HAFNI report on finalized components on task: RELATIONAL |
Totally 2 copes finalized |
Subject: 1 |
GLM result cope: 01, threshold: 3.5
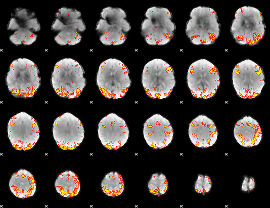 |
GLM result cope: 02, threshold: 4
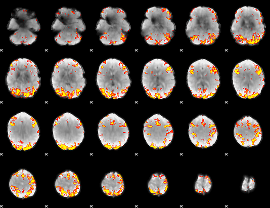 |
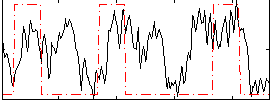
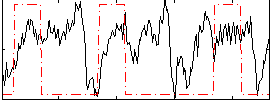
HAFNI result component: 087
Overlap rate: 0.43519
 |
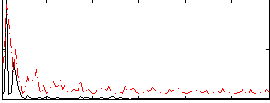
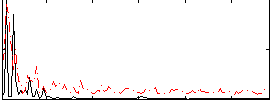
HAFNI result component: 087
Overlap rate: 0.31669
 |
Correlation: 0.27178
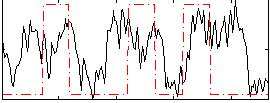 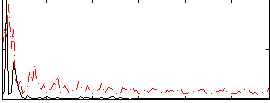 |
Correlation: 0.29962
  |
Subject: 2 |
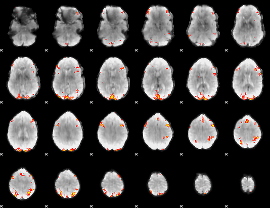
GLM result cope: 01, threshold: 3.5
 |
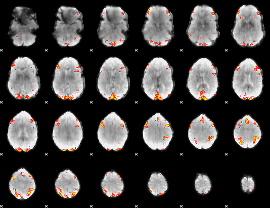
GLM result cope: 02, threshold: 3.5
 |
HAFNI result component: 234
Overlap rate: 0.45335
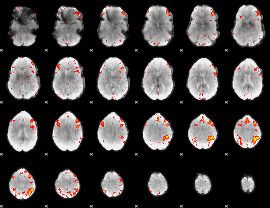 |
HAFNI result component: 234
Overlap rate: 0.44978
 |
Correlation: 0.18555
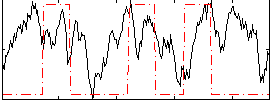  |
Correlation: 0.24285
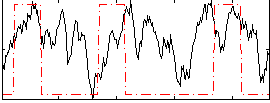  |
Subject: 3 |
GLM result cope: 01, threshold: 3.5
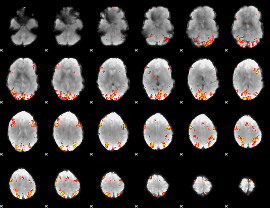 |
GLM result cope: 02, threshold: 3.5
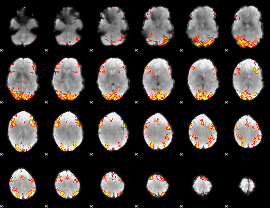 |
HAFNI result component: 268
Overlap rate: 0.59576
 |
HAFNI result component: 161
Overlap rate: 0.36721
 |
Correlation: 0.26572
  |
Correlation: 0.13423
  |
Subject: 4 |
GLM result cope: 01, threshold: 4
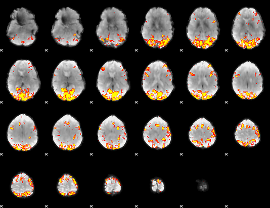 |
GLM result cope: 02, threshold: 4
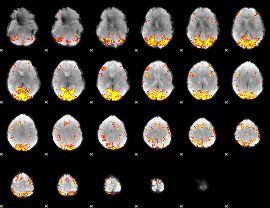 |
HAFNI result component: 384
Overlap rate: 0.44366
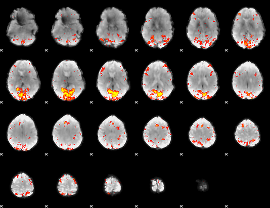 |
HAFNI result component: 384
Overlap rate: 0.45464
 |
Correlation: 0.25852
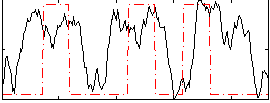 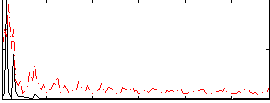 |
Correlation: 0.17056
  |
Subject: 5 |
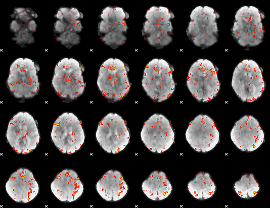
GLM result cope: 01, threshold: 3.5
 |
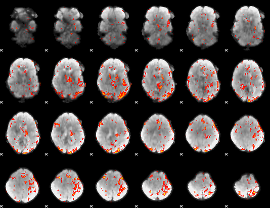
GLM result cope: 02, threshold: 3.5
 |
HAFNI result component: 316
Overlap rate: 0.39495
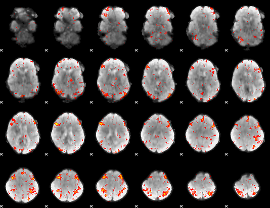 |
HAFNI result component: 316
Overlap rate: 0.30818
 |
Correlation: 0.22677
  |
Correlation: 0.2723
  |
Subject: 6 |
GLM result cope: 01, threshold: 3.5
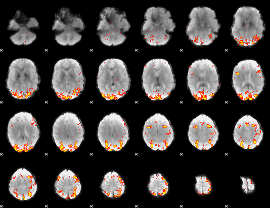 |
GLM result cope: 02, threshold: 3.5
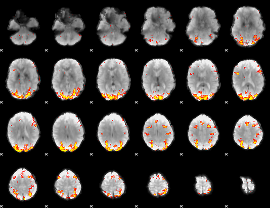 |
HAFNI result component: 157
Overlap rate: 0.57145
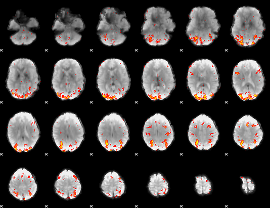 |
HAFNI result component: 157
Overlap rate: 0.57573
 |
Correlation: 0.27302
  |
Correlation: 0.24875
  |
Subject: 7 |
GLM result cope: 01, threshold: 3.5
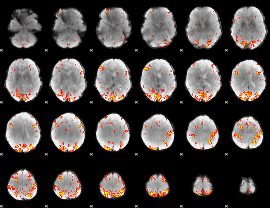 |
GLM result cope: 02, threshold: 3.5
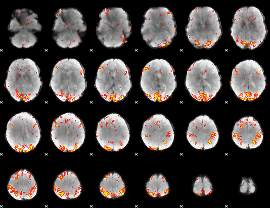 |
HAFNI result component: 217
Overlap rate: 0.47966
 |
HAFNI result component: 069
Overlap rate: 0.338
 |
Correlation: 0.18936
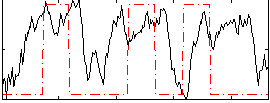 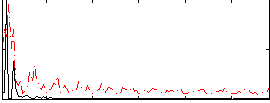 |
Correlation: 0.16414
  |
Subject: 8 |
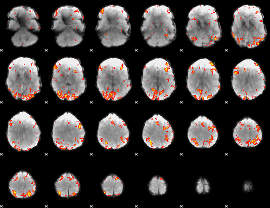
GLM result cope: 01, threshold: 3.5
 |
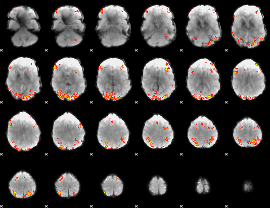
GLM result cope: 02, threshold: 4
 |
HAFNI result component: 370
Overlap rate: 0.422
 |
HAFNI result component: 287
Overlap rate: 0.34145
 |
Correlation: 0.21961
  |
Correlation: 0.1822
  |
Subject: 9 |
GLM result cope: 01, threshold: 4
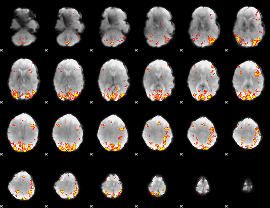 |
GLM result cope: 02, threshold: 3.5
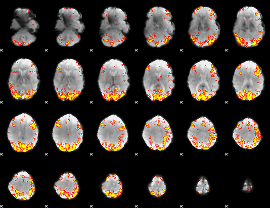 |
HAFNI result component: 337
Overlap rate: 0.34984
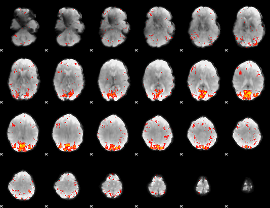 |
HAFNI result component: 337
Overlap rate: 0.28135
 |
Correlation: 0.21875
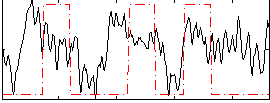  |
Correlation: 0.33918
  |
Subject: 10 |
GLM result cope: 01, threshold: 3.5
 |
GLM result cope: 02, threshold: 3.5
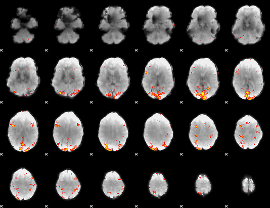 |
HAFNI result component: 179
Overlap rate: 0.31451
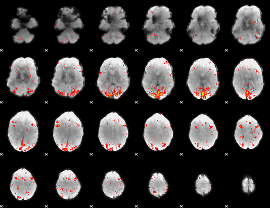 |
HAFNI result component: 371
Overlap rate: 0.52921
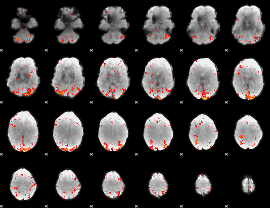 |
Correlation: 0.22976
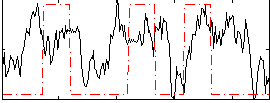  |
Correlation: 0.046982
  |
Subject: 11 |
GLM result cope: 01, threshold: 3.5
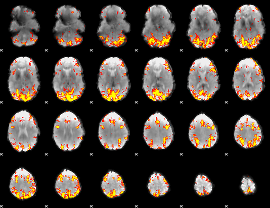 |
GLM result cope: 02, threshold: 3.5
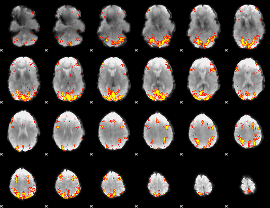 |
HAFNI result component: 300
Overlap rate: 0.35783
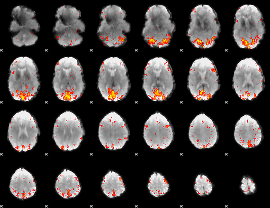 |
HAFNI result component: 300
Overlap rate: 0.49228
 |
Correlation: 0.29895
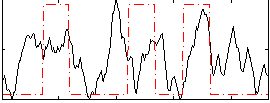 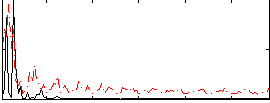 |
Correlation: 0.25507
  |
Subject: 12 |
GLM result cope: 01, threshold: 3.5
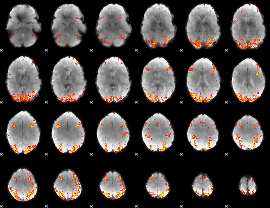 |
GLM result cope: 02, threshold: 3.5
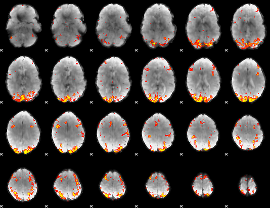 |
HAFNI result component: 181
Overlap rate: 0.4755
 |
HAFNI result component: 075
Overlap rate: 0.40274
 |
Correlation: 0.34526
  |
Correlation: 0.34359
  |
Subject: 13 |
GLM result cope: 01, threshold: 3.5
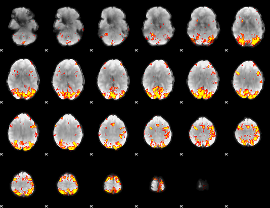 |
GLM result cope: 02, threshold: 4
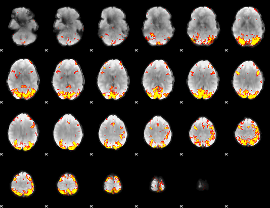 |
HAFNI result component: 199
Overlap rate: 0.45144
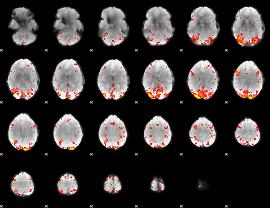 |
HAFNI result component: 199
Overlap rate: 0.40768
 |
Correlation: 0.10945
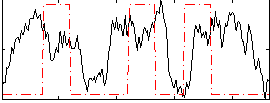  |
Correlation: 0.35203
  |
Subject: 14 |
GLM result cope: 01, threshold: 3.5
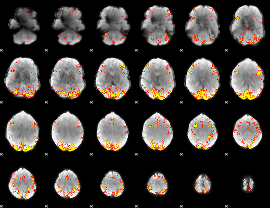 |
GLM result cope: 02, threshold: 3.5
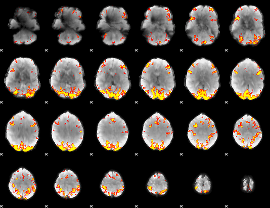 |
HAFNI result component: 343
Overlap rate: 0.50974
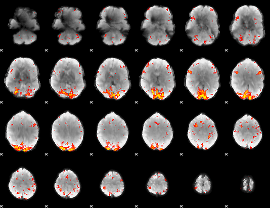 |
HAFNI result component: 343
Overlap rate: 0.49558
 |
Correlation: 0.1632
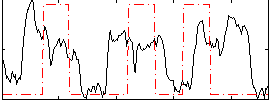 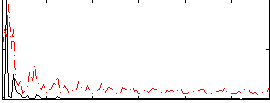 |
Correlation: 0.33779
  |
Subject: 15 |
GLM result cope: 01, threshold: 3.5
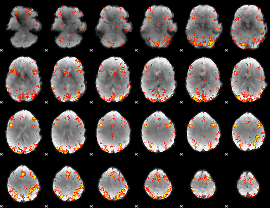 |
GLM result cope: 02, threshold: 4
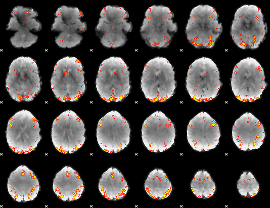 |
HAFNI result component: 232
Overlap rate: 0.23325
 |
HAFNI result component: 362
Overlap rate: 0.24389
 |
Correlation: -0.090768
  |
Correlation: 0.46361
  |
Subject: 16 |
GLM result cope: 01, threshold: 3.5
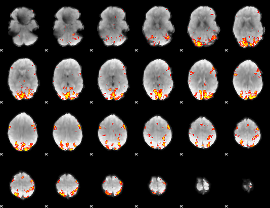 |
GLM result cope: 02, threshold: 3.5
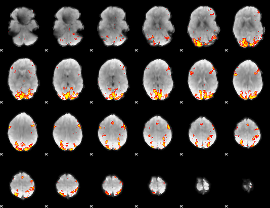 |
HAFNI result component: 010
Overlap rate: 0.41559
 |
HAFNI result component: 378
Overlap rate: 0.41551
 |
Correlation: 0.2946
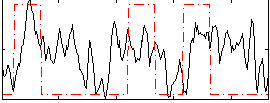 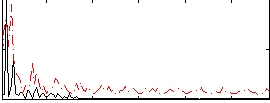 |
Correlation: 0.19494
  |
Subject: 17 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 18 |
GLM result cope: 01, threshold: 3.5
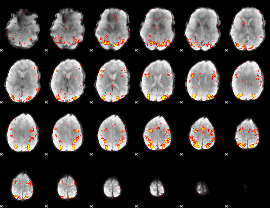 |
GLM result cope: 02, threshold: 3.5
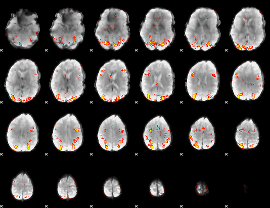 |
HAFNI result component: 123
Overlap rate: 0.42464
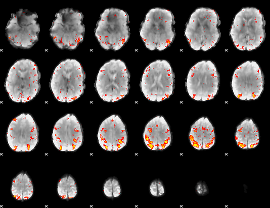 |
HAFNI result component: 123
Overlap rate: 0.40876
 |
Correlation: 0.30015
  |
Correlation: -0.047876
 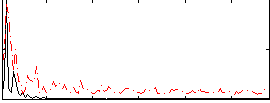 |
Subject: 19 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 20 |
GLM result cope: 01, threshold: 4
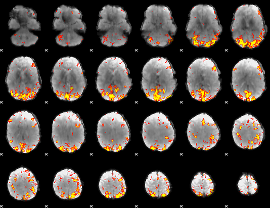 |
GLM result cope: 02, threshold: 4
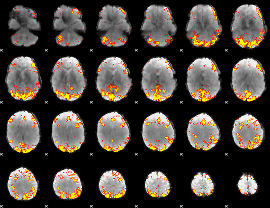 |
HAFNI result component: 163
Overlap rate: 0.36604
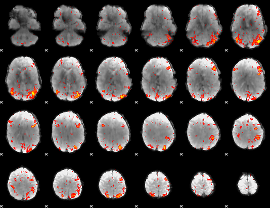 |
HAFNI result component: 063
Overlap rate: 0.27054
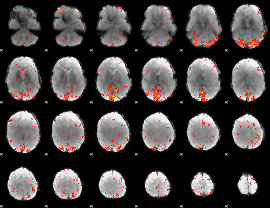 |
Correlation: 0.13895
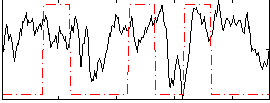 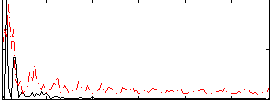 |
Correlation: 0.19687
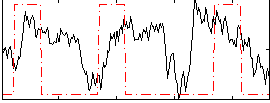  |
Subject: 21 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 22 |
GLM result cope: 01, threshold: 3
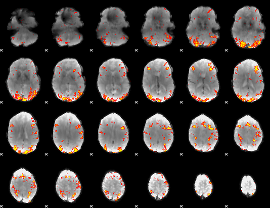 |
GLM result cope: 02, threshold: 2.5
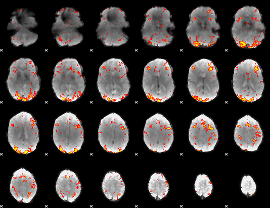 |
HAFNI result component: 322
Overlap rate: 0.27818
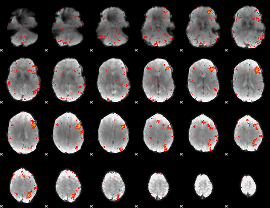 |
HAFNI result component: 096
Overlap rate: 0.5592
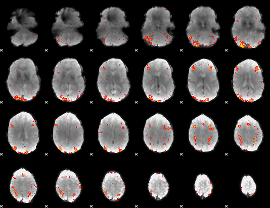 |
Correlation: 0.33352
  |
Correlation: 0.023038
  |
Subject: 23 |
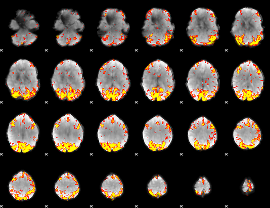
GLM result cope: 01, threshold: 3.5
 |
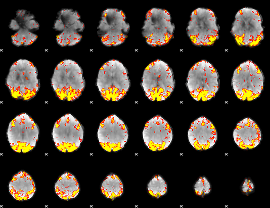
GLM result cope: 02, threshold: 3.5
 |
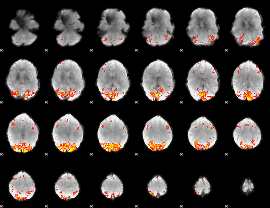
HAFNI result component: 345
Overlap rate: 0.40495
 |
HAFNI result component: 345
Overlap rate: 0.35956
 |
Correlation: 0.27556
  |
Correlation: 0.30028
  |
Subject: 24 |
GLM result cope: 01, threshold: 3
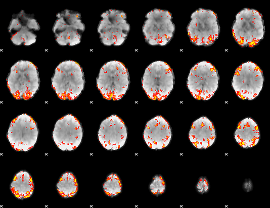 |
GLM result cope: 02, threshold: 3
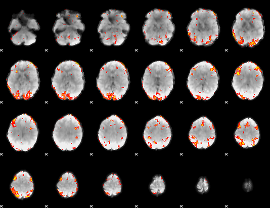 |
HAFNI result component: 103
Overlap rate: 0.3789
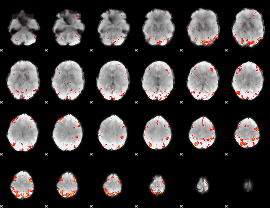 |
HAFNI result component: 350
Overlap rate: 0.30767
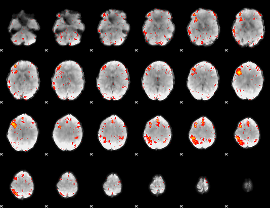 |
Correlation: 0.32626
  |
Correlation: 0.15576
  |
Subject: 25 |
GLM result cope: 01, threshold: 3.5
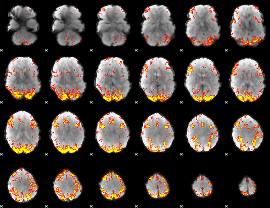 |
GLM result cope: 02, threshold: 3.5
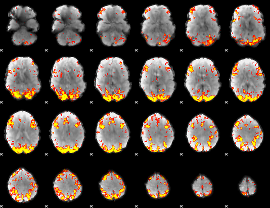 |
HAFNI result component: 012
Overlap rate: 0.35381
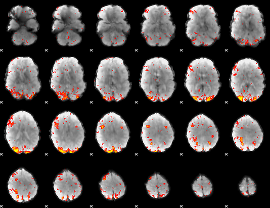 |
HAFNI result component: 383
Overlap rate: 0.34573
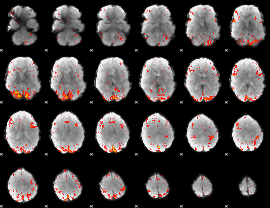 |
Correlation: 0.31935
  |
Correlation: 0.10713
  |
Subject: 26 |
GLM result cope: 01, threshold: 4
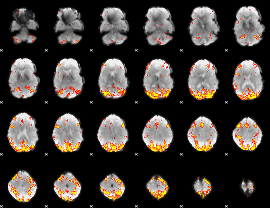 |
GLM result cope: 02, threshold: 3.5
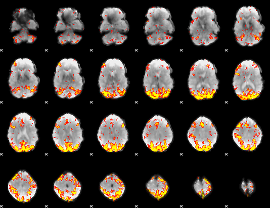 |
HAFNI result component: 142
Overlap rate: 0.28851
 |
HAFNI result component: 331
Overlap rate: 0.28616
 |
Correlation: 0.19764
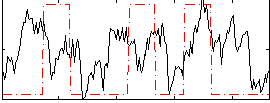 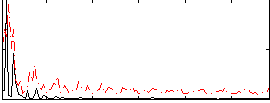 |
Correlation: 0.20172
  |
Subject: 27 |
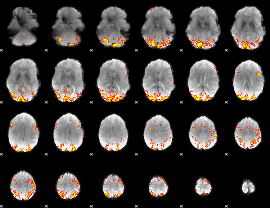
GLM result cope: 01, threshold: 3.5
 |
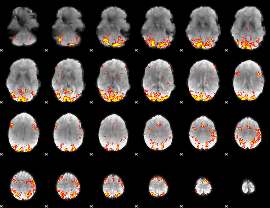
GLM result cope: 02, threshold: 3
 |
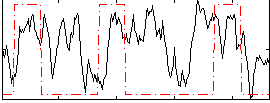
HAFNI result component: 275
Overlap rate: 0.45062
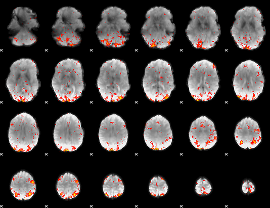 |
HAFNI result component: 275
Overlap rate: 0.45103
 |
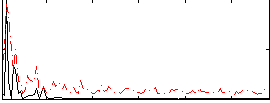
Correlation: 0.24167
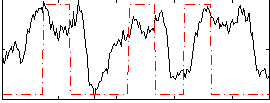  |
Correlation: 0.041772
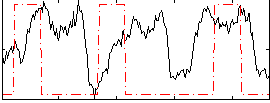  |
Subject: 28 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 29 |
GLM result cope: 01, threshold: 3.5
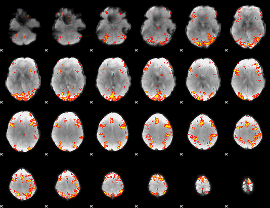 |
GLM result cope: 02, threshold: 4
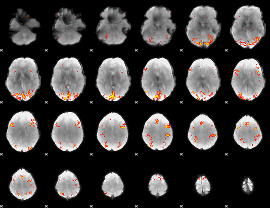 |
HAFNI result component: 397
Overlap rate: 0.48272
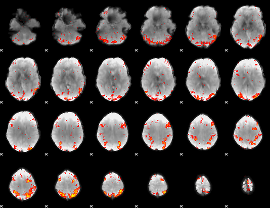 |
HAFNI result component: 379
Overlap rate: 0.49238
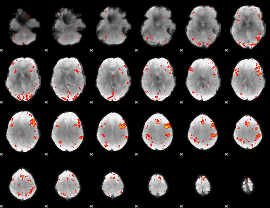 |
Correlation: 0.36184
  |
Correlation: 0.059687
  |
Subject: 30 |
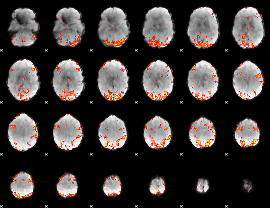
GLM result cope: 01, threshold: 3
 |
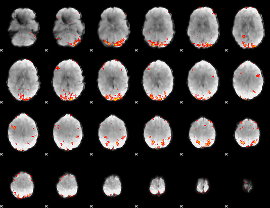
GLM result cope: 02, threshold: 3
 |
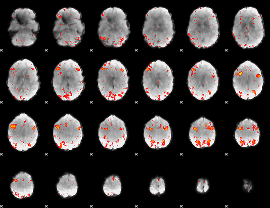
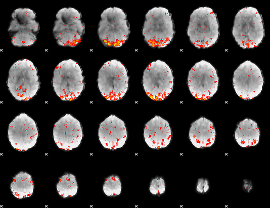
HAFNI result component: 308
Overlap rate: 0.41057
 |
HAFNI result component: 341
Overlap rate: 0.6953
 |
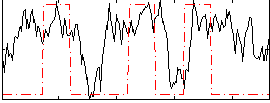
Correlation: 0.3277
  |
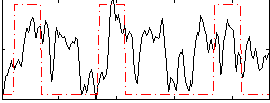
Correlation: 0.27895
  |
Subject: 31 |
GLM result cope: 01, threshold: 3
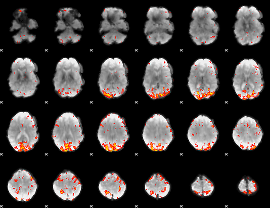 |
GLM result cope: 02, threshold: 2.5
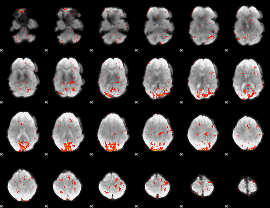 |
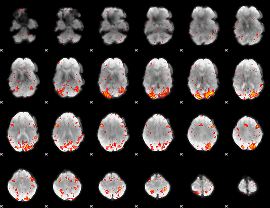
HAFNI result component: 249
Overlap rate: 0.58059
 |
HAFNI result component: 321
Overlap rate: 0.61449
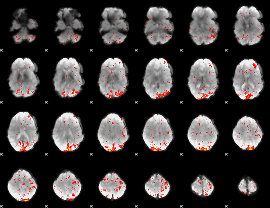 |
Correlation: 0.28431
  |
Correlation: 0.25818
  |
Subject: 32 |
GLM result cope: 01, threshold: 3
 |
GLM result cope: 02, threshold: 2.5
 |
HAFNI result component: 261
Overlap rate: 0.38653
 |
HAFNI result component: 261
Overlap rate: 0.51412
 |
Correlation: 0.17412
  |
Correlation: 0.17103
  |
Subject: 33 |
GLM result cope: 01, threshold: 3.5
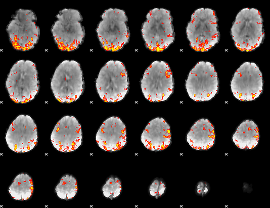 |
GLM result cope: 02, threshold: 3.5
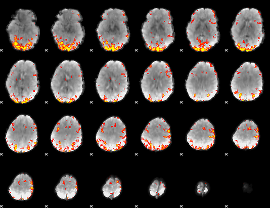 |
HAFNI result component: 274
Overlap rate: 0.41942
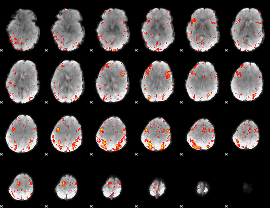 |
HAFNI result component: 144
Overlap rate: 0.46228
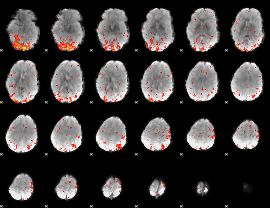 |
Correlation: 0.11271
  |
Correlation: 0.40347
  |
Subject: 34 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 35 |
GLM result cope: 01, threshold: 3.5
 |
GLM result cope: 02, threshold: 3.5
 |
HAFNI result component: 111
Overlap rate: 0.6617
 |
HAFNI result component: 059
Overlap rate: 0.31928
 |
Correlation: 0.28292
  |
Correlation: 0.06909
  |
Subject: 36 |
GLM result cope: 01, threshold: 4
 |
GLM result cope: 02, threshold: 4
 |
HAFNI result component: 087
Overlap rate: 0.77318
 |
HAFNI result component: 087
Overlap rate: 0.59584
 |
Correlation: 0.11899
  |
Correlation: 0.28348
  |
Subject: 37 |
GLM result cope: 01, threshold: 3.5
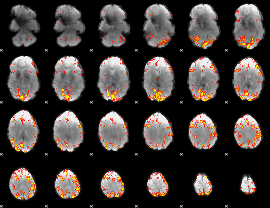 |
GLM result cope: 02, threshold: 4
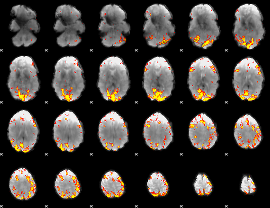 |
HAFNI result component: 327
Overlap rate: 0.35704
 |
HAFNI result component: 079
Overlap rate: 0.40573
 |
Correlation: 0.18412
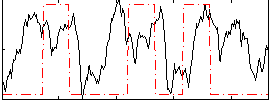  |
Correlation: 0.20297
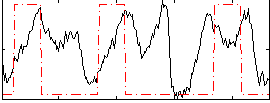  |
Subject: 38 |
GLM result cope: 01, threshold: 3.5
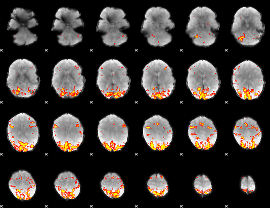 |
GLM result cope: 02, threshold: 3.5
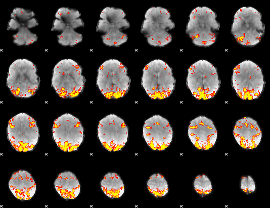 |
HAFNI result component: 274
Overlap rate: 0.51149
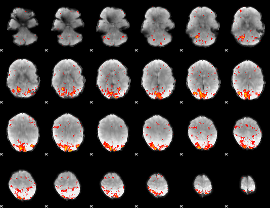 |
HAFNI result component: 353
Overlap rate: 0.48868
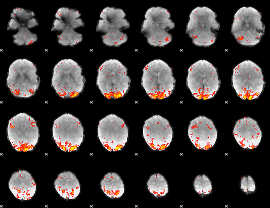 |
Correlation: 0.18544
  |
Correlation: 0.26429
  |
Subject: 39 |
GLM result cope: 01, threshold: 3.5
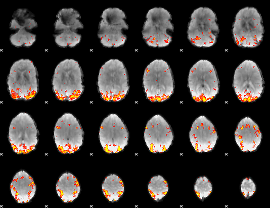 |
GLM result cope: 02, threshold: 3.5
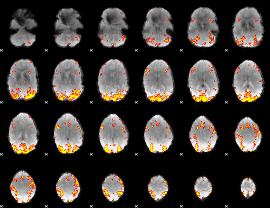 |
HAFNI result component: 042
Overlap rate: 0.39373
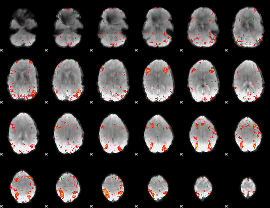 |
HAFNI result component: 163
Overlap rate: 0.37867
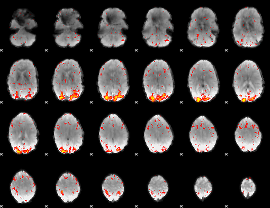 |
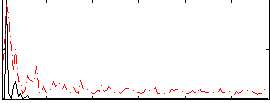
Correlation: 0.28823
  |
Correlation: 0.38812
  |
Subject: 40 |
GLM result cope: 01, threshold: 3.5
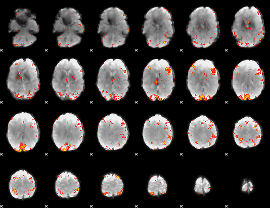 |
GLM result cope: 02, threshold: 3.5
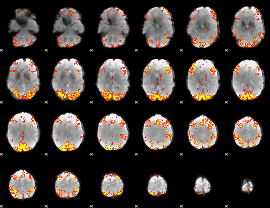 |
HAFNI result component: 280
Overlap rate: 0.32166
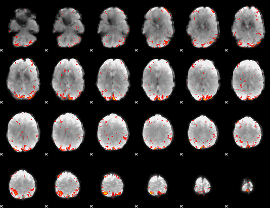 |
HAFNI result component: 153
Overlap rate: 0.31278
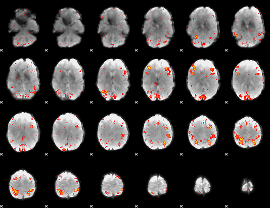 |
Correlation: 0.31272
  |
Correlation: 0.27404
  |
Subject: 41 |
GLM result cope: 01, threshold: 4
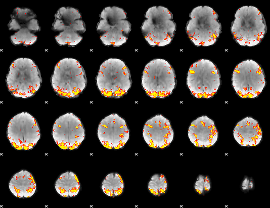 |
GLM result cope: 02, threshold: 3.5
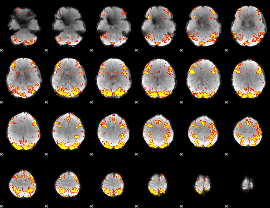 |
HAFNI result component: 053
Overlap rate: 0.40533
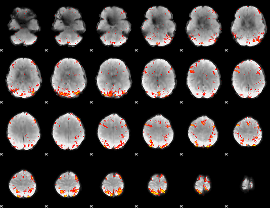 |
HAFNI result component: 246
Overlap rate: 0.38426
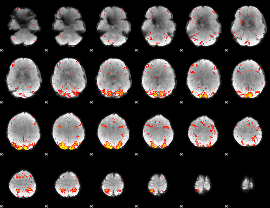 |
Correlation: 0.20374
  |
Correlation: 0.368
  |
Subject: 42 |
GLM result cope: 01, threshold: 3.5
 |
GLM result cope: 02, threshold: 4
 |
HAFNI result component: 206
Overlap rate: 0.35996
 |
HAFNI result component: 331
Overlap rate: 0.34483
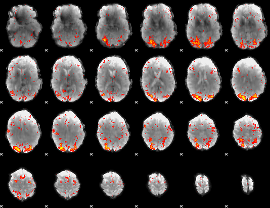 |
Correlation: 0.20153
  |
Correlation: 0.29253
  |
Subject: 43 |
GLM result cope: 01, threshold: 3.5
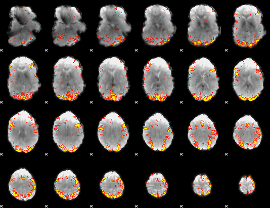 |
GLM result cope: 02, threshold: 4
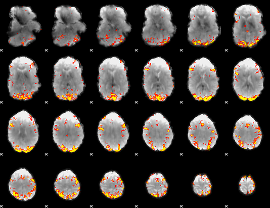 |
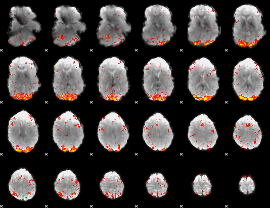
HAFNI result component: 141
Overlap rate: 0.40646
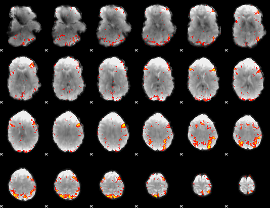 |
HAFNI result component: 245
Overlap rate: 0.51527
 |
Correlation: 0.24803
  |
Correlation: 0.34706
  |
Subject: 44 |
GLM result cope: 01, threshold: 3.5
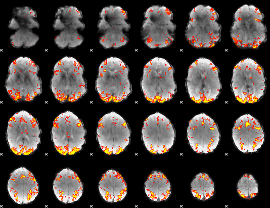 |
GLM result cope: 02, threshold: 3.5
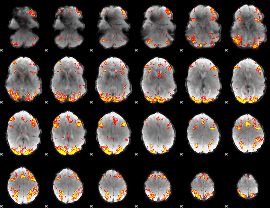 |
HAFNI result component: 294
Overlap rate: 0.32121
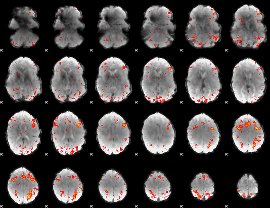 |
HAFNI result component: 303
Overlap rate: 0.40767
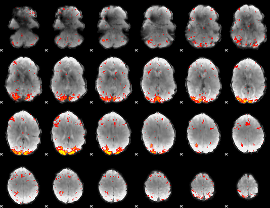 |
Correlation: 0.22724
  |
Correlation: 0.25751
  |
Subject: 45 |
GLM result cope: 01, threshold: 3.5
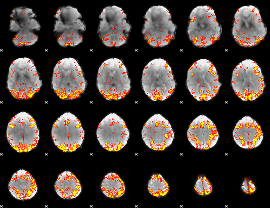 |
GLM result cope: 02, threshold: 4
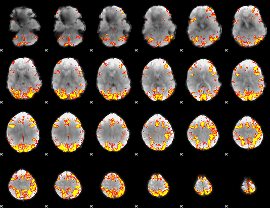 |
HAFNI result component: 343
Overlap rate: 0.34645
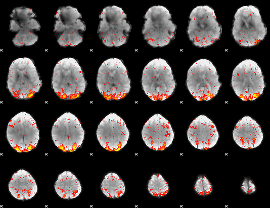 |
HAFNI result component: 104
Overlap rate: 0.26942
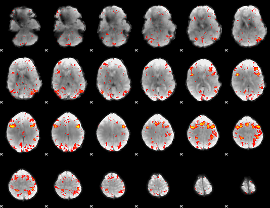 |
Correlation: 0.2897
  |
Correlation: 0.27159
  |
Subject: 46 |
GLM result cope: 01, threshold: 4
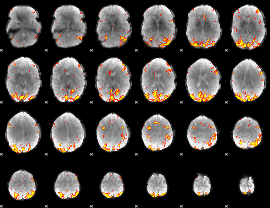 |
GLM result cope: 02, threshold: 3.5
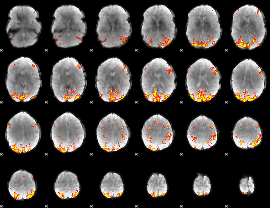 |
HAFNI result component: 019
Overlap rate: 0.46754
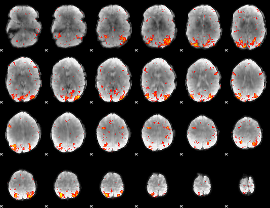 |
HAFNI result component: 228
Overlap rate: 0.4779
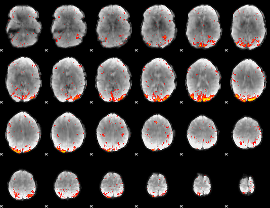 |
Correlation: 0.27844
  |
Correlation: 0.3442
  |
Subject: 47 |
GLM result cope: 01, threshold: 3.5
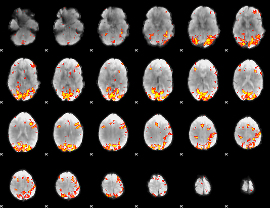 |
GLM result cope: 02, threshold: 3.5
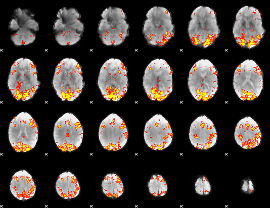 |
HAFNI result component: 072
Overlap rate: 0.4181
 |
HAFNI result component: 036
Overlap rate: 0.31469
 |
Correlation: 0.34976
  |
Correlation: 0.41803
  |
Subject: 48 |
GLM result cope: 01, threshold: 3.5
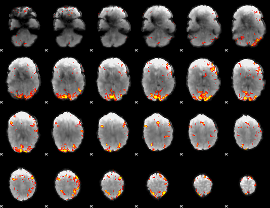 |
GLM result cope: 02, threshold: 3.5
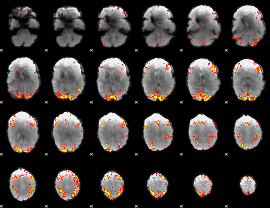 |
HAFNI result component: 390
Overlap rate: 0.5917
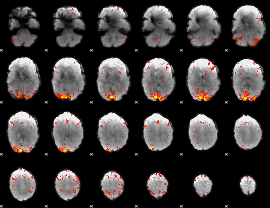 |
HAFNI result component: 351
Overlap rate: 0.36376
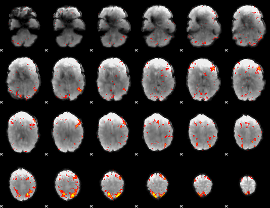 |
Correlation: 0.25003
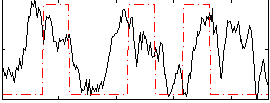  |
Correlation: 0.33573
  |
Subject: 49 |
GLM result cope: 01, threshold: 3.5
 |
GLM result cope: 02, threshold: 4
 |
HAFNI result component: 179
Overlap rate: 0.42671
 |
HAFNI result component: 160
Overlap rate: 0.26285
 |
Correlation: 0.3236
  |
Correlation: 0.34564
  |
Subject: 50 |
GLM result cope: 01, threshold: 4
 |
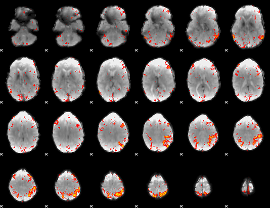
GLM result cope: 02, threshold: 4
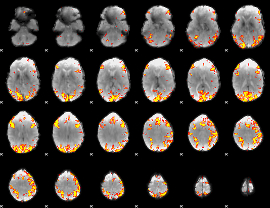 |
HAFNI result component: 167
Overlap rate: 0.33035
 |
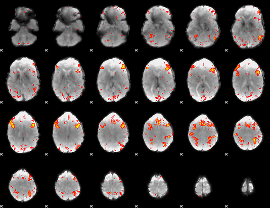
HAFNI result component: 122
Overlap rate: 0.355
 |
Correlation: 0.37406
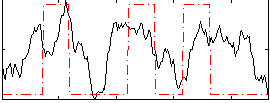  |
Correlation: 0.2572
  |
Subject: 51 |
GLM result cope: 01, threshold: 4
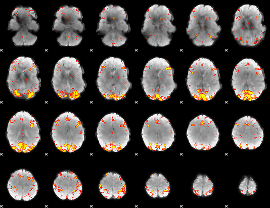 |
GLM result cope: 02, threshold: 4
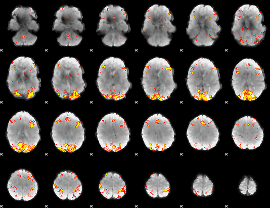 |
HAFNI result component: 114
Overlap rate: 0.41037
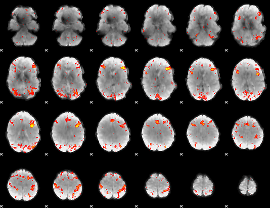 |
HAFNI result component: 373
Overlap rate: 0.27004
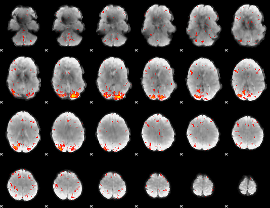 |
Correlation: 0.32299
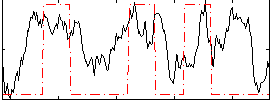  |
Correlation: 0.25191
  |
Subject: 52 |
GLM result cope: 01, threshold: 4
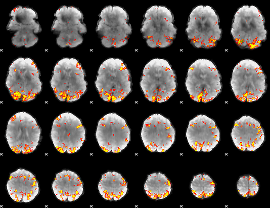 |
GLM result cope: 02, threshold: 4
 |
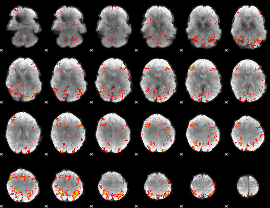
HAFNI result component: 250
Overlap rate: 0.45387
 |
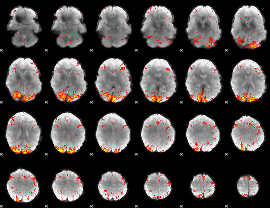
HAFNI result component: 245
Overlap rate: 0.39855
 |
Correlation: 0.3163
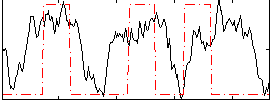  |
Correlation: 0.30309
  |
Subject: 53 |
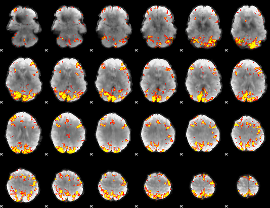
GLM result cope: 01, threshold: 4
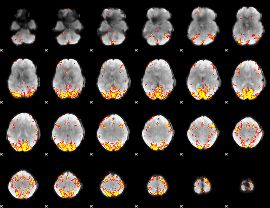 |
GLM result cope: 02, threshold: 4
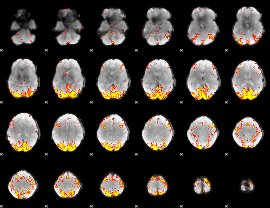 |
HAFNI result component: 384
Overlap rate: 0.30839
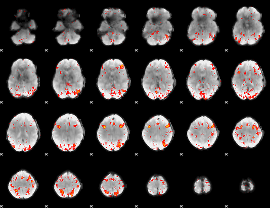 |
HAFNI result component: 207
Overlap rate: 0.32521
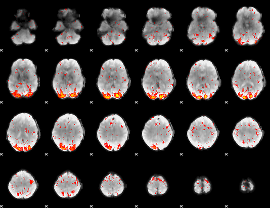 |
Correlation: 0.21442
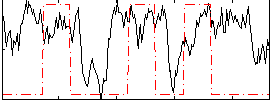  |
Correlation: 0.45426
  |
Subject: 54 |
GLM result cope: 01, threshold: 3.5
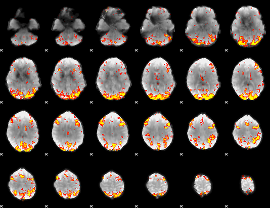 |
GLM result cope: 02, threshold: 4
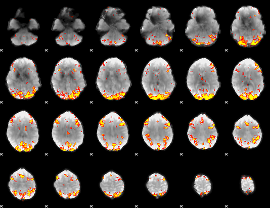 |
HAFNI result component: 308
Overlap rate: 0.37342
 |
HAFNI result component: 188
Overlap rate: 0.3382
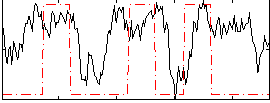
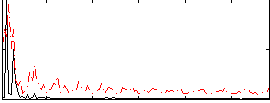
 |
Correlation: 0.26953
  |
Correlation: 0.20545
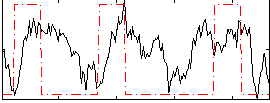  |
Subject: 55 |
GLM result cope: 01, threshold: 4
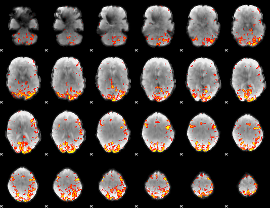 |
GLM result cope: 02, threshold: 4
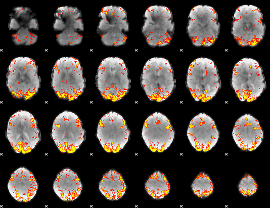 |
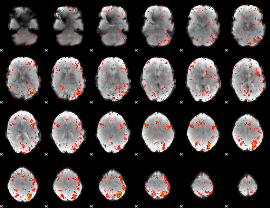
HAFNI result component: 271
Overlap rate: 0.28578
 |
HAFNI result component: 022
Overlap rate: 0.26875
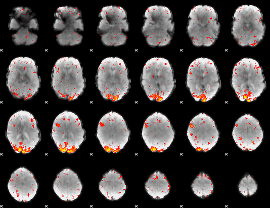 |
Correlation: 0.32932
  |
Correlation: 0.39171
  |
Subject: 56 |
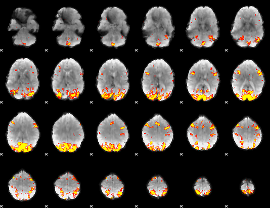
GLM result cope: 01, threshold: 3.5
 |
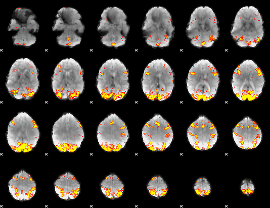
GLM result cope: 02, threshold: 4
 |
HAFNI result component: 387
Overlap rate: 0.55282
 |
HAFNI result component: 052
Overlap rate: 0.38122
 |
Correlation: 0.25716
  |
Correlation: 0.11615
  |
Subject: 57 |
GLM result cope: 01, threshold: 4
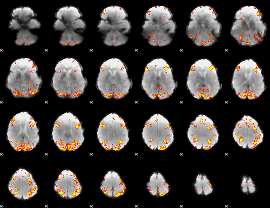 |
GLM result cope: 02, threshold: 4
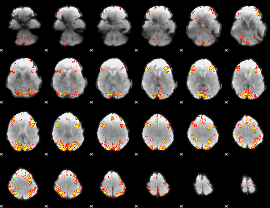 |
HAFNI result component: 114
Overlap rate: 0.43063
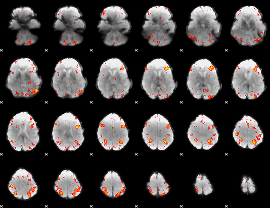 |
HAFNI result component: 186
Overlap rate: 0.36653
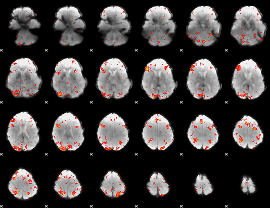 |
Correlation: 0.394
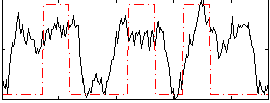  |
Correlation: 0.20949
  |
Subject: 58 |
GLM result cope: 01, threshold: 3.5
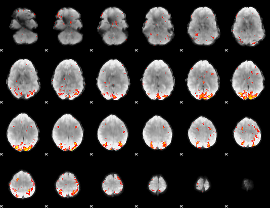 |
GLM result cope: 02, threshold: 4
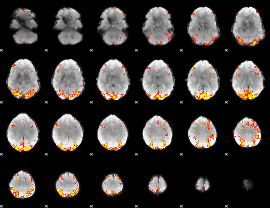 |
HAFNI result component: 203
Overlap rate: 0.4958
 |
HAFNI result component: 043
Overlap rate: 0.37635
 |
Correlation: 0.10881
  |
Correlation: 0.4519
  |
Subject: 59 |
GLM result cope: 01, threshold: 4
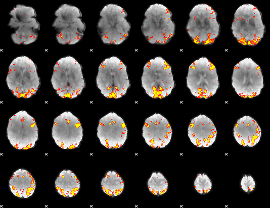 |
GLM result cope: 02, threshold: 4
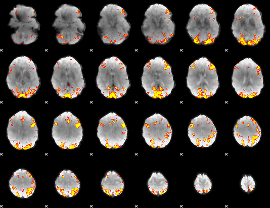 |
HAFNI result component: 304
Overlap rate: 0.43988
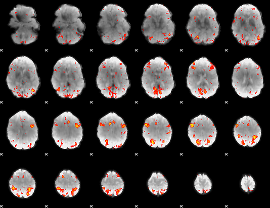 |
HAFNI result component: 375
Overlap rate: 0.49307
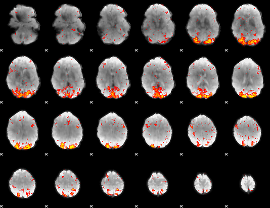 |
Correlation: 0.3426
  |
Correlation: 0.26898
  |
Subject: 60 |
GLM result cope: 01, threshold: 4
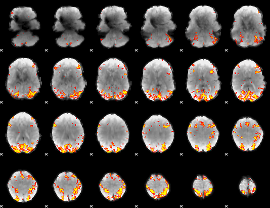 |
GLM result cope: 02, threshold: 4
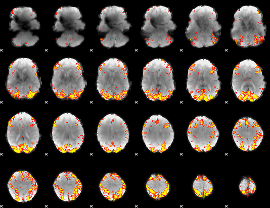 |
HAFNI result component: 050
Overlap rate: 0.43941
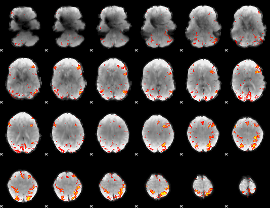 |
HAFNI result component: 364
Overlap rate: 0.27154
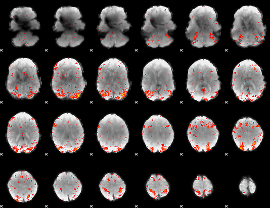 |
Correlation: 0.1343
  |
Correlation: 0.27306
  |
Subject: 61 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 62 |
GLM result cope: 01, threshold: 3.5
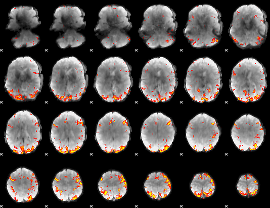 |
GLM result cope: 02, threshold: 3.5
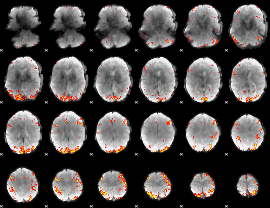 |
HAFNI result component: 213
Overlap rate: 0.39984
 |
HAFNI result component: 255
Overlap rate: 0.50381
 |
Correlation: 0.28131
  |
Correlation: 0.19433
  |
Subject: 63 |
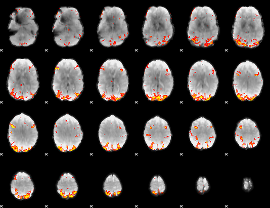
GLM result cope: 01, threshold: 3.5
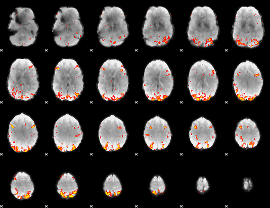 |
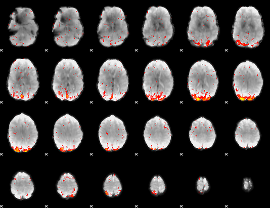
GLM result cope: 02, threshold: 3.5
 |
HAFNI result component: 167
Overlap rate: 0.3927
 |
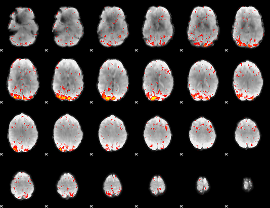
HAFNI result component: 157
Overlap rate: 0.36656
 |
Correlation: 0.2788
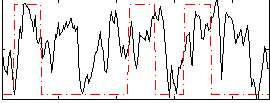  |
Correlation: 0.30492
  |
Subject: 64 |
GLM result cope: 01, threshold: 3.5
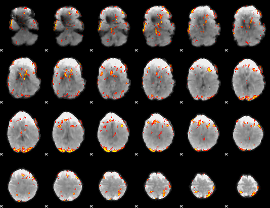 |
GLM result cope: 02, threshold: 4
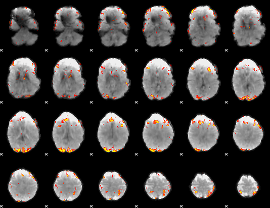 |
HAFNI result component: 363
Overlap rate: 0.26104
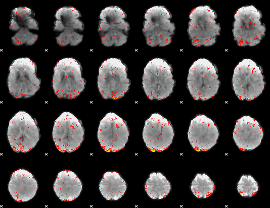 |
HAFNI result component: 226
Overlap rate: 0.26291
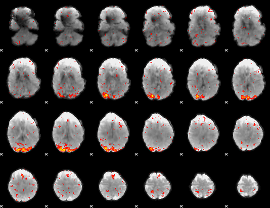 |
Correlation: 0.18711
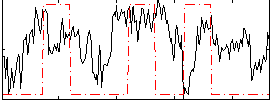  |
Correlation: 0.37281
  |
Subject: 65 |
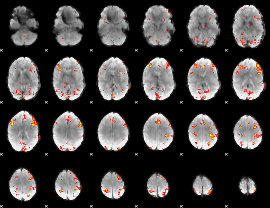
GLM result cope: 01, threshold: 3.5
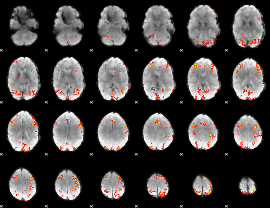 |
GLM result cope: 02, threshold: 3.5
 |
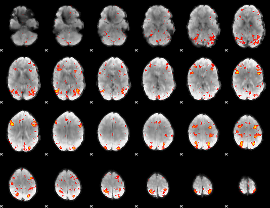
HAFNI result component: 114
Overlap rate: 0.56309
 |
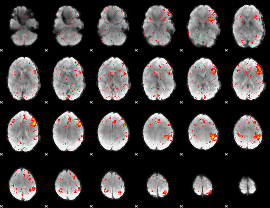
HAFNI result component: 018
Overlap rate: 0.44359
 |
Correlation: 0.28647
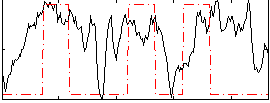  |
Correlation: 0.15763
  |
Subject: 66 |
GLM result cope: 01, threshold: 4
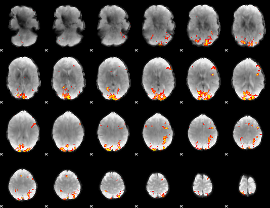 |
GLM result cope: 02, threshold: 4
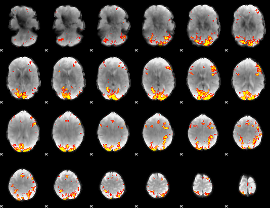 |
HAFNI result component: 388
Overlap rate: 0.50322
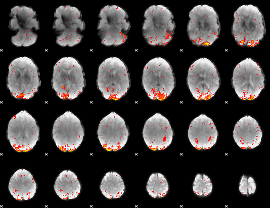 |
HAFNI result component: 029
Overlap rate: 0.45532
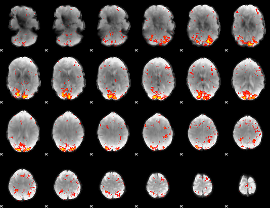 |
Correlation: 0.27904
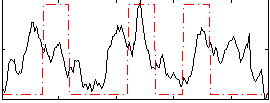  |
Correlation: 0.41662
  |
Subject: 67 |
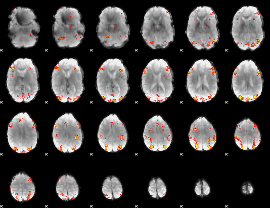
GLM result cope: 01, threshold: 3.5
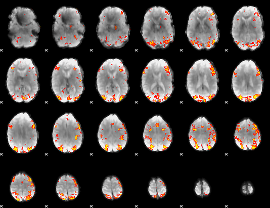 |
GLM result cope: 02, threshold: 3.5
 |
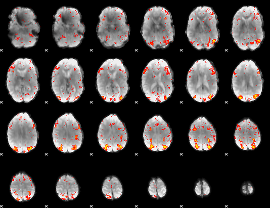
HAFNI result component: 279
Overlap rate: 0.57869
 |
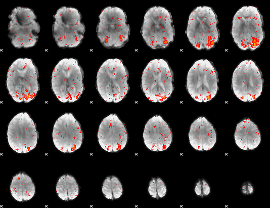
HAFNI result component: 326
Overlap rate: 0.22576
 |
Correlation: 0.34828
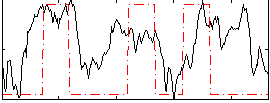  |
Correlation: 0.22798
  |
Subject: 68 |
GLM result cope: 01, threshold: 4
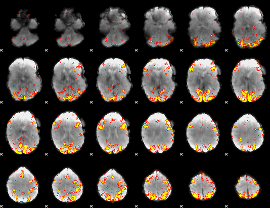 |
GLM result cope: 02, threshold: 4
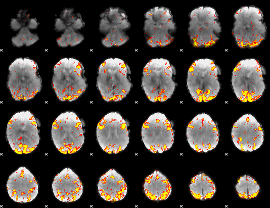 |
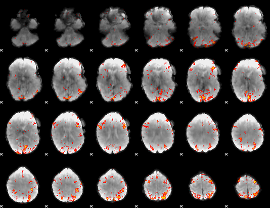
HAFNI result component: 186
Overlap rate: 0.36737
 |
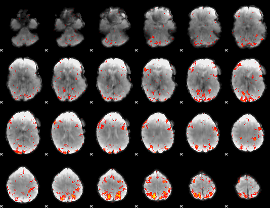
HAFNI result component: 379
Overlap rate: 0.29963
 |
Correlation: 0.13369
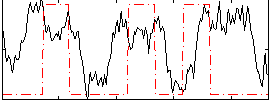  |
Correlation: 0.34692
  |































































































































































































































































































































































































































































































































































