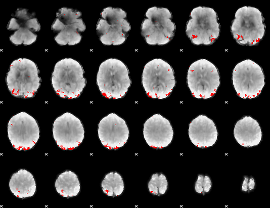 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
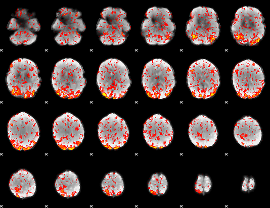 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
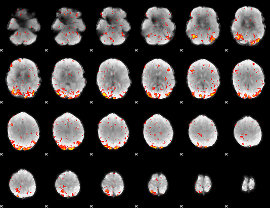 |
GLM Z-score result thresholded by 3
 |
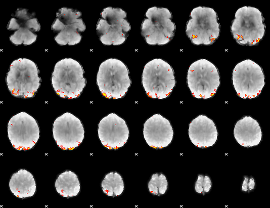
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:148; Jaccard:0.11988
 |
Component:312; Jaccard:0.15141
 |
Component:312; Jaccard:0.18487
 |
Component:312; Jaccard:0.20495
 |
Component:115; Jaccard:0.19762
 |
Component:115; Jaccard:0.17872
 |
Component:374; Jaccard:0.15517
 |
Correlation:0.10359
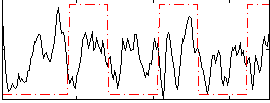  |
Correlation:0.61887
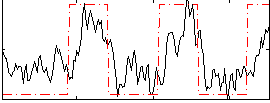 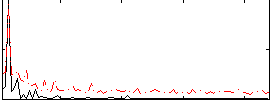 |
Correlation:0.61887
  |
Correlation:0.61887
  |
Correlation:0.49375
 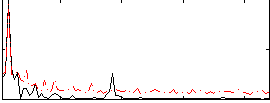 |
Correlation:0.49375
  |
Correlation:0.50465
  |
Component:312; Jaccard:0.11868
 |
Component:148; Jaccard:0.14455
 |
Component:115; Jaccard:0.17528
 |
Component:115; Jaccard:0.19877
 |
Component:312; Jaccard:0.19578
 |
Component:374; Jaccard:0.17411
 |
Component:115; Jaccard:0.15359
 |
Correlation:0.61887
  |
Correlation:0.10359
  |
Correlation:0.49375
  |
Correlation:0.49375
  |
Correlation:0.61887
  |
Correlation:0.50465
  |
Correlation:0.49375
  |
Component:115; Jaccard:0.10856
 |
Component:115; Jaccard:0.14219
 |
Component:148; Jaccard:0.16423
 |
Component:374; Jaccard:0.17572
 |
Component:374; Jaccard:0.18491
 |
Component:312; Jaccard:0.16776
 |
Component:312; Jaccard:0.13954
 |
Correlation:0.49375
  |
Correlation:0.49375
  |
Correlation:0.10359
  |
Correlation:0.50465
  |
Correlation:0.50465
  |
Correlation:0.61887
  |
Correlation:0.61887
  |
Component:251; Jaccard:0.10073
 |
Component:251; Jaccard:0.12288
 |
Component:374; Jaccard:0.1483
 |
Component:148; Jaccard:0.16625
 |
Component:255; Jaccard:0.16323
 |
Component:255; Jaccard:0.15111
 |
Component:255; Jaccard:0.12706
 |
Correlation:0.10526
  |
Correlation:0.10526
  |
Correlation:0.50465
  |
Correlation:0.10359
  |
Correlation:0.048431
 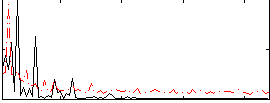 |
Correlation:0.048431
  |
Correlation:0.048431
  |
Component:186; Jaccard:0.091598
 |
Component:374; Jaccard:0.11773
 |
Component:251; Jaccard:0.14332
 |
Component:255; Jaccard:0.15829
 |
Component:148; Jaccard:0.14972
 |
Component:264; Jaccard:0.12513
 |
Component:264; Jaccard:0.10434
 |
Correlation:0.07091
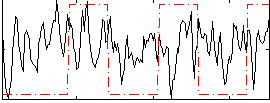 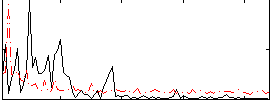 |
Correlation:0.50465
  |
Correlation:0.10526
  |
Correlation:0.048431
  |
Correlation:0.10359
  |
Correlation:0.15628
  |
Correlation:0.15628
  |
Component:051; Jaccard:0.09031
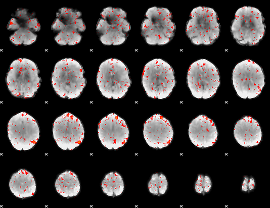 |
Component:264; Jaccard:0.10849
 |
Component:255; Jaccard:0.13208
 |
Component:251; Jaccard:0.15053
 |
Component:251; Jaccard:0.14094
 |
Component:148; Jaccard:0.12289
 |
Component:251; Jaccard:0.10025
 |
Correlation:0.19333
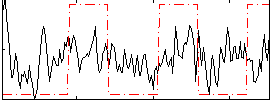  |
Correlation:0.15628
  |
Correlation:0.048431
  |
Correlation:0.10526
  |
Correlation:0.10526
  |
Correlation:0.10359
  |
Correlation:0.10526
  |
Component:374; Jaccard:0.08894
 |
Component:255; Jaccard:0.10287
 |
Component:264; Jaccard:0.12874
 |
Component:264; Jaccard:0.1398
 |
Component:264; Jaccard:0.13858
 |
Component:251; Jaccard:0.12168
 |
Component:148; Jaccard:0.095837
 |
Correlation:0.50465
  |
Correlation:0.048431
  |
Correlation:0.15628
  |
Correlation:0.15628
  |
Correlation:0.15628
  |
Correlation:0.10526
  |
Correlation:0.10359
  |
Component:264; Jaccard:0.088429
 |
Component:186; Jaccard:0.10218
 |
Component:186; Jaccard:0.10451
 |
Component:197; Jaccard:0.099082
 |
Component:197; Jaccard:0.10057
 |
Component:197; Jaccard:0.091082
 |
Component:197; Jaccard:0.079137
 |
Correlation:0.15628
  |
Correlation:0.07091
  |
Correlation:0.07091
  |
Correlation:-0.20089
  |
Correlation:-0.20089
  |
Correlation:-0.20089
  |
Correlation:-0.20089
  |
Component:255; Jaccard:0.079285
 |
Component:051; Jaccard:0.094168
 |
Component:197; Jaccard:0.087014
 |
Component:186; Jaccard:0.091331
 |
Component:153; Jaccard:0.071486
 |
Component:187; Jaccard:0.058324
 |
Component:187; Jaccard:0.048184
 |
Correlation:0.048431
  |
Correlation:0.19333
  |
Correlation:-0.20089
  |
Correlation:0.07091
  |
Correlation:0.29525
  |
Correlation:0.66622
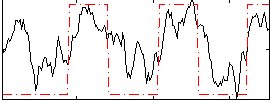  |
Correlation:0.66622
  |
Component:387; Jaccard:0.07464
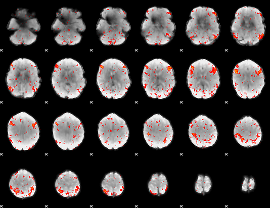 |
Component:153; Jaccard:0.081255
 |
Component:051; Jaccard:0.085536
 |
Component:153; Jaccard:0.081647
 |
Component:186; Jaccard:0.070097
 |
Component:153; Jaccard:0.055162
 |
Component:153; Jaccard:0.042297
 |
Correlation:0.14035
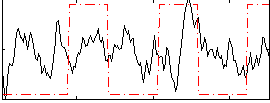 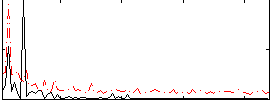 |
Correlation:0.29525
  |
Correlation:0.19333
  |
Correlation:0.29525
  |
Correlation:0.07091
  |
Correlation:0.29525
  |
Correlation:0.29525
  |
Cope: 02:"SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
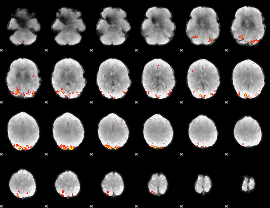 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
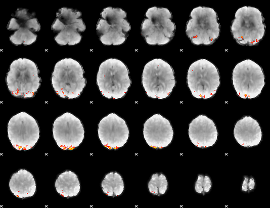 |
Component:148; Jaccard:0.14641
 |
Component:148; Jaccard:0.17822
 |
Component:255; Jaccard:0.21138
 |
Component:255; Jaccard:0.23417
 |
Component:255; Jaccard:0.22046
 |
Component:255; Jaccard:0.18623
 |
Component:255; Jaccard:0.14843
 |
Correlation:0.018069
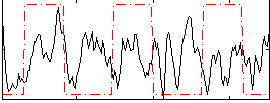  |
Correlation:0.018069
  |
Correlation:0.20321
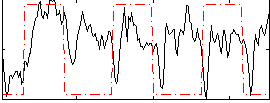  |
Correlation:0.20321
  |
Correlation:0.20321
  |
Correlation:0.20321
  |
Correlation:0.20321
  |
Component:251; Jaccard:0.12425
 |
Component:255; Jaccard:0.16595
 |
Component:148; Jaccard:0.18793
 |
Component:264; Jaccard:0.18024
 |
Component:197; Jaccard:0.16492
 |
Component:197; Jaccard:0.14515
 |
Component:197; Jaccard:0.11866
 |
Correlation:0.067781
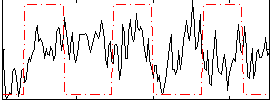  |
Correlation:0.20321
  |
Correlation:0.018069
  |
Correlation:0.034301
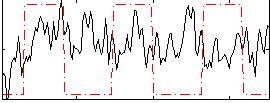 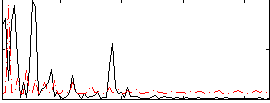 |
Correlation:0.3704
 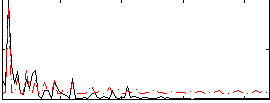 |
Correlation:0.3704
  |
Correlation:0.3704
  |
Component:264; Jaccard:0.12366
 |
Component:264; Jaccard:0.15882
 |
Component:264; Jaccard:0.18486
 |
Component:197; Jaccard:0.17843
 |
Component:264; Jaccard:0.15826
 |
Component:264; Jaccard:0.12941
 |
Component:264; Jaccard:0.10081
 |
Correlation:0.034301
  |
Correlation:0.034301
  |
Correlation:0.034301
  |
Correlation:0.3704
  |
Correlation:0.034301
  |
Correlation:0.034301
  |
Correlation:0.034301
  |
Component:255; Jaccard:0.12078
 |
Component:251; Jaccard:0.15301
 |
Component:197; Jaccard:0.1675
 |
Component:148; Jaccard:0.169
 |
Component:148; Jaccard:0.13149
 |
Component:251; Jaccard:0.10305
 |
Component:251; Jaccard:0.078325
 |
Correlation:0.20321
  |
Correlation:0.067781
  |
Correlation:0.3704
  |
Correlation:0.018069
  |
Correlation:0.018069
  |
Correlation:0.067781
  |
Correlation:0.067781
  |
Component:197; Jaccard:0.103
 |
Component:197; Jaccard:0.13771
 |
Component:251; Jaccard:0.16272
 |
Component:251; Jaccard:0.1541
 |
Component:251; Jaccard:0.12856
 |
Component:148; Jaccard:0.099585
 |
Component:148; Jaccard:0.073655
 |
Correlation:0.3704
  |
Correlation:0.3704
  |
Correlation:0.067781
  |
Correlation:0.067781
  |
Correlation:0.067781
  |
Correlation:0.018069
  |
Correlation:0.018069
  |
Component:186; Jaccard:0.096986
 |
Component:186; Jaccard:0.098809
 |
Component:374; Jaccard:0.10403
 |
Component:374; Jaccard:0.10552
 |
Component:374; Jaccard:0.089634
 |
Component:374; Jaccard:0.073348
 |
Component:374; Jaccard:0.060864
 |
Correlation:-0.060929
  |
Correlation:-0.060929
  |
Correlation:-0.28279
  |
Correlation:-0.28279
  |
Correlation:-0.28279
  |
Correlation:-0.28279
  |
Correlation:-0.28279
  |
Component:115; Jaccard:0.077404
 |
Component:374; Jaccard:0.090484
 |
Component:115; Jaccard:0.092998
 |
Component:115; Jaccard:0.087416
 |
Component:115; Jaccard:0.068637
 |
Component:115; Jaccard:0.051624
 |
Component:115; Jaccard:0.039857
 |
Correlation:-0.3675
  |
Correlation:-0.28279
  |
Correlation:-0.3675
  |
Correlation:-0.3675
  |
Correlation:-0.3675
  |
Correlation:-0.3675
  |
Correlation:-0.3675
  |
Component:374; Jaccard:0.074428
 |
Component:115; Jaccard:0.089292
 |
Component:186; Jaccard:0.089272
 |
Component:186; Jaccard:0.065818
 |
Component:312; Jaccard:0.050692
 |
Component:312; Jaccard:0.038339
 |
Component:312; Jaccard:0.027702
 |
Correlation:-0.28279
  |
Correlation:-0.3675
  |
Correlation:-0.060929
  |
Correlation:-0.060929
  |
Correlation:-0.54307
  |
Correlation:-0.54307
  |
Correlation:-0.54307
  |
Component:051; Jaccard:0.073613
 |
Component:312; Jaccard:0.076833
 |
Component:312; Jaccard:0.075415
 |
Component:312; Jaccard:0.065608
 |
Component:186; Jaccard:0.046658
 |
Component:186; Jaccard:0.033848
 |
Component:186; Jaccard:0.025249
 |
Correlation:-0.2696
  |
Correlation:-0.54307
  |
Correlation:-0.54307
  |
Correlation:-0.54307
  |
Correlation:-0.060929
  |
Correlation:-0.060929
  |
Correlation:-0.060929
  |
Component:312; Jaccard:0.072098
 |
Component:153; Jaccard:0.068883
 |
Component:153; Jaccard:0.064567
 |
Component:153; Jaccard:0.054919
 |
Component:153; Jaccard:0.043173
 |
Component:314; Jaccard:0.031698
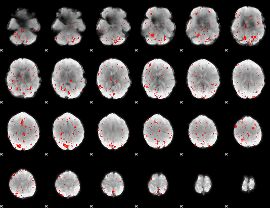 |
Component:314; Jaccard:0.023651
 |
Correlation:-0.54307
  |
Correlation:-0.22517
  |
Correlation:-0.22517
  |
Correlation:-0.22517
  |
Correlation:-0.22517
  |
Correlation:0.32337
  |
Correlation:0.32337
  |
Cope: 03:"FACES-SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
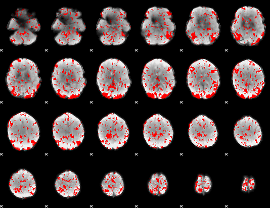 |
GLM binary result thresholded by 2.5
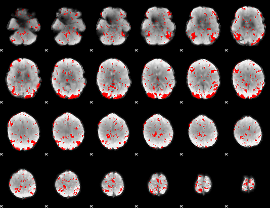 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
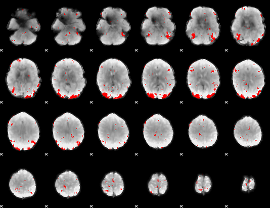 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
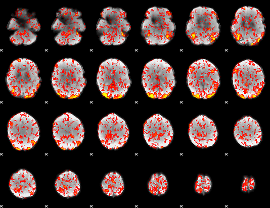 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
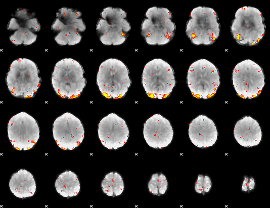 |
GLM Z-score result thresholded by 4
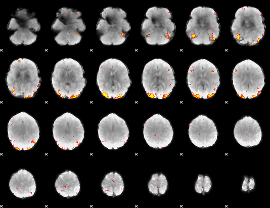 |
Component:187; Jaccard:0.12
 |
Component:187; Jaccard:0.15549
 |
Component:312; Jaccard:0.19529
 |
Component:312; Jaccard:0.23315
 |
Component:312; Jaccard:0.24837
 |
Component:312; Jaccard:0.23324
 |
Component:312; Jaccard:0.2012
 |
Correlation:0.73571
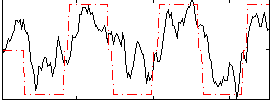 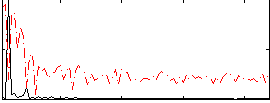 |
Correlation:0.73571
  |
Correlation:0.63026
  |
Correlation:0.63026
  |
Correlation:0.63026
  |
Correlation:0.63026
  |
Correlation:0.63026
  |
Component:312; Jaccard:0.11294
 |
Component:312; Jaccard:0.15167
 |
Component:187; Jaccard:0.19188
 |
Component:187; Jaccard:0.22128
 |
Component:187; Jaccard:0.23028
 |
Component:187; Jaccard:0.21615
 |
Component:187; Jaccard:0.19231
 |
Correlation:0.63026
  |
Correlation:0.63026
  |
Correlation:0.73571
  |
Correlation:0.73571
  |
Correlation:0.73571
  |
Correlation:0.73571
  |
Correlation:0.73571
  |
Component:115; Jaccard:0.093929
 |
Component:115; Jaccard:0.12158
 |
Component:115; Jaccard:0.15573
 |
Component:115; Jaccard:0.18756
 |
Component:115; Jaccard:0.20075
 |
Component:115; Jaccard:0.19333
 |
Component:115; Jaccard:0.17188
 |
Correlation:0.46715
  |
Correlation:0.46715
  |
Correlation:0.46715
  |
Correlation:0.46715
  |
Correlation:0.46715
  |
Correlation:0.46715
  |
Correlation:0.46715
  |
Component:387; Jaccard:0.088723
 |
Component:387; Jaccard:0.10056
 |
Component:374; Jaccard:0.11494
 |
Component:374; Jaccard:0.13856
 |
Component:374; Jaccard:0.15208
 |
Component:374; Jaccard:0.15137
 |
Component:374; Jaccard:0.14023
 |
Correlation:0.11753
  |
Correlation:0.11753
  |
Correlation:0.42711
  |
Correlation:0.42711
  |
Correlation:0.42711
  |
Correlation:0.42711
  |
Correlation:0.42711
  |
Component:243; Jaccard:0.088495
 |
Component:243; Jaccard:0.094776
 |
Component:387; Jaccard:0.10694
 |
Component:387; Jaccard:0.10478
 |
Component:387; Jaccard:0.090419
 |
Component:387; Jaccard:0.068468
 |
Component:387; Jaccard:0.049019
 |
Correlation:0.13894
  |
Correlation:0.13894
  |
Correlation:0.11753
  |
Correlation:0.11753
  |
Correlation:0.11753
  |
Correlation:0.11753
  |
Correlation:0.11753
  |
Component:238; Jaccard:0.075741
 |
Component:374; Jaccard:0.091394
 |
Component:243; Jaccard:0.093866
 |
Component:243; Jaccard:0.084686
 |
Component:090; Jaccard:0.070463
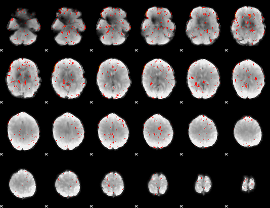 |
Component:235; Jaccard:0.05771
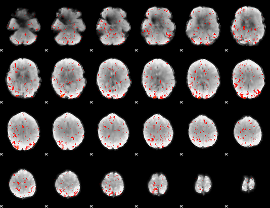 |
Component:235; Jaccard:0.048216
 |
Correlation:0.13957
  |
Correlation:0.42711
  |
Correlation:0.13894
  |
Correlation:0.13894
  |
Correlation:0.23239
  |
Correlation:0.28886
  |
Correlation:0.28886
  |
Component:017; Jaccard:0.074885
 |
Component:017; Jaccard:0.080284
 |
Component:133; Jaccard:0.086269
 |
Component:090; Jaccard:0.082856
 |
Component:133; Jaccard:0.069949
 |
Component:153; Jaccard:0.05444
 |
Component:153; Jaccard:0.044554
 |
Correlation:0.23586
  |
Correlation:0.23586
  |
Correlation:0.27081
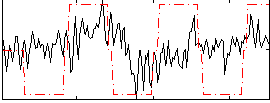  |
Correlation:0.23239
  |
Correlation:0.27081
  |
Correlation:0.28228
  |
Correlation:0.28228
  |
Component:051; Jaccard:0.074775
 |
Component:133; Jaccard:0.079155
 |
Component:090; Jaccard:0.084179
 |
Component:133; Jaccard:0.082545
 |
Component:243; Jaccard:0.067172
 |
Component:133; Jaccard:0.053713
 |
Component:133; Jaccard:0.038085
 |
Correlation:0.2511
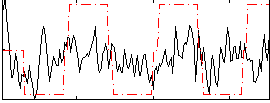 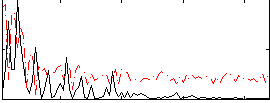 |
Correlation:0.27081
  |
Correlation:0.23239
  |
Correlation:0.27081
  |
Correlation:0.13894
  |
Correlation:0.27081
  |
Correlation:0.27081
  |
Component:047; Jaccard:0.073076
 |
Component:051; Jaccard:0.07903
 |
Component:017; Jaccard:0.083303
 |
Component:017; Jaccard:0.076724
 |
Component:235; Jaccard:0.06611
 |
Component:090; Jaccard:0.053404
 |
Component:251; Jaccard:0.03705
 |
Correlation:0.12123
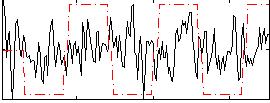  |
Correlation:0.2511
  |
Correlation:0.23586
  |
Correlation:0.23586
  |
Correlation:0.28886
  |
Correlation:0.23239
  |
Correlation:0.020328
  |
Component:374; Jaccard:0.071814
 |
Component:238; Jaccard:0.076685
 |
Component:051; Jaccard:0.07727
 |
Component:235; Jaccard:0.070005
 |
Component:017; Jaccard:0.065056
 |
Component:017; Jaccard:0.048993
 |
Component:148; Jaccard:0.036695
 |
Correlation:0.42711
  |
Correlation:0.13957
  |
Correlation:0.2511
  |
Correlation:0.28886
  |
Correlation:0.23586
  |
Correlation:0.23586
  |
Correlation:0.04639
  |
Cope: 04:"neg_FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
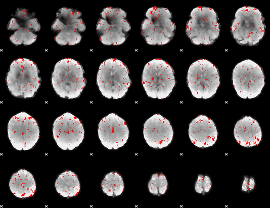 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
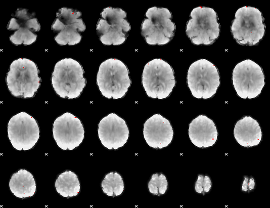 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:063; Jaccard:0.10683
 |
Component:063; Jaccard:0.12681
 |
Component:063; Jaccard:0.12566
 |
Component:063; Jaccard:0.10507
 |
Component:063; Jaccard:0.063641
 |
Component:063; Jaccard:0.032275
 |
Component:063; Jaccard:0.01469
 |
Correlation:0.27497
  |
Correlation:0.27497
  |
Correlation:0.27497
  |
Correlation:0.27497
  |
Correlation:0.27497
  |
Correlation:0.27497
  |
Correlation:0.27497
  |
Component:097; Jaccard:0.075007
 |
Component:326; Jaccard:0.072293
 |
Component:326; Jaccard:0.063314
 |
Component:326; Jaccard:0.046452
 |
Component:390; Jaccard:0.026827
 |
Component:390; Jaccard:0.016287
 |
Component:390; Jaccard:0.008492
 |
Correlation:0.23835
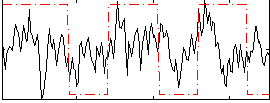 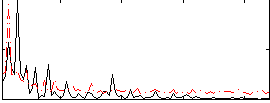 |
Correlation:0.1843
  |
Correlation:0.1843
  |
Correlation:0.1843
  |
Correlation:0.13421
  |
Correlation:0.13421
  |
Correlation:0.13421
  |
Component:342; Jaccard:0.070774
 |
Component:342; Jaccard:0.072141
 |
Component:296; Jaccard:0.063224
 |
Component:242; Jaccard:0.046163
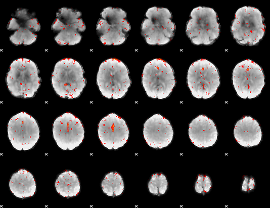 |
Component:242; Jaccard:0.026219
 |
Component:076; Jaccard:0.014501
 |
Component:076; Jaccard:0.008066
 |
Correlation:0.031155
  |
Correlation:0.031155
  |
Correlation:-0.036364
  |
Correlation:-0.033291
  |
Correlation:-0.033291
  |
Correlation:0.11964
  |
Correlation:0.11964
  |
Component:326; Jaccard:0.07064
 |
Component:097; Jaccard:0.07208
 |
Component:234; Jaccard:0.062048
 |
Component:234; Jaccard:0.043262
 |
Component:076; Jaccard:0.026159
 |
Component:242; Jaccard:0.013931
 |
Component:168; Jaccard:0.007786
 |
Correlation:0.1843
  |
Correlation:0.23835
  |
Correlation:0.091667
  |
Correlation:0.091667
  |
Correlation:0.11964
  |
Correlation:-0.033291
  |
Correlation:0.12728
  |
Component:234; Jaccard:0.070243
 |
Component:234; Jaccard:0.070624
 |
Component:097; Jaccard:0.061696
 |
Component:097; Jaccard:0.042865
 |
Component:326; Jaccard:0.025381
 |
Component:168; Jaccard:0.013834
 |
Component:242; Jaccard:0.006929
 |
Correlation:0.091667
  |
Correlation:0.091667
  |
Correlation:0.23835
  |
Correlation:0.23835
  |
Correlation:0.1843
  |
Correlation:0.12728
  |
Correlation:-0.033291
  |
Component:100; Jaccard:0.068233
 |
Component:229; Jaccard:0.068678
 |
Component:342; Jaccard:0.060015
 |
Component:296; Jaccard:0.041704
 |
Component:234; Jaccard:0.023876
 |
Component:326; Jaccard:0.013047
 |
Component:234; Jaccard:0.005522
 |
Correlation:0.11668
  |
Correlation:-0.025675
  |
Correlation:0.031155
  |
Correlation:-0.036364
  |
Correlation:0.091667
  |
Correlation:0.1843
  |
Correlation:0.091667
  |
Component:220; Jaccard:0.066404
 |
Component:296; Jaccard:0.068607
 |
Component:242; Jaccard:0.059897
 |
Component:076; Jaccard:0.041693
 |
Component:168; Jaccard:0.022968
 |
Component:234; Jaccard:0.010807
 |
Component:151; Jaccard:0.005096
 |
Correlation:-0.026565
  |
Correlation:-0.036364
  |
Correlation:-0.033291
  |
Correlation:0.11964
  |
Correlation:0.12728
  |
Correlation:0.091667
  |
Correlation:0.062899
  |
Component:229; Jaccard:0.065625
 |
Component:065; Jaccard:0.067625
 |
Component:229; Jaccard:0.058874
 |
Component:390; Jaccard:0.040475
 |
Component:097; Jaccard:0.021017
 |
Component:089; Jaccard:0.010384
 |
Component:065; Jaccard:0.004942
 |
Correlation:-0.025675
  |
Correlation:0.12831
  |
Correlation:-0.025675
  |
Correlation:0.13421
  |
Correlation:0.23835
  |
Correlation:0.20045
  |
Correlation:0.12831
  |
Component:065; Jaccard:0.064641
 |
Component:100; Jaccard:0.065188
 |
Component:065; Jaccard:0.056596
 |
Component:065; Jaccard:0.038981
 |
Component:296; Jaccard:0.020657
 |
Component:097; Jaccard:0.00982
 |
Component:326; Jaccard:0.004625
 |
Correlation:0.12831
  |
Correlation:0.11668
  |
Correlation:0.12831
  |
Correlation:0.12831
  |
Correlation:-0.036364
  |
Correlation:0.23835
  |
Correlation:0.1843
  |
Component:151; Jaccard:0.063512
 |
Component:242; Jaccard:0.063803
 |
Component:220; Jaccard:0.05242
 |
Component:229; Jaccard:0.038645
 |
Component:220; Jaccard:0.020615
 |
Component:220; Jaccard:0.009626
 |
Component:219; Jaccard:0.004424
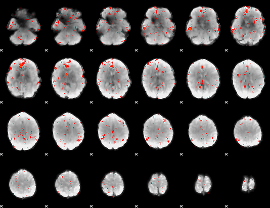 |
Correlation:0.062899
  |
Correlation:-0.033291
  |
Correlation:-0.026565
  |
Correlation:-0.025675
  |
Correlation:-0.026565
  |
Correlation:-0.026565
  |
Correlation:-0.0053087
  |
Cope: 05:"neg_SHAPES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
 |
GLM binary result thresholded by 1.5
 |
GLM binary result thresholded by 2
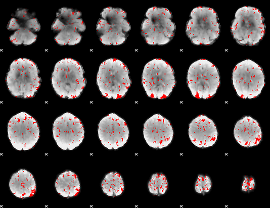 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
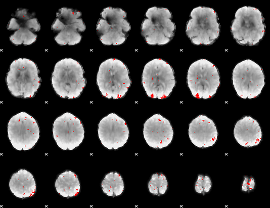 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
 |
GLM Z-score result thresholded by 1
 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:220; Jaccard:0.096554
 |
Component:220; Jaccard:0.10609
 |
Component:187; Jaccard:0.10569
 |
Component:187; Jaccard:0.087613
 |
Component:187; Jaccard:0.062255
 |
Component:187; Jaccard:0.040492
 |
Component:187; Jaccard:0.024131
 |
Correlation:0.25204
  |
Correlation:0.25204
  |
Correlation:0.69014
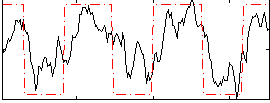  |
Correlation:0.69014
  |
Correlation:0.69014
  |
Correlation:0.69014
  |
Correlation:0.69014
  |
Component:151; Jaccard:0.096285
 |
Component:151; Jaccard:0.10502
 |
Component:220; Jaccard:0.10476
 |
Component:220; Jaccard:0.085118
 |
Component:220; Jaccard:0.056039
 |
Component:220; Jaccard:0.031447
 |
Component:220; Jaccard:0.018222
 |
Correlation:0.095024
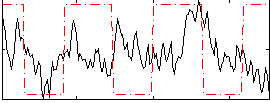  |
Correlation:0.095024
  |
Correlation:0.25204
  |
Correlation:0.25204
  |
Correlation:0.25204
  |
Correlation:0.25204
  |
Correlation:0.25204
  |
Component:187; Jaccard:0.09337
 |
Component:187; Jaccard:0.10469
 |
Component:151; Jaccard:0.09843
 |
Component:063; Jaccard:0.075563
 |
Component:063; Jaccard:0.046826
 |
Component:063; Jaccard:0.022977
 |
Component:151; Jaccard:0.011442
 |
Correlation:0.69014
  |
Correlation:0.69014
  |
Correlation:0.095024
  |
Correlation:-0.15495
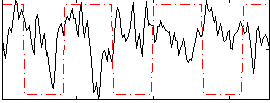  |
Correlation:-0.15495
  |
Correlation:-0.15495
  |
Correlation:0.095024
  |
Component:238; Jaccard:0.089162
 |
Component:063; Jaccard:0.095773
 |
Component:063; Jaccard:0.09452
 |
Component:151; Jaccard:0.072545
 |
Component:097; Jaccard:0.042987
 |
Component:151; Jaccard:0.022793
 |
Component:063; Jaccard:0.01113
 |
Correlation:0.25838
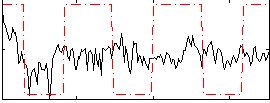  |
Correlation:-0.15495
  |
Correlation:-0.15495
  |
Correlation:0.095024
  |
Correlation:-0.16426
  |
Correlation:0.095024
  |
Correlation:-0.15495
  |
Component:097; Jaccard:0.084546
 |
Component:097; Jaccard:0.094295
 |
Component:097; Jaccard:0.089314
 |
Component:097; Jaccard:0.067686
 |
Component:151; Jaccard:0.042231
 |
Component:097; Jaccard:0.021423
 |
Component:224; Jaccard:0.00844
 |
Correlation:-0.16426
  |
Correlation:-0.16426
  |
Correlation:-0.16426
  |
Correlation:-0.16426
  |
Correlation:0.095024
  |
Correlation:-0.16426
  |
Correlation:0.030998
  |
Component:063; Jaccard:0.084331
 |
Component:238; Jaccard:0.089652
 |
Component:238; Jaccard:0.074631
 |
Component:224; Jaccard:0.05223
 |
Component:224; Jaccard:0.031231
 |
Component:224; Jaccard:0.017044
 |
Component:238; Jaccard:0.008418
 |
Correlation:-0.15495
  |
Correlation:0.25838
  |
Correlation:0.25838
  |
Correlation:0.030998
  |
Correlation:0.030998
  |
Correlation:0.030998
  |
Correlation:0.25838
  |
Component:243; Jaccard:0.078505
 |
Component:243; Jaccard:0.081047
 |
Component:243; Jaccard:0.069491
 |
Component:238; Jaccard:0.049522
 |
Component:243; Jaccard:0.02598
 |
Component:361; Jaccard:0.013192
 |
Component:097; Jaccard:0.007754
 |
Correlation:0.22032
  |
Correlation:0.22032
  |
Correlation:0.22032
  |
Correlation:0.25838
  |
Correlation:0.22032
  |
Correlation:0.18609
  |
Correlation:-0.16426
  |
Component:277; Jaccard:0.073403
 |
Component:224; Jaccard:0.073018
 |
Component:224; Jaccard:0.068629
 |
Component:243; Jaccard:0.047063
 |
Component:238; Jaccard:0.025387
 |
Component:238; Jaccard:0.012782
 |
Component:361; Jaccard:0.007439
 |
Correlation:0.15235
  |
Correlation:0.030998
  |
Correlation:0.030998
  |
Correlation:0.22032
  |
Correlation:0.25838
  |
Correlation:0.25838
  |
Correlation:0.18609
  |
Component:291; Jaccard:0.072887
 |
Component:373; Jaccard:0.070727
 |
Component:342; Jaccard:0.062827
 |
Component:275; Jaccard:0.044245
 |
Component:229; Jaccard:0.024953
 |
Component:243; Jaccard:0.012063
 |
Component:242; Jaccard:0.005831
 |
Correlation:0.1578
  |
Correlation:0.2292
  |
Correlation:-0.0056497
  |
Correlation:-0.019566
  |
Correlation:0.14312
  |
Correlation:0.22032
  |
Correlation:0.068517
  |
Component:373; Jaccard:0.07221
 |
Component:277; Jaccard:0.06974
 |
Component:373; Jaccard:0.059945
 |
Component:342; Jaccard:0.043588
 |
Component:361; Jaccard:0.024792
 |
Component:229; Jaccard:0.011963
 |
Component:277; Jaccard:0.005457
 |
Correlation:0.2292
  |
Correlation:0.15235
  |
Correlation:0.2292
  |
Correlation:-0.0056497
  |
Correlation:0.18609
  |
Correlation:0.14312
  |
Correlation:0.15235
  |
Cope: 06:"SHAPES-FACES" BACK
|
GLM result group-wise
 |
GLM binary result thresholded by 1
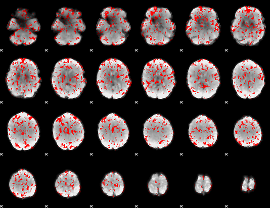 |
GLM binary result thresholded by 1.5
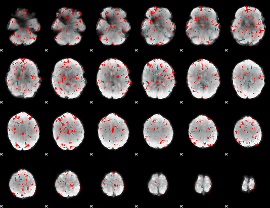 |
GLM binary result thresholded by 2
 |
GLM binary result thresholded by 2.5
 |
GLM binary result thresholded by 3
 |
GLM binary result thresholded by 3.5
 |
GLM binary result thresholded by 4
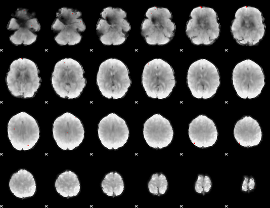 |
GLM Z-score result thresholded by 1
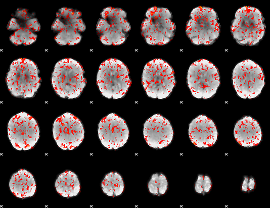 |
GLM Z-score result thresholded by 1.5
 |
GLM Z-score result thresholded by 2
 |
GLM Z-score result thresholded by 2.5
 |
GLM Z-score result thresholded by 3
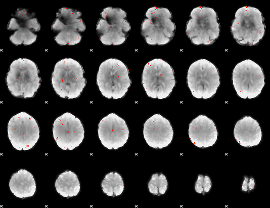 |
GLM Z-score result thresholded by 3.5
 |
GLM Z-score result thresholded by 4
 |
Component:063; Jaccard:0.073154
 |
Component:101; Jaccard:0.076272
 |
Component:168; Jaccard:0.068502
 |
Component:168; Jaccard:0.055167
 |
Component:168; Jaccard:0.031239
 |
Component:168; Jaccard:0.016514
 |
Component:168; Jaccard:0.009497
 |
Correlation:0.23319
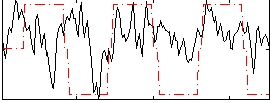 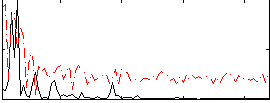 |
Correlation:0.18022
 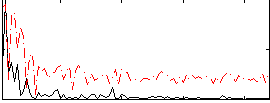 |
Correlation:0.13828
  |
Correlation:0.13828
  |
Correlation:0.13828
  |
Correlation:0.13828
  |
Correlation:0.13828
  |
Component:101; Jaccard:0.072595
 |
Component:063; Jaccard:0.0735
 |
Component:101; Jaccard:0.064603
 |
Component:063; Jaccard:0.042696
 |
Component:390; Jaccard:0.02481
 |
Component:390; Jaccard:0.01492
 |
Component:390; Jaccard:0.009086
 |
Correlation:0.18022
  |
Correlation:0.23319
  |
Correlation:0.18022
  |
Correlation:0.23319
  |
Correlation:0.13153
  |
Correlation:0.13153
  |
Correlation:0.13153
  |
Component:188; Jaccard:0.07106
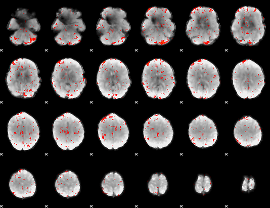 |
Component:168; Jaccard:0.071746
 |
Component:063; Jaccard:0.06222
 |
Component:050; Jaccard:0.040451
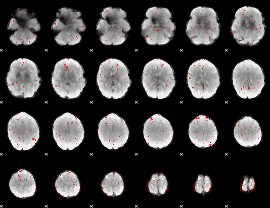 |
Component:076; Jaccard:0.02331
 |
Component:076; Jaccard:0.013095
 |
Component:076; Jaccard:0.007203
 |
Correlation:0.12614
  |
Correlation:0.13828
  |
Correlation:0.23319
  |
Correlation:0.26278
  |
Correlation:0.16722
  |
Correlation:0.16722
  |
Correlation:0.16722
  |
Component:326; Jaccard:0.069718
 |
Component:188; Jaccard:0.068589
 |
Component:326; Jaccard:0.057297
 |
Component:390; Jaccard:0.039862
 |
Component:063; Jaccard:0.023038
 |
Component:063; Jaccard:0.012622
 |
Component:063; Jaccard:0.006268
 |
Correlation:0.18342
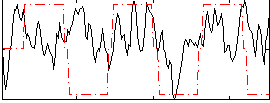  |
Correlation:0.12614
  |
Correlation:0.18342
  |
Correlation:0.13153
  |
Correlation:0.23319
  |
Correlation:0.23319
  |
Correlation:0.23319
  |
Component:234; Jaccard:0.066972
 |
Component:326; Jaccard:0.068106
 |
Component:050; Jaccard:0.056864
 |
Component:076; Jaccard:0.03876
 |
Component:050; Jaccard:0.02053
 |
Component:338; Jaccard:0.010037
 |
Component:338; Jaccard:0.006064
 |
Correlation:0.039382
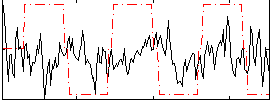  |
Correlation:0.18342
  |
Correlation:0.26278
  |
Correlation:0.16722
  |
Correlation:0.26278
  |
Correlation:0.066075
  |
Correlation:0.066075
  |
Component:065; Jaccard:0.06677
 |
Component:065; Jaccard:0.066494
 |
Component:370; Jaccard:0.055691
 |
Component:370; Jaccard:0.038425
 |
Component:338; Jaccard:0.019675
 |
Component:320; Jaccard:0.008681
 |
Component:089; Jaccard:0.004422
 |
Correlation:0.075413
  |
Correlation:0.075413
  |
Correlation:0.23875
  |
Correlation:0.23875
  |
Correlation:0.066075
  |
Correlation:0.13546
  |
Correlation:0.17921
  |
Component:168; Jaccard:0.064213
 |
Component:234; Jaccard:0.06429
 |
Component:076; Jaccard:0.054485
 |
Component:101; Jaccard:0.03826
 |
Component:320; Jaccard:0.019608
 |
Component:050; Jaccard:0.008373
 |
Component:320; Jaccard:0.004325
 |
Correlation:0.13828
  |
Correlation:0.039382
  |
Correlation:0.16722
  |
Correlation:0.18022
  |
Correlation:0.13546
  |
Correlation:0.26278
  |
Correlation:0.13546
  |
Component:209; Jaccard:0.063789
 |
Component:076; Jaccard:0.061142
 |
Component:188; Jaccard:0.051966
 |
Component:326; Jaccard:0.036799
 |
Component:065; Jaccard:0.019286
 |
Component:089; Jaccard:0.008353
 |
Component:219; Jaccard:0.004141
 |
Correlation:0.13588
  |
Correlation:0.16722
  |
Correlation:0.12614
  |
Correlation:0.18342
  |
Correlation:0.075413
  |
Correlation:0.17921
  |
Correlation:0.0027183
  |
Component:283; Jaccard:0.061364
 |
Component:370; Jaccard:0.059896
 |
Component:065; Jaccard:0.051909
 |
Component:320; Jaccard:0.036565
 |
Component:370; Jaccard:0.019243
 |
Component:065; Jaccard:0.008337
 |
Component:352; Jaccard:0.003881
 |
Correlation:0.0090153
  |
Correlation:0.23875
  |
Correlation:0.075413
  |
Correlation:0.13546
  |
Correlation:0.23875
  |
Correlation:0.075413
  |
Correlation:-0.027099
  |
Component:076; Jaccard:0.060669
 |
Component:320; Jaccard:0.059764
 |
Component:234; Jaccard:0.051572
 |
Component:065; Jaccard:0.033832
 |
Component:326; Jaccard:0.017792
 |
Component:188; Jaccard:0.008304
 |
Component:065; Jaccard:0.003769
 |
Correlation:0.16722
  |
Correlation:0.13546
  |
Correlation:0.039382
  |
Correlation:0.075413
  |
Correlation:0.18342
  |
Correlation:0.12614
  |
Correlation:0.075413
  |





































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































