HAFNI report on finalized components on task: LANGUAGE |
Totally 2 copes finalized |
Subject: 1 |
GLM result cope: 02, threshold: 2.5
 |
GLM result cope: 03, threshold: 3
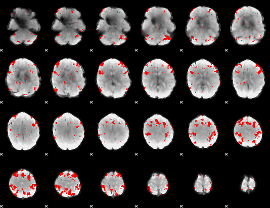 |
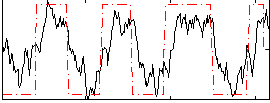
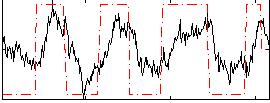
HAFNI result component: 220
Overlap rate: 0.90545
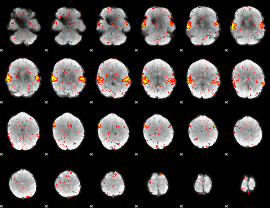 |
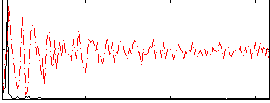
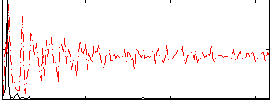
HAFNI result component: 311
Overlap rate: 0.5267
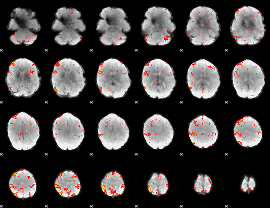 |
Correlation: 0.40686
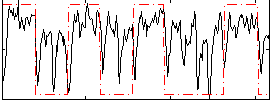  |
Correlation: 0.49951
  |
Subject: 2 |
GLM result cope: 02, threshold: 2.5
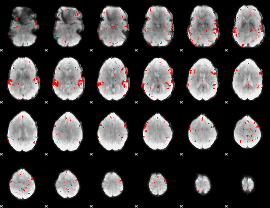 |
GLM result cope: 03, threshold: 4
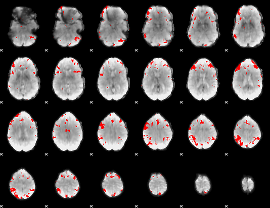 |
HAFNI result component: 009
Overlap rate: 0.75624
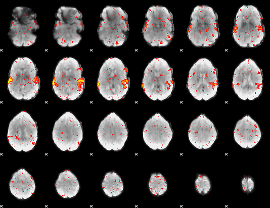 |
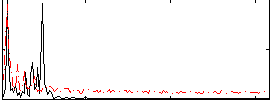
HAFNI result component: 316
Overlap rate: 0.55358
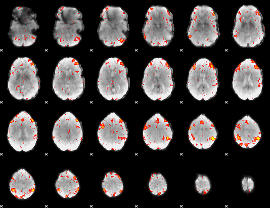 |
Correlation: 0.3096
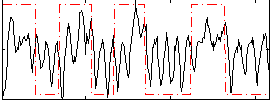  |
Correlation: 0.54896
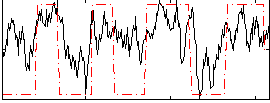 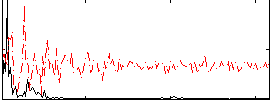 |
Subject: 3 |
GLM result cope: 02, threshold: 1.5
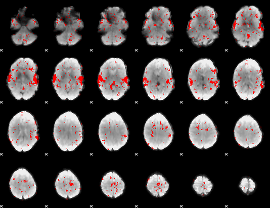 |
GLM result cope: 03, threshold: 4
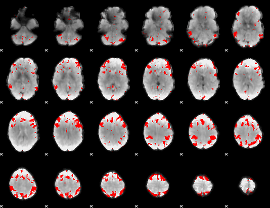 |
HAFNI result component: 277
Overlap rate: 0.95008
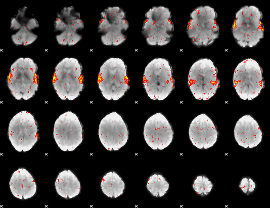 |
HAFNI result component: 150
Overlap rate: 0.36918
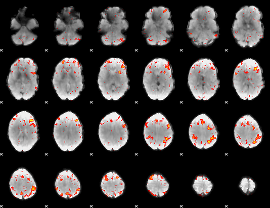 |
Correlation: 0.27207
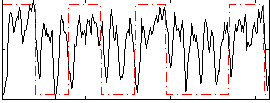  |
Correlation: 0.63766
  |
Subject: 4 |
GLM result cope: 02, threshold: 3.5
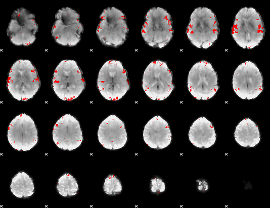 |
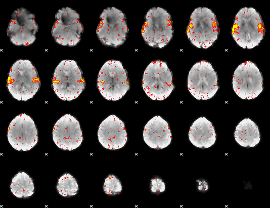
GLM result cope: 03, threshold: 4
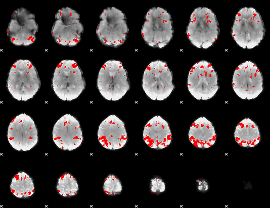 |
HAFNI result component: 124
Overlap rate: 0.63331
 |
HAFNI result component: 029
Overlap rate: 0.38072
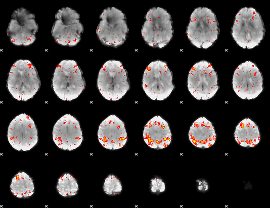 |
Correlation: 0.41849
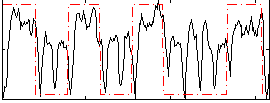  |
Correlation: 0.57597
  |
Subject: 5 |
GLM result cope: 02, threshold: 2
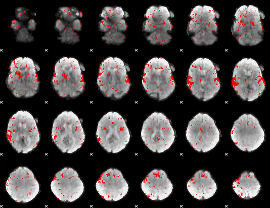 |
GLM result cope: 03, threshold: 3
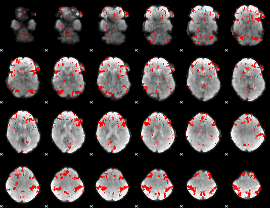 |
HAFNI result component: 172
Overlap rate: 0.83482
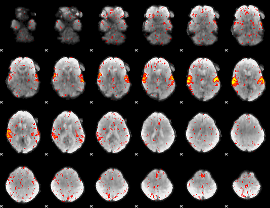 |
HAFNI result component: 298
Overlap rate: 0.32237
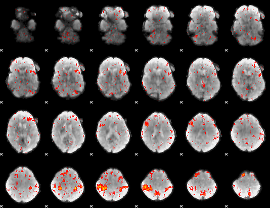 |
Correlation: 0.28129
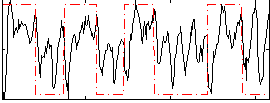  |
Correlation: 0.40449
  |
Subject: 6 |
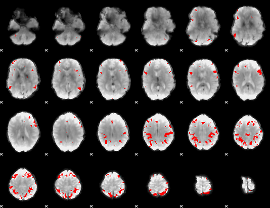
GLM result cope: 02, threshold: 4
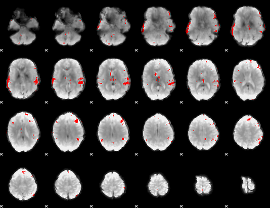 |
GLM result cope: 03, threshold: 4
 |
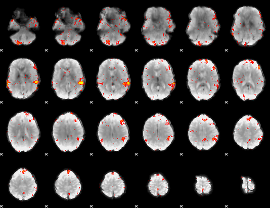
HAFNI result component: 003
Overlap rate: 0.49775
 |
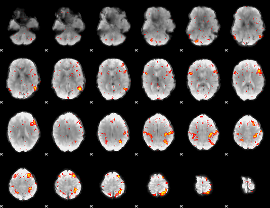
HAFNI result component: 358
Overlap rate: 0.5168
 |
Correlation: 0.34414
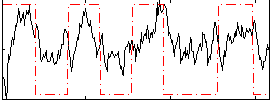  |
Correlation: 0.60641
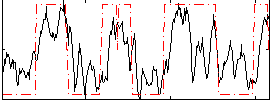  |
Subject: 7 |
GLM result cope: 02, threshold: 1.5
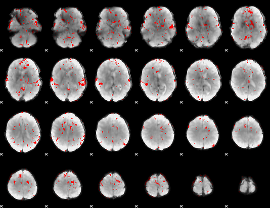 |
GLM result cope: 03, threshold: 4
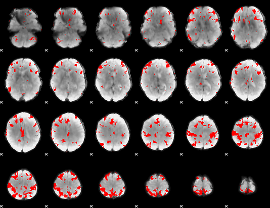 |
HAFNI result component: 187
Overlap rate: 0.7
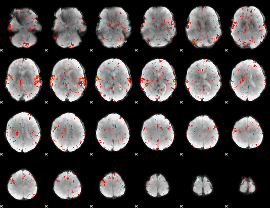 |
HAFNI result component: 241
Overlap rate: 0.40563
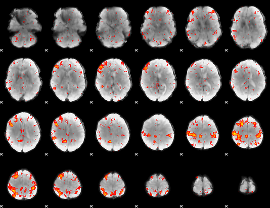 |
Correlation: 0.21232
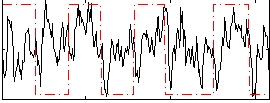 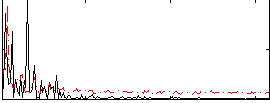 |
Correlation: 0.46692
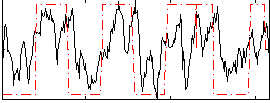 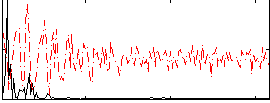 |
Subject: 8 |
GLM result cope: 02, threshold: 4
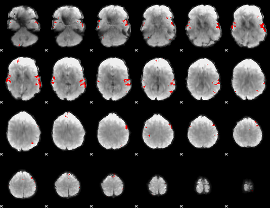 |
GLM result cope: 03, threshold: 4
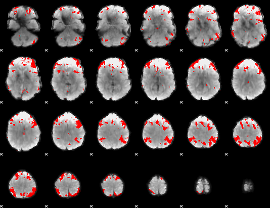 |
HAFNI result component: 374
Overlap rate: 0.76526
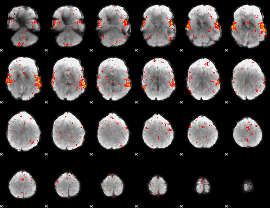 |
HAFNI result component: 306
Overlap rate: 0.40185
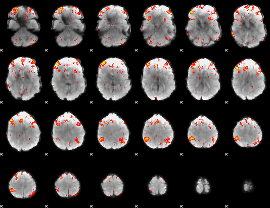 |
Correlation: 0.2592
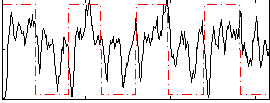  |
Correlation: 0.58021
  |
Subject: 9 |
GLM result cope: 02, threshold: 2
 |
GLM result cope: 03, threshold: 4
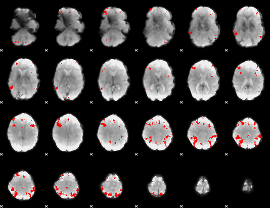 |
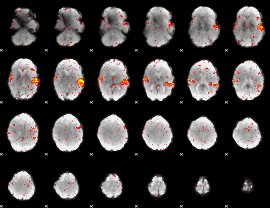
HAFNI result component: 291
Overlap rate: 0.688
 |
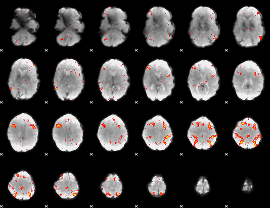
HAFNI result component: 065
Overlap rate: 0.53796
 |
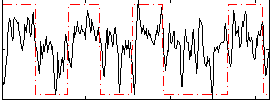
Correlation: 0.41679
 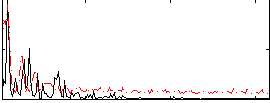 |
Correlation: 0.55955
  |
Subject: 10 |
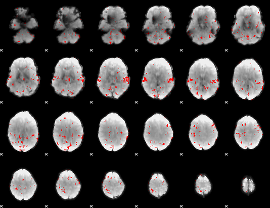
GLM result cope: 02, threshold: 2
 |
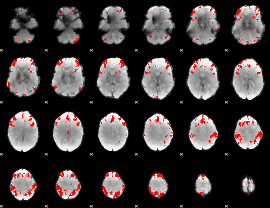
GLM result cope: 03, threshold: 4
 |
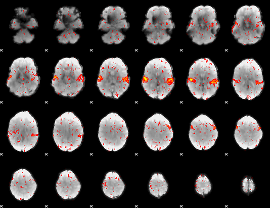
HAFNI result component: 233
Overlap rate: 0.84375
 |
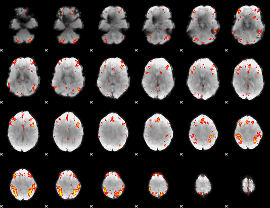
HAFNI result component: 289
Overlap rate: 0.52267
 |
Correlation: 0.21956
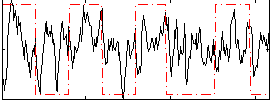 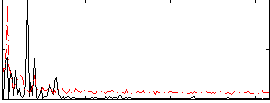 |
Correlation: 0.64697
  |
Subject: 11 |
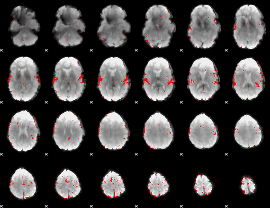
GLM result cope: 02, threshold: 3
 |
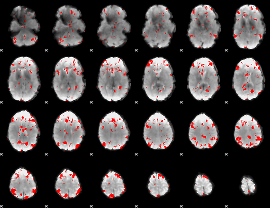
GLM result cope: 03, threshold: 2
 |
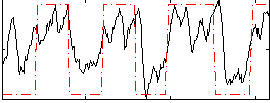
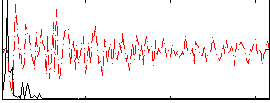
HAFNI result component: 347
Overlap rate: 0.47532
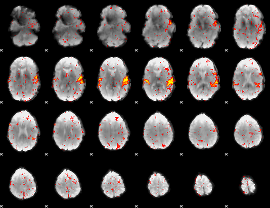 |
HAFNI result component: 213
Overlap rate: 0.58439
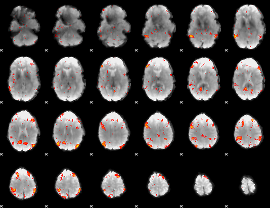 |
Correlation: 0.32871
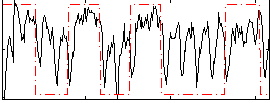 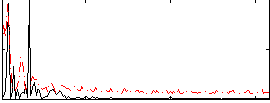 |
Correlation: 0.41139
  |
Subject: 12 |
GLM result cope: 02, threshold: 2
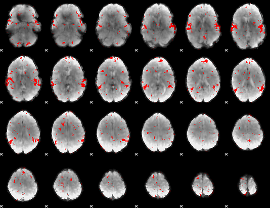 |
GLM result cope: 03, threshold: 4
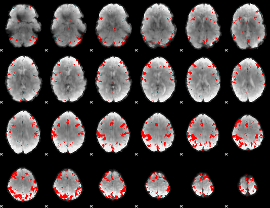 |
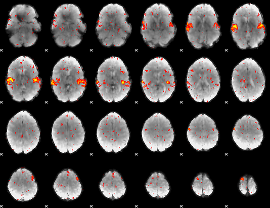
HAFNI result component: 080
Overlap rate: 0.53003
 |
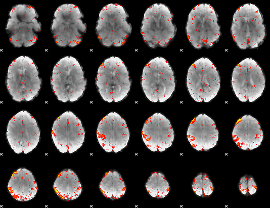
HAFNI result component: 304
Overlap rate: 0.45125
 |
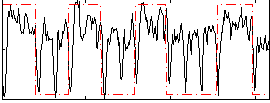
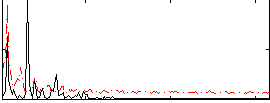
Correlation: 0.38479
  |
Correlation: 0.63081
  |
Subject: 13 |
GLM result cope: 02, threshold: 1.5
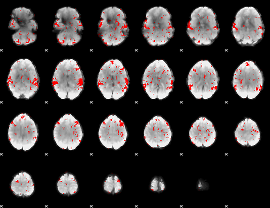 |
GLM result cope: 03, threshold: 4
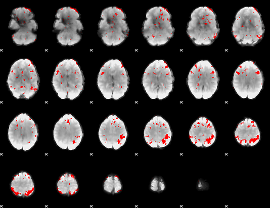 |
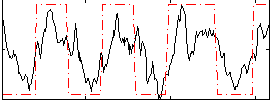
HAFNI result component: 069
Overlap rate: 0.83203
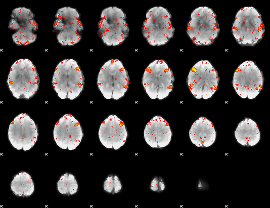 |
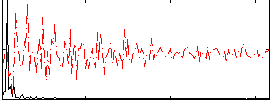
HAFNI result component: 048
Overlap rate: 0.44228
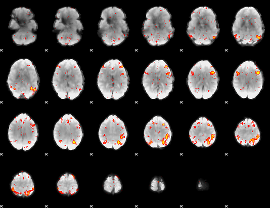 |
Correlation: 0.55311
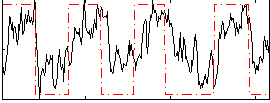 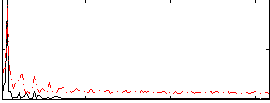 |
Correlation: 0.62897
  |
Subject: 14 |
GLM result cope: 02, threshold: 1
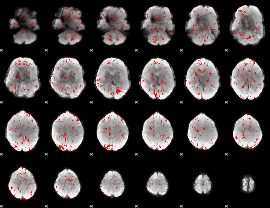 |
GLM result cope: 03, threshold: 4
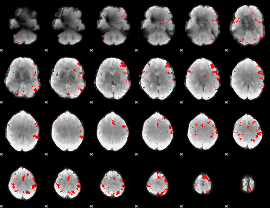 |
HAFNI result component: 040
Overlap rate: 1
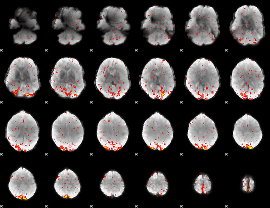 |
HAFNI result component: 051
Overlap rate: 0.55085
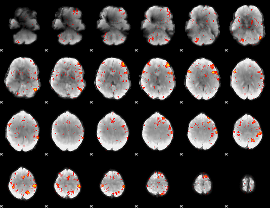 |
Correlation: 0.02231
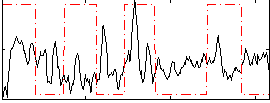 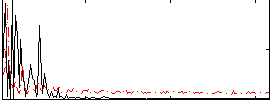 |
Correlation: 0.65238
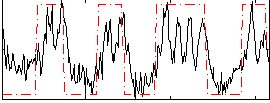  |
Subject: 15 |
GLM result cope: 02, threshold: 3
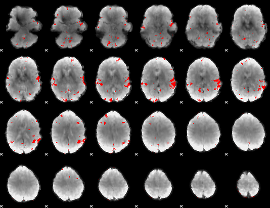 |
GLM result cope: 03, threshold: 4
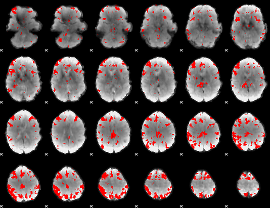 |
HAFNI result component: 233
Overlap rate: 0.62615
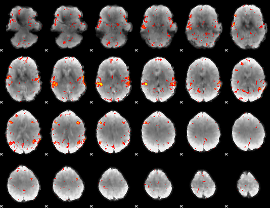 |
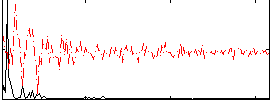
HAFNI result component: 085
Overlap rate: 0.45598
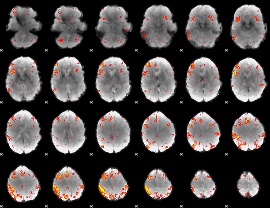 |
Correlation: 0.61341
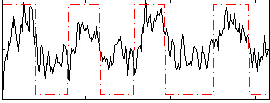  |
Correlation: 0.71572
  |
Subject: 16 |
GLM result cope: 02, threshold: 2
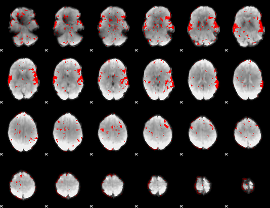 |
GLM result cope: 03, threshold: 4
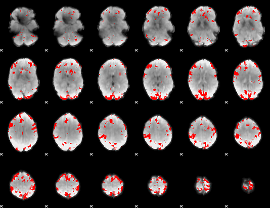 |
HAFNI result component: 360
Overlap rate: 0.55138
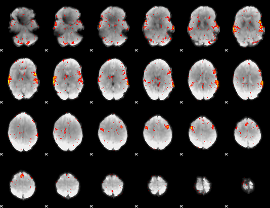 |
HAFNI result component: 193
Overlap rate: 0.30207
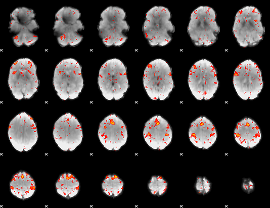 |
Correlation: -0.057451
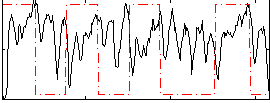 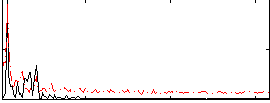 |
Correlation: 0.63287
  |
Subject: 17 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 03, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 18 |
GLM result cope: 02, threshold: 2
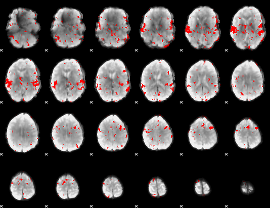 |
GLM result cope: 03, threshold: 4
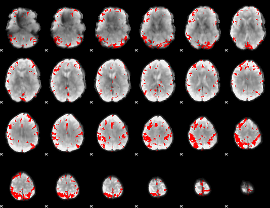 |
HAFNI result component: 161
Overlap rate: 0.87638
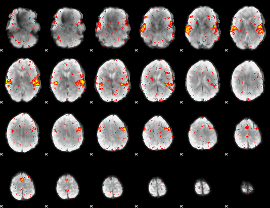 |
HAFNI result component: 238
Overlap rate: 0.3646
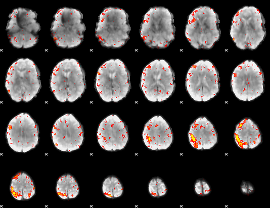 |
Correlation: 0.040891
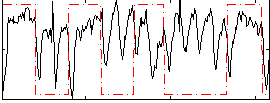 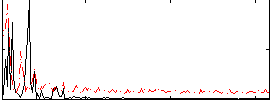 |
Correlation: 0.67723
  |
Subject: 19 |
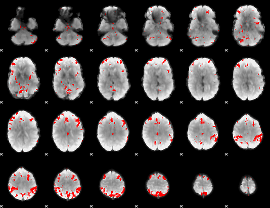
GLM result cope: 02, threshold: 2.5
 |
GLM result cope: 03, threshold: 4
 |
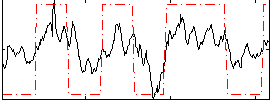
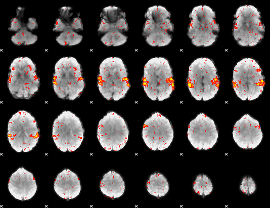
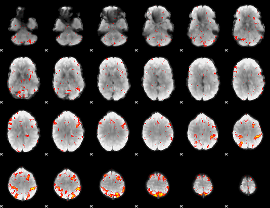
HAFNI result component: 033
Overlap rate: 0.75639
 |
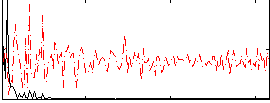
HAFNI result component: 319
Overlap rate: 0.3905
 |
Correlation: 0.30846
  |
Correlation: 0.56373
  |
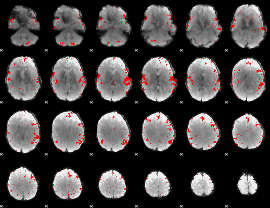
Subject: 20 |
GLM result cope: 02, threshold: 4
 |
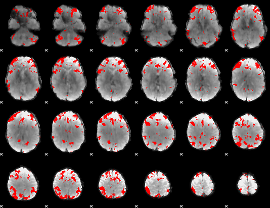
GLM result cope: 03, threshold: 4
 |
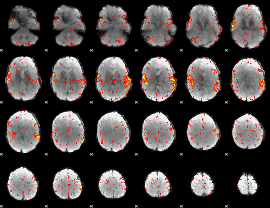
HAFNI result component: 368
Overlap rate: 0.54293
 |
HAFNI result component: 190
Overlap rate: 0.44293
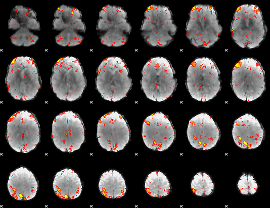 |
Correlation: 0.19446
  |
Correlation: 0.67115
  |
Subject: 21 |
GLM result cope: 02, threshold: 2
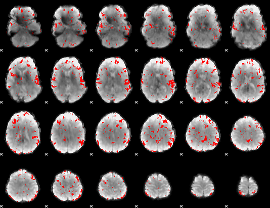 |
GLM result cope: 03, threshold: 2.5
 |
HAFNI result component: 066
Overlap rate: 0.61
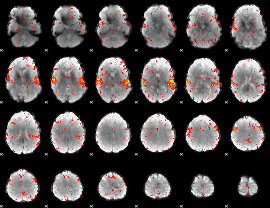 |
HAFNI result component: 288
Overlap rate: 0.18605
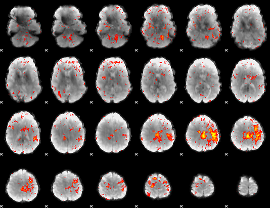 |
Correlation: 0.059614
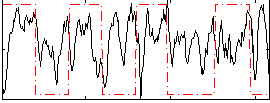 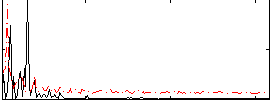 |
Correlation: 0.32786
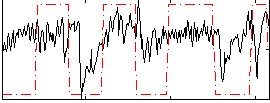 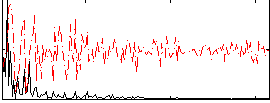 |
Subject: 22 |
GLM result cope: 02, threshold: 2
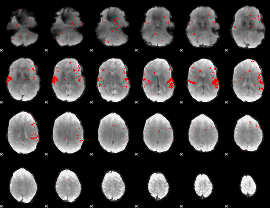 |
GLM result cope: 03, threshold: 2.5
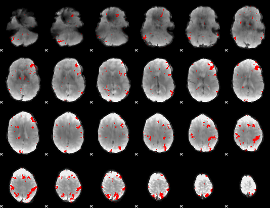 |
HAFNI result component: 212
Overlap rate: 0.9402
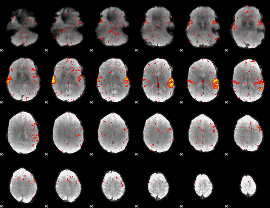 |
HAFNI result component: 382
Overlap rate: 0.66756
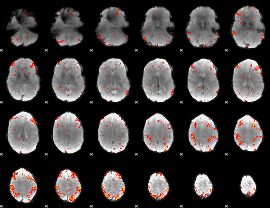 |
Correlation: 0.31172
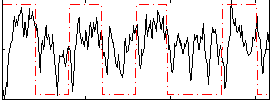  |
Correlation: 0.43609
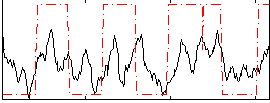 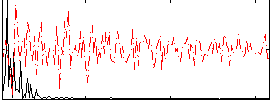 |
Subject: 23 |
GLM result cope: 02, threshold: 2
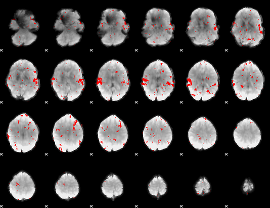 |
GLM result cope: 03, threshold: 4
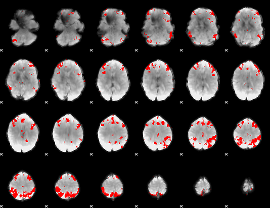 |
HAFNI result component: 341
Overlap rate: 0.84926
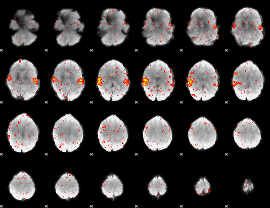 |
HAFNI result component: 339
Overlap rate: 0.45445
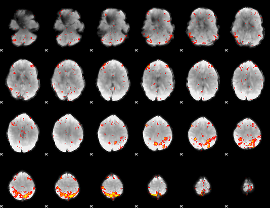 |
Correlation: 0.37464
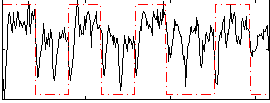  |
Correlation: 0.46092
  |
Subject: 24 |
GLM result cope: 02, threshold: 2.5
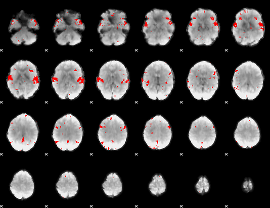 |
GLM result cope: 03, threshold: 3.5
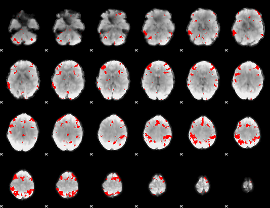 |
HAFNI result component: 312
Overlap rate: 0.86026
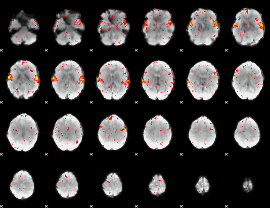 |
HAFNI result component: 220
Overlap rate: 0.51168
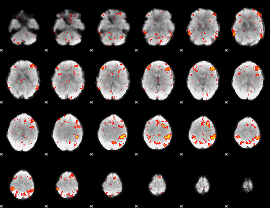 |
Correlation: 0.38897
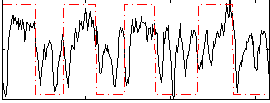  |
Correlation: 0.53309
  |
Subject: 25 |
GLM result cope: 02, threshold: 2
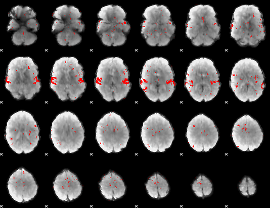 |
GLM result cope: 03, threshold: 4
 |
HAFNI result component: 149
Overlap rate: 0.97763
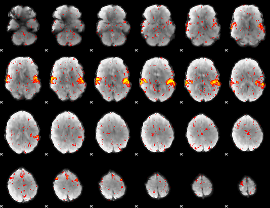 |
HAFNI result component: 050
Overlap rate: 0.38206
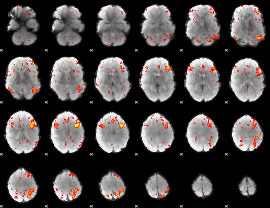 |
Correlation: 0.22589
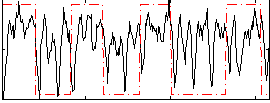 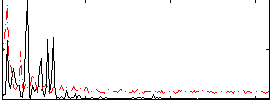 |
Correlation: 0.61443
  |
Subject: 26 |
GLM result cope: 02, threshold: 2
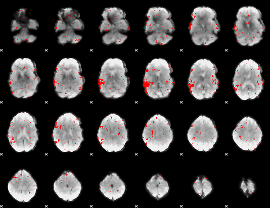 |
GLM result cope: 03, threshold: 2
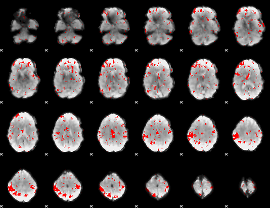 |
HAFNI result component: 001
Overlap rate: 0.88532
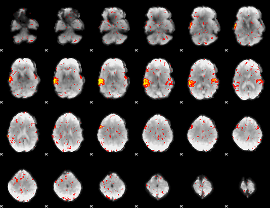 |
HAFNI result component: 297
Overlap rate: 0.64121
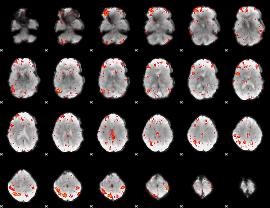 |
Correlation: 0.33463
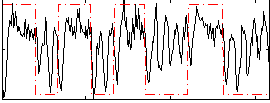 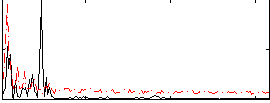 |
Correlation: 0.53491
  |
Subject: 27 |
GLM result cope: 02, threshold: 2
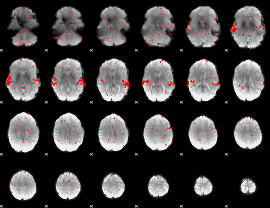 |
GLM result cope: 03, threshold: 3.5
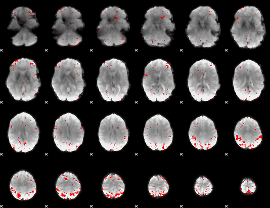 |
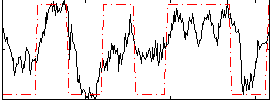
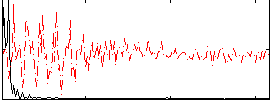
HAFNI result component: 246
Overlap rate: 0.93286
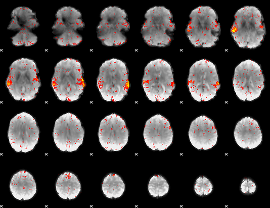 |
HAFNI result component: 296
Overlap rate: 0.4878
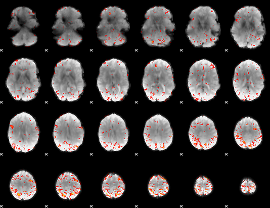 |
Correlation: 0.081554
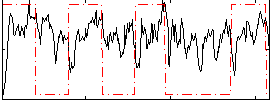 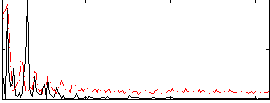 |
Correlation: 0.63099
  |
Subject: 28 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 03, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 29 |
GLM result cope: 02, threshold: 3
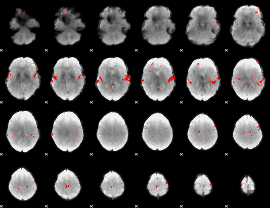 |
GLM result cope: 03, threshold: 4
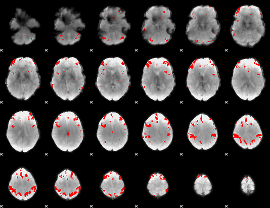 |
HAFNI result component: 248
Overlap rate: 0.67497
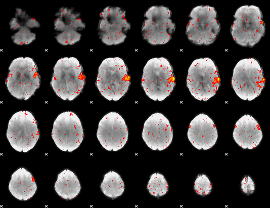 |
HAFNI result component: 331
Overlap rate: 0.61703
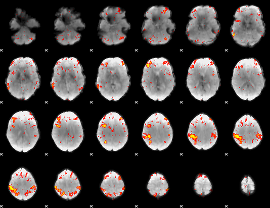 |
Correlation: 0.27668
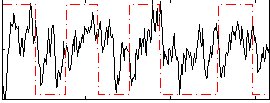 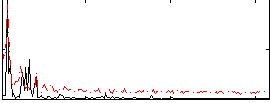 |
Correlation: 0.52428
  |
Subject: 30 |
GLM result cope: 02, threshold: 2.5
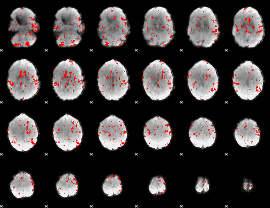 |
GLM result cope: 03, threshold: 3
 |
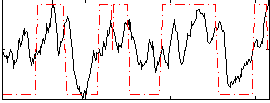
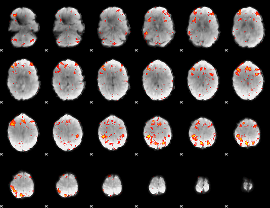
HAFNI result component: 380
Overlap rate: 0.45578
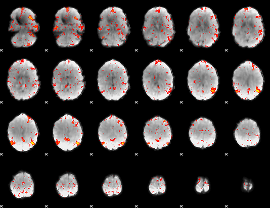 |
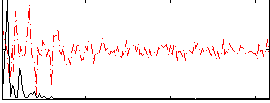
HAFNI result component: 391
Overlap rate: 0.68521
 |
Correlation: 0.50153
  |
Correlation: 0.47254
  |
Subject: 31 |
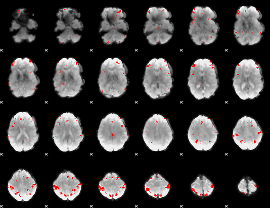
GLM result cope: 02, threshold: 2
 |
GLM result cope: 03, threshold: 4
 |
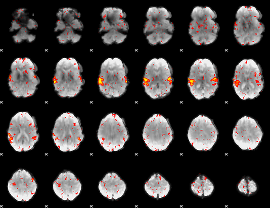
HAFNI result component: 353
Overlap rate: 0.8483
 |
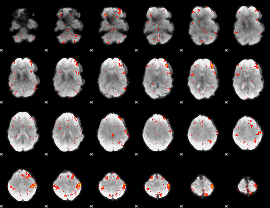
HAFNI result component: 037
Overlap rate: 0.50479
 |
Correlation: 0.3525
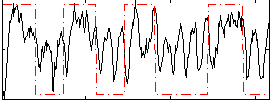 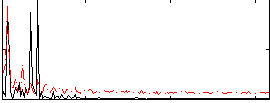 |
Correlation: 0.53142
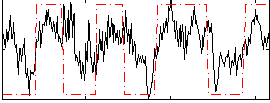 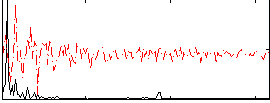 |
Subject: 32 |
GLM result cope: 02, threshold: 3.5
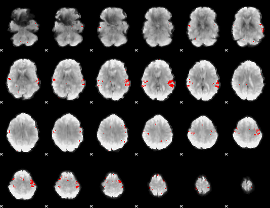 |
GLM result cope: 03, threshold: 4
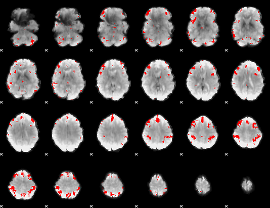 |
HAFNI result component: 199
Overlap rate: 0.50632
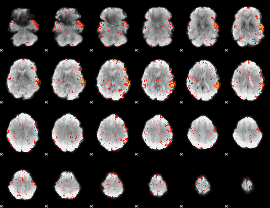 |
HAFNI result component: 180
Overlap rate: 0.64406
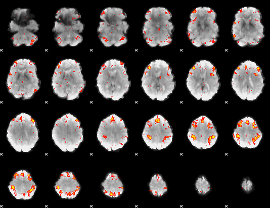 |
Correlation: 0.52575
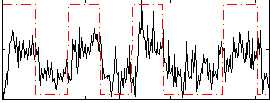 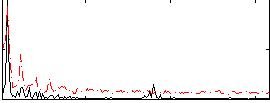 |
Correlation: 0.55655
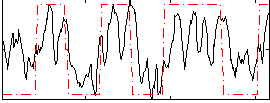 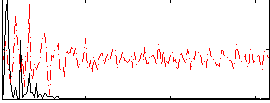 |
Subject: 33 |
GLM result cope: 02, threshold: 1.5
 |
GLM result cope: 03, threshold: 3.5
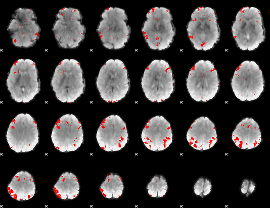 |
HAFNI result component: 062
Overlap rate: 0.81481
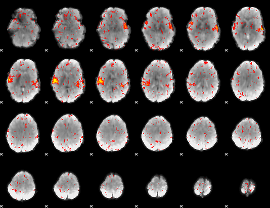 |
HAFNI result component: 202
Overlap rate: 0.55027
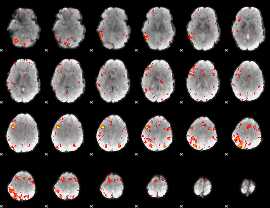 |
Correlation: 0.32087
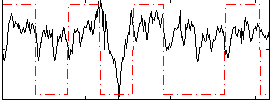  |
Correlation: 0.54555
  |
Subject: 34 |
GLM result cope: 02, threshold: 3.5
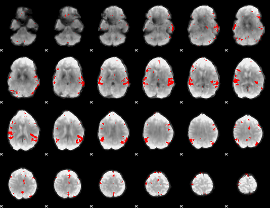 |
GLM result cope: 03, threshold: 4
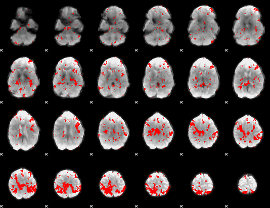 |
HAFNI result component: 137
Overlap rate: 0.48099
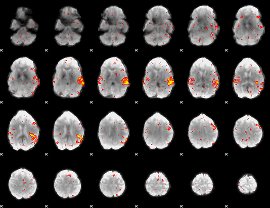 |
HAFNI result component: 329
Overlap rate: 0.17336
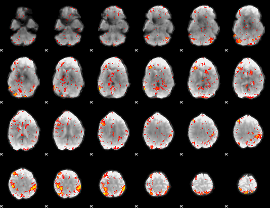 |
Correlation: 0.30138
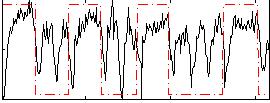  |
Correlation: 0.52739
  |
Subject: 35 |
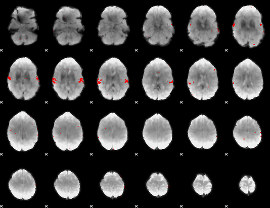
GLM result cope: 02, threshold: 3
 |
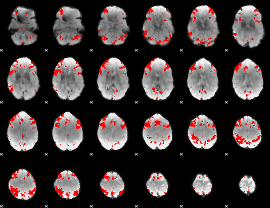
GLM result cope: 03, threshold: 3.5
 |
HAFNI result component: 324
Overlap rate: 0.95957
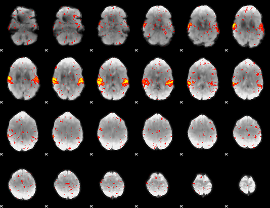 |
HAFNI result component: 319
Overlap rate: 0.52726
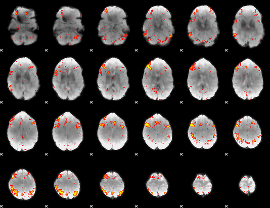 |
Correlation: 0.35846
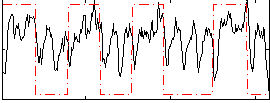 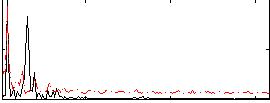 |
Correlation: 0.70346
  |
Subject: 36 |
GLM result cope: 02, threshold: 3
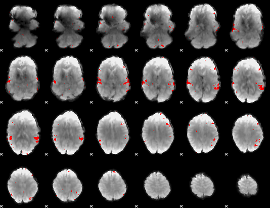 |
GLM result cope: 03, threshold: 3.5
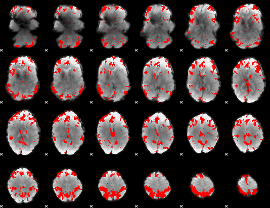 |
HAFNI result component: 312
Overlap rate: 0.69828
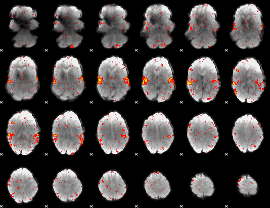 |
HAFNI result component: 071
Overlap rate: 0.33591
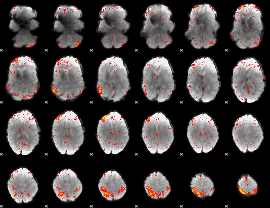 |
Correlation: 0.3689
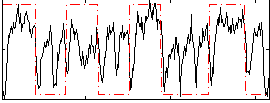 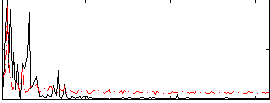 |
Correlation: 0.52703
  |
Subject: 37 |
GLM result cope: 02, threshold: 3
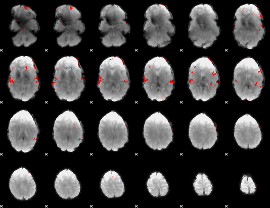 |
GLM result cope: 03, threshold: 3
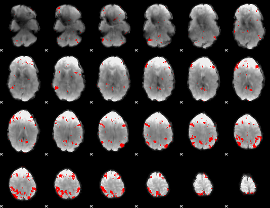 |
HAFNI result component: 228
Overlap rate: 0.73544
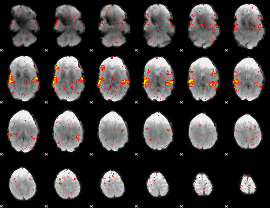 |
HAFNI result component: 391
Overlap rate: 0.48305
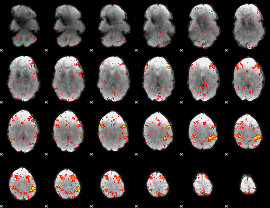 |
Correlation: 0.29528
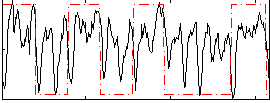 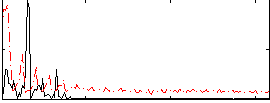 |
Correlation: 0.49322
  |
Subject: 38 |
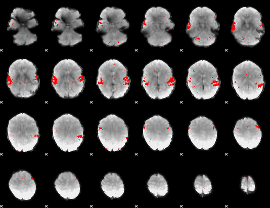
GLM result cope: 02, threshold: 3
 |
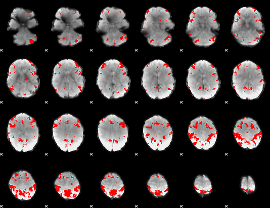
GLM result cope: 03, threshold: 3.5
 |
HAFNI result component: 034
Overlap rate: 0.93475
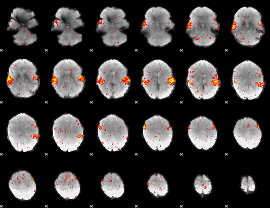 |
HAFNI result component: 021
Overlap rate: 0.49426
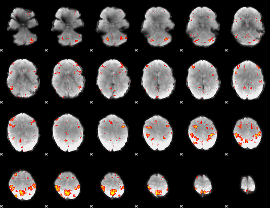 |
Correlation: 0.29012
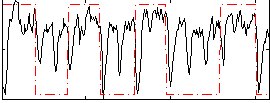 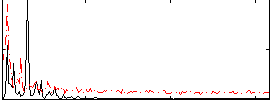 |
Correlation: 0.6634
  |
Subject: 39 |
GLM result cope: 02, threshold: 3
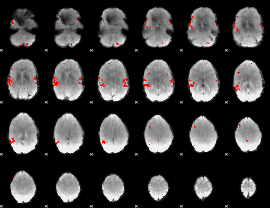 |
GLM result cope: 03, threshold: 3.5
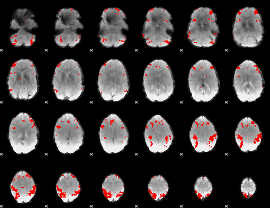 |
HAFNI result component: 088
Overlap rate: 0.53425
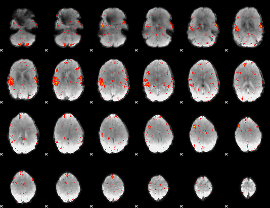 |
HAFNI result component: 022
Overlap rate: 0.50638
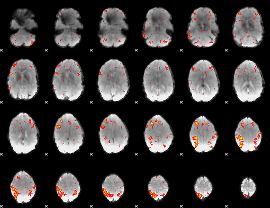 |
Correlation: 0.51062
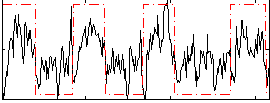 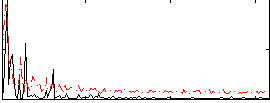 |
Correlation: 0.63603
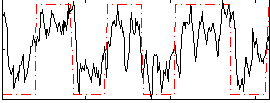 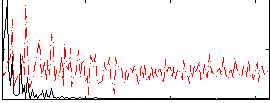 |
Subject: 40 |
GLM result cope: 02, threshold: 2.5
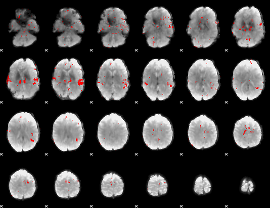 |
GLM result cope: 03, threshold: 3.5
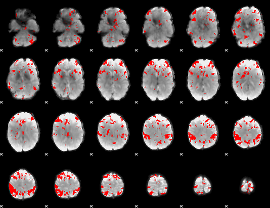 |
HAFNI result component: 353
Overlap rate: 0.49029
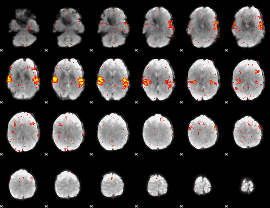 |
HAFNI result component: 363
Overlap rate: 0.46108
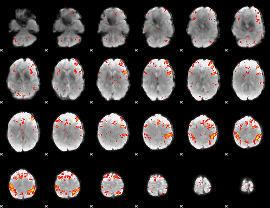 |
Correlation: 0.33227
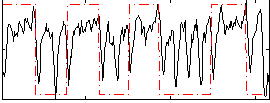 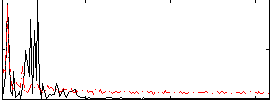 |
Correlation: 0.62503
  |
Subject: 41 |
GLM result cope: 02, threshold: 2
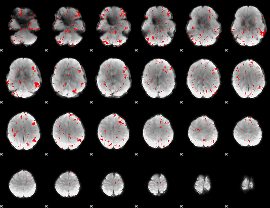 |
GLM result cope: 03, threshold: 3.5
 |
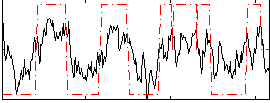
HAFNI result component: 362
Overlap rate: 0.57692
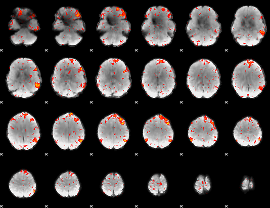 |
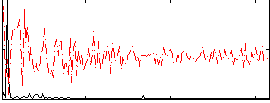
HAFNI result component: 360
Overlap rate: 0.52723
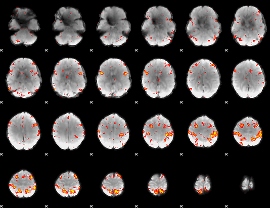 |
Correlation: 0.32097
  |
Correlation: 0.72746
  |
Subject: 42 |
GLM result cope: 02, threshold: 2.5
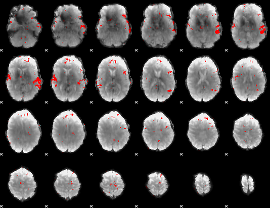 |
GLM result cope: 03, threshold: 3.5
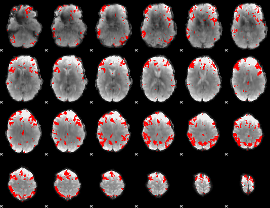 |
HAFNI result component: 112
Overlap rate: 0.63853
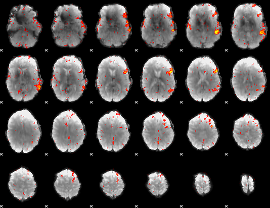 |
HAFNI result component: 338
Overlap rate: 0.46542
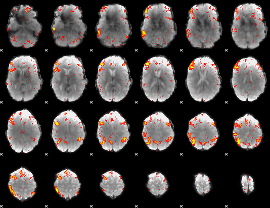 |
Correlation: 0.37638
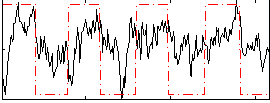  |
Correlation: 0.52281
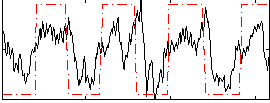  |
Subject: 43 |
GLM result cope: 02, threshold: 2.5
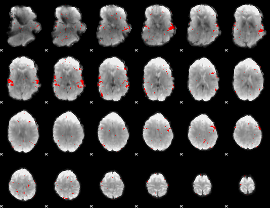 |
GLM result cope: 03, threshold: 3.5
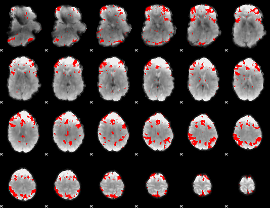 |
HAFNI result component: 055
Overlap rate: 0.73084
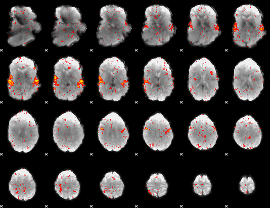 |
HAFNI result component: 228
Overlap rate: 0.24877
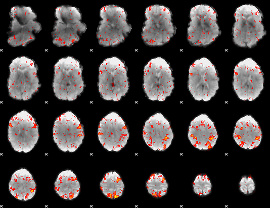 |
Correlation: 0.37414
  |
Correlation: 0.50487
  |
Subject: 44 |
GLM result cope: 02, threshold: 3
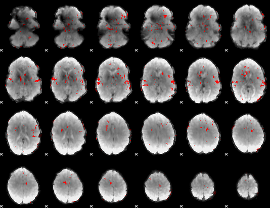 |
GLM result cope: 03, threshold: 3.5
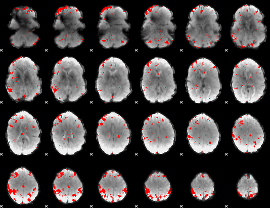 |
HAFNI result component: 343
Overlap rate: 0.31269
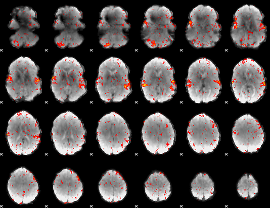 |
HAFNI result component: 227
Overlap rate: 0.41057
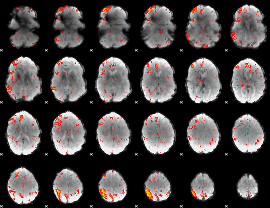 |
Correlation: 0.24547
  |
Correlation: 0.52233
  |
Subject: 45 |
GLM result cope: 02, threshold: 3.5
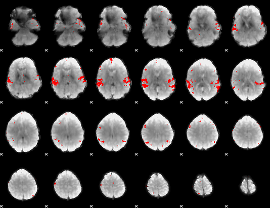 |
GLM result cope: 03, threshold: 3.5
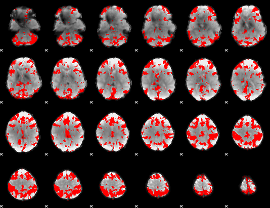 |
HAFNI result component: 156
Overlap rate: 0.60984
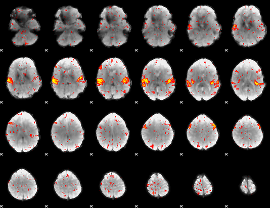 |
HAFNI result component: 140
Overlap rate: 0.22231
 |
Correlation: 0.15637
  |
Correlation: 0.62151
  |
Subject: 46 |
GLM result cope: 02, threshold: 1.5
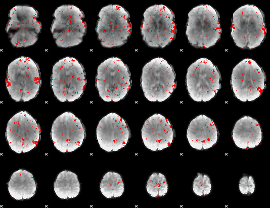 |
GLM result cope: 03, threshold: 3
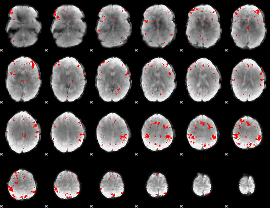 |
HAFNI result component: 269
Overlap rate: 0.58915
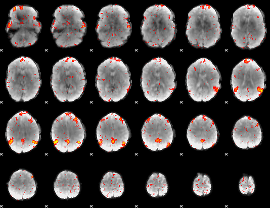 |
HAFNI result component: 292
Overlap rate: 0.62926
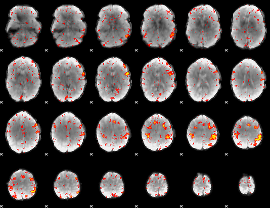 |
Correlation: 0.24252
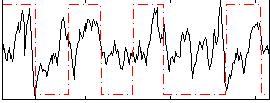 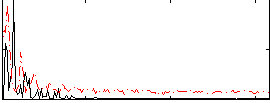 |
Correlation: 0.66033
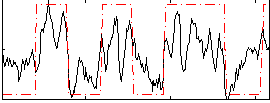  |
Subject: 47 |
GLM result cope: 02, threshold: 2.5
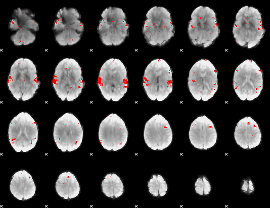 |
GLM result cope: 03, threshold: 3
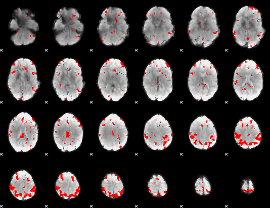 |
HAFNI result component: 258
Overlap rate: 0.74123
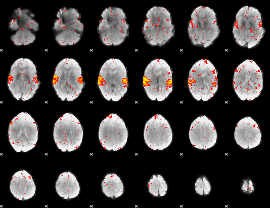 |
HAFNI result component: 267
Overlap rate: 0.50765
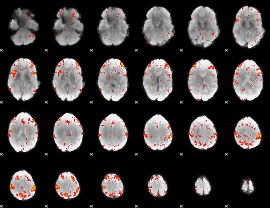 |
Correlation: 0.33615
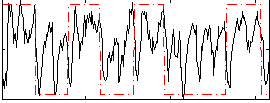 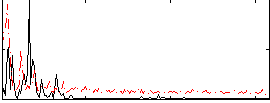 |
Correlation: 0.60363
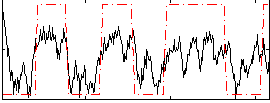  |
Subject: 48 |
GLM result cope: 02, threshold: 2
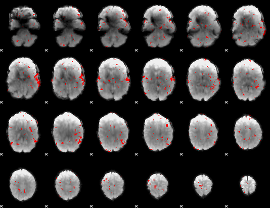 |
GLM result cope: 03, threshold: 3
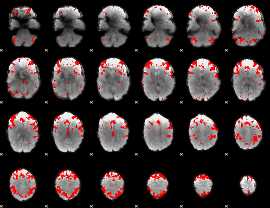 |
HAFNI result component: 397
Overlap rate: 0.5102
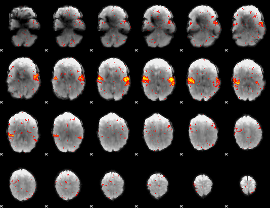 |
HAFNI result component: 383
Overlap rate: 0.40797
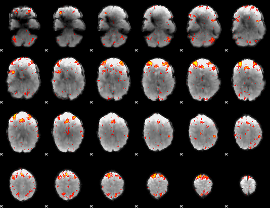 |
Correlation: 0.25216
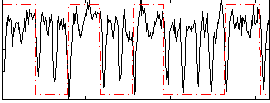 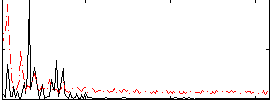 |
Correlation: 0.591
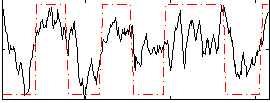  |
Subject: 49 |
GLM result cope: 02, threshold: 2.5
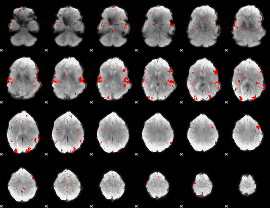 |
GLM result cope: 03, threshold: 3.5
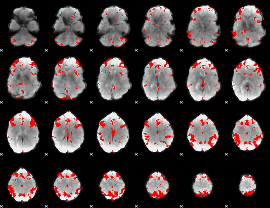 |
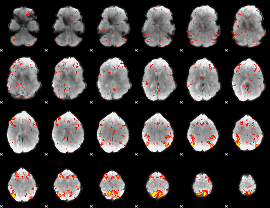
HAFNI result component: 204
Overlap rate: 0.58835
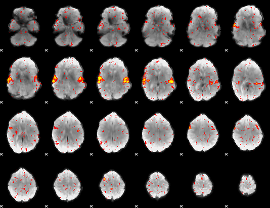 |
HAFNI result component: 160
Overlap rate: 0.37718
 |
Correlation: 0.33903
  |
Correlation: 0.48897
  |
Subject: 50 |
GLM result cope: 02, threshold: 3
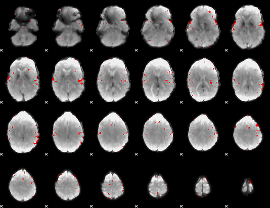 |
GLM result cope: 03, threshold: 3.5
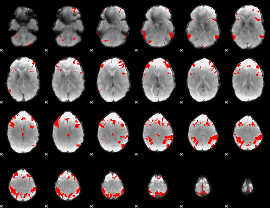 |
HAFNI result component: 182
Overlap rate: 0.63124
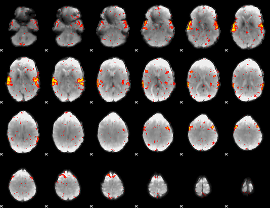 |
HAFNI result component: 224
Overlap rate: 0.52099
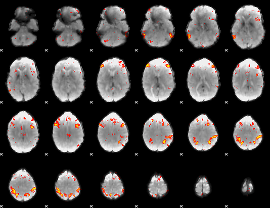 |
Correlation: 0.36123
  |
Correlation: 0.70842
  |
Subject: 51 |
GLM result cope: 02, threshold: 2.5
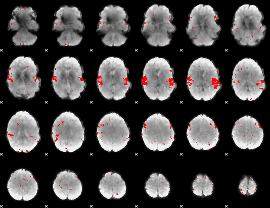 |
GLM result cope: 03, threshold: 3.5
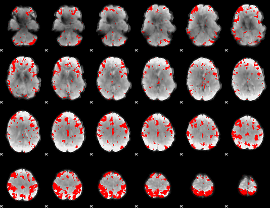 |
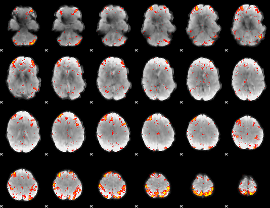
HAFNI result component: 118
Overlap rate: 0.70131
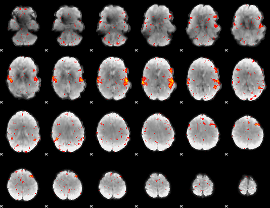 |
HAFNI result component: 261
Overlap rate: 0.44077
 |
Correlation: 0.38313
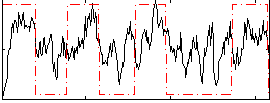 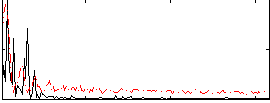 |
Correlation: 0.49206
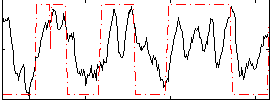  |
Subject: 52 |
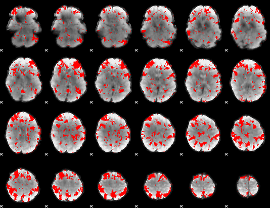
GLM result cope: 02, threshold: 2
 |
GLM result cope: 03, threshold: 2
 |
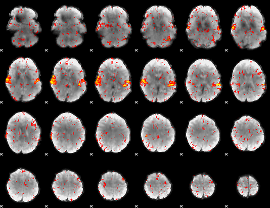
HAFNI result component: 353
Overlap rate: 0.86471
 |
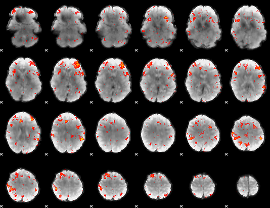
HAFNI result component: 311
Overlap rate: 0.4718
 |
Correlation: 0.35105
  |
Correlation: 0.62515
  |
Subject: 53 |
GLM result cope: 02, threshold: 2
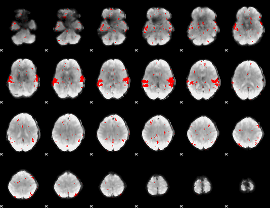 |
GLM result cope: 03, threshold: 2.5
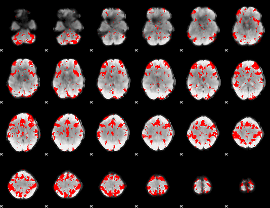 |
HAFNI result component: 386
Overlap rate: 0.93354
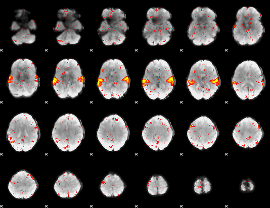 |
HAFNI result component: 307
Overlap rate: 0.34652
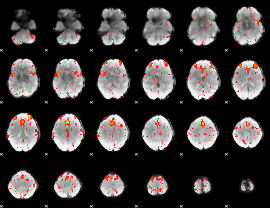 |
Correlation: 0.2556
  |
Correlation: 0.50741
  |
Subject: 54 |
GLM result cope: 02, threshold: 2
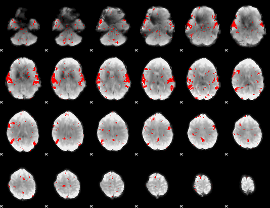 |
GLM result cope: 03, threshold: 3
 |
HAFNI result component: 024
Overlap rate: 0.86142
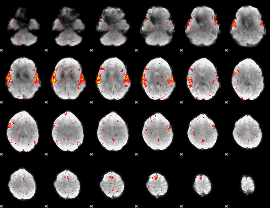 |
HAFNI result component: 209
Overlap rate: 0.53279
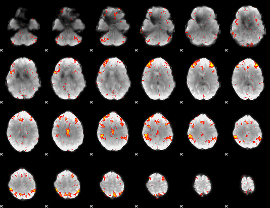 |
Correlation: 0.12622
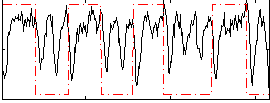 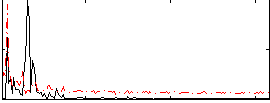 |
Correlation: 0.57735
  |
Subject: 55 |
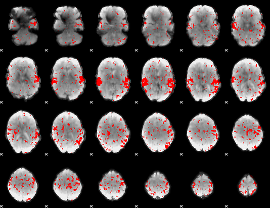
GLM result cope: 02, threshold: 2
 |
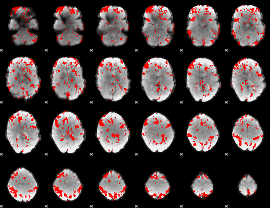
GLM result cope: 03, threshold: 2.5
 |
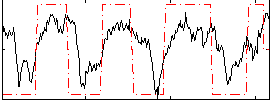
HAFNI result component: 233
Overlap rate: 0.91163
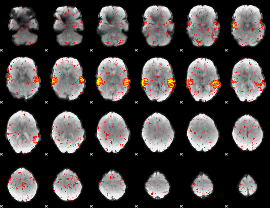 |
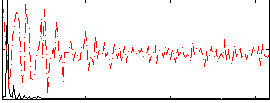
HAFNI result component: 384
Overlap rate: 0.50256
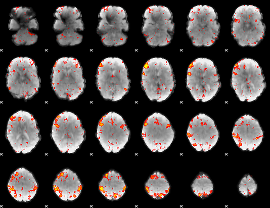 |
Correlation: 0.26914
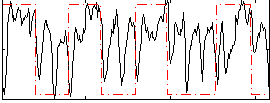 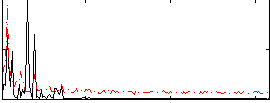 |
Correlation: 0.58671
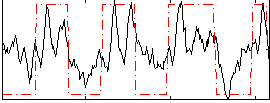 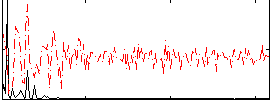 |
Subject: 56 |
GLM result cope: 02, threshold: 2
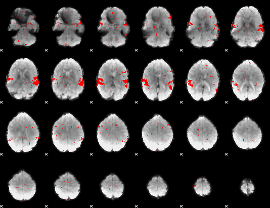 |
GLM result cope: 03, threshold: 2.5
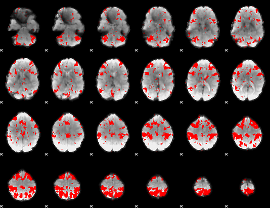 |
HAFNI result component: 330
Overlap rate: 0.88493
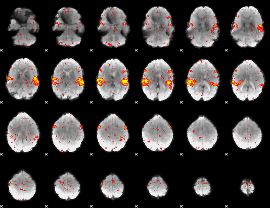 |
HAFNI result component: 187
Overlap rate: 0.45091
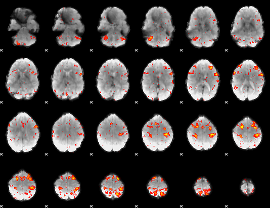 |
Correlation: 0.39158
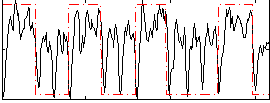 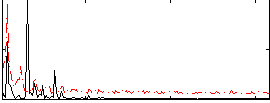 |
Correlation: 0.66615
  |
Subject: 57 |
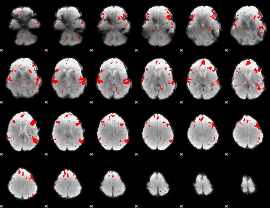
GLM result cope: 02, threshold: 2
 |
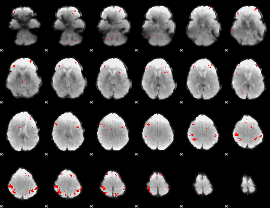
GLM result cope: 03, threshold: 3.5
 |
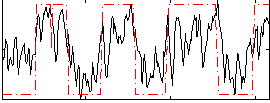
HAFNI result component: 205
Overlap rate: 0.52355
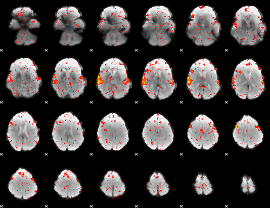 |
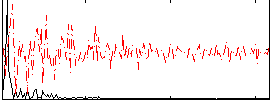
HAFNI result component: 124
Overlap rate: 0.77925
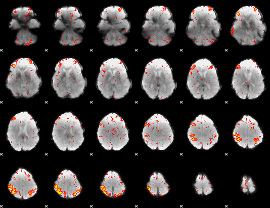 |
Correlation: 0.34304
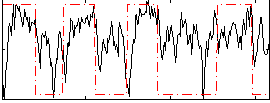 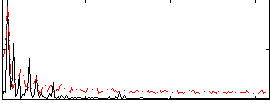 |
Correlation: 0.64402
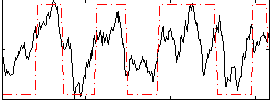 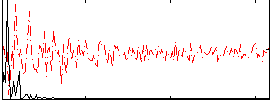 |
Subject: 58 |
GLM result cope: 02, threshold: 2.5
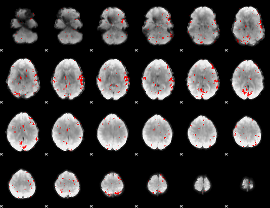 |
GLM result cope: 03, threshold: 3.5
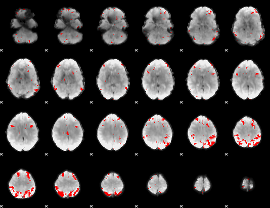 |
HAFNI result component: 269
Overlap rate: 0.59513
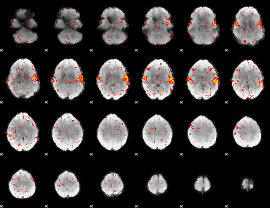 |
HAFNI result component: 153
Overlap rate: 0.5925
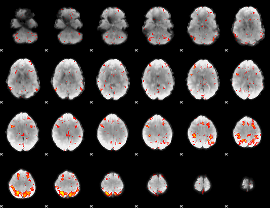 |
Correlation: 0.31638
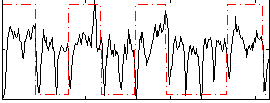 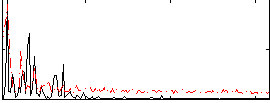 |
Correlation: 0.64471
  |
Subject: 59 |
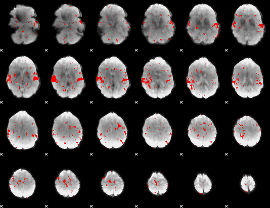
GLM result cope: 02, threshold: 2
 |
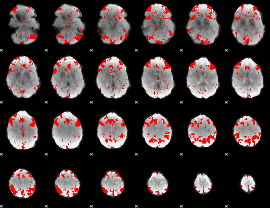
GLM result cope: 03, threshold: 2.5
 |
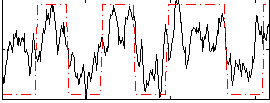
HAFNI result component: 392
Overlap rate: 0.94408
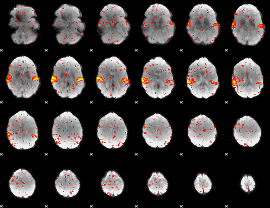 |
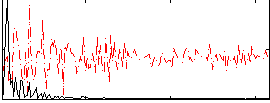
HAFNI result component: 210
Overlap rate: 0.60794
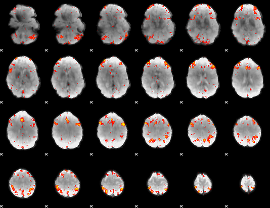 |
Correlation: 0.30899
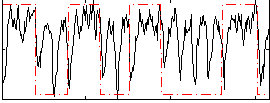 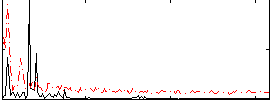 |
Correlation: 0.6649
  |
Subject: 60 |
GLM result cope: 02, threshold: 1.5
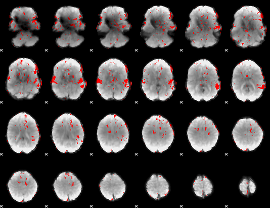 |
GLM result cope: 03, threshold: 3
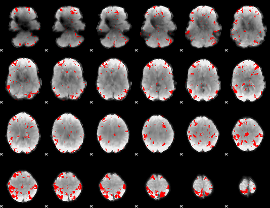 |
HAFNI result component: 307
Overlap rate: 0.33657
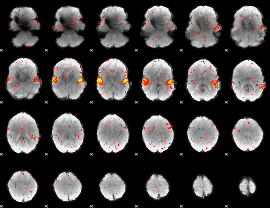 |
HAFNI result component: 313
Overlap rate: 0.41227
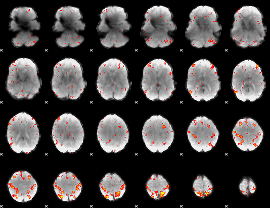 |
Correlation: 0.40671
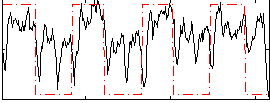 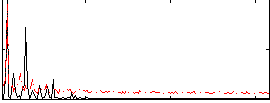 |
Correlation: 0.64651
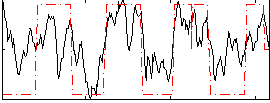 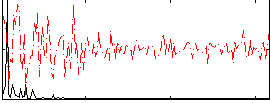 |
Subject: 61 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 03, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 62 |
GLM result cope: 02, threshold: 1.5
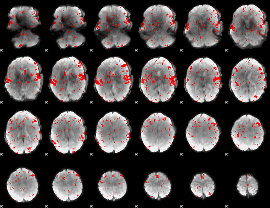 |
GLM result cope: 03, threshold: 3.5
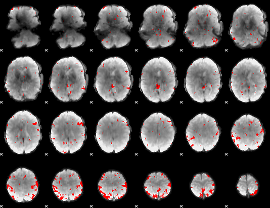 |
HAFNI result component: 223
Overlap rate: 0.88483
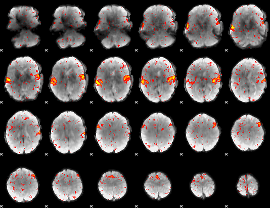 |
HAFNI result component: 287
Overlap rate: 0.6955
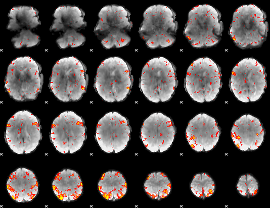 |
Correlation: 0.12738
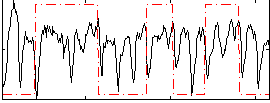 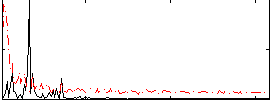 |
Correlation: 0.55471
  |
Subject: 63 |
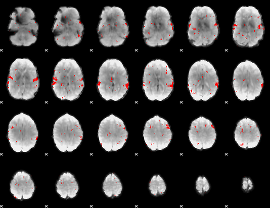
GLM result cope: 02, threshold: 2.5
 |
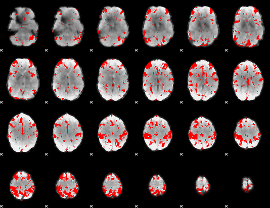
GLM result cope: 03, threshold: 3.5
 |
HAFNI result component: 212
Overlap rate: 0.74175
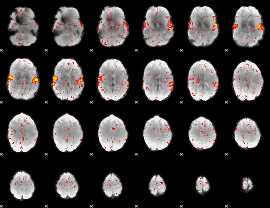 |
HAFNI result component: 353
Overlap rate: 0.37334
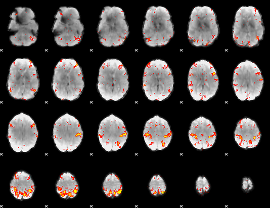 |
Correlation: 0.50691
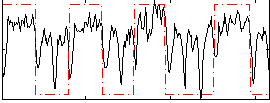 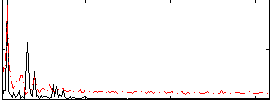 |
Correlation: 0.68603
  |
Subject: 64 |
GLM result cope: 02, threshold: 2.5
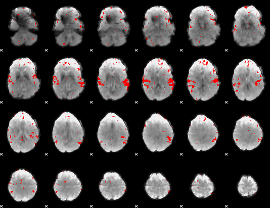 |
GLM result cope: 03, threshold: 2.5
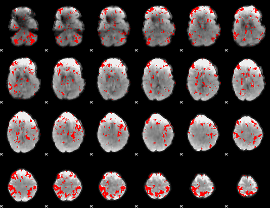 |
HAFNI result component: 037
Overlap rate: 0.83803
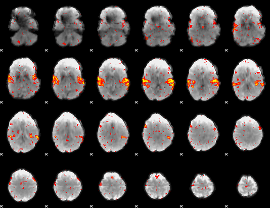 |
HAFNI result component: 236
Overlap rate: 0.48454
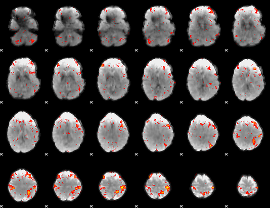 |
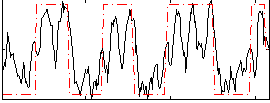
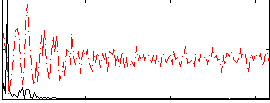
Correlation: 0.34236
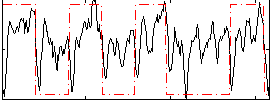 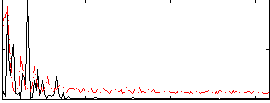 |
Correlation: 0.6188
  |
Subject: 65 |
GLM result cope: 02, threshold: 2.5
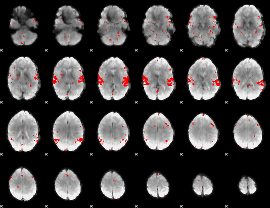 |
GLM result cope: 03, threshold: 3
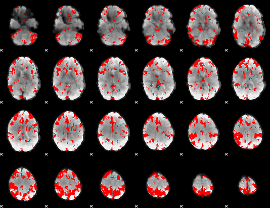 |
HAFNI result component: 362
Overlap rate: 0.5706
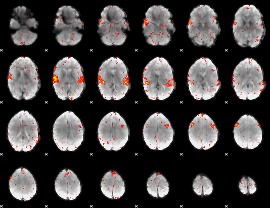 |
HAFNI result component: 162
Overlap rate: 0.38324
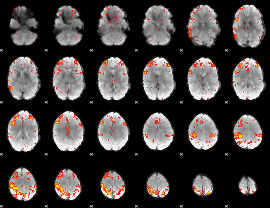 |
Correlation: 0.32764
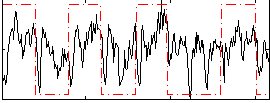 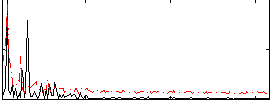 |
Correlation: 0.70869
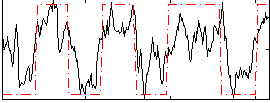  |
Subject: 66 |
GLM result cope: 02, threshold: 3
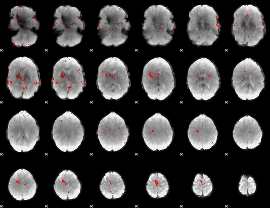 |
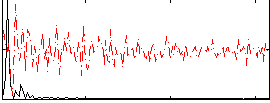
GLM result cope: 03, threshold: 2
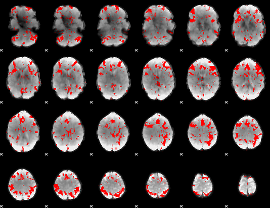 |
HAFNI result component: 315
Overlap rate: 0.41985
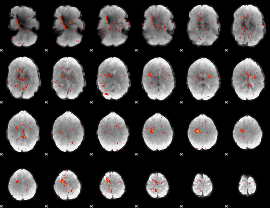 |
HAFNI result component: 207
Overlap rate: 0.5087
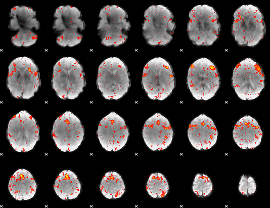 |
Correlation: 0.090856
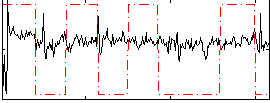 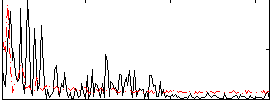 |
Correlation: 0.68685
  |
Subject: 67 |
GLM result cope: 02, threshold: 2
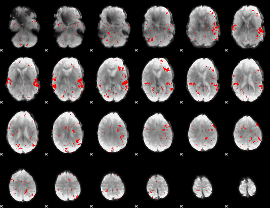 |
GLM result cope: 03, threshold: 3.5
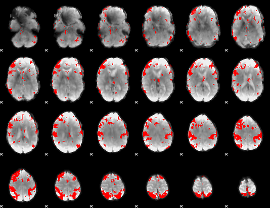 |
HAFNI result component: 397
Overlap rate: 0.80383
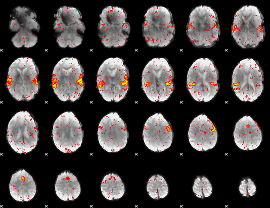 |
HAFNI result component: 202
Overlap rate: 0.43468
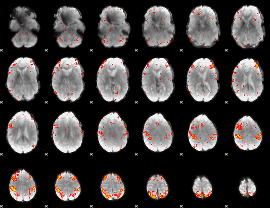 |
Correlation: 0.12548
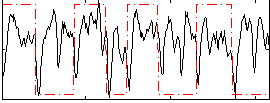 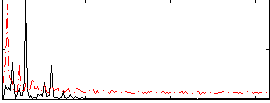 |
Correlation: 0.68206
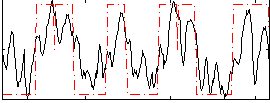 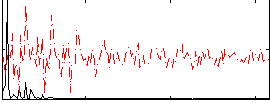 |
Subject: 68 |
GLM result cope: 02, threshold: 3.5
 |
GLM result cope: 03, threshold: 3
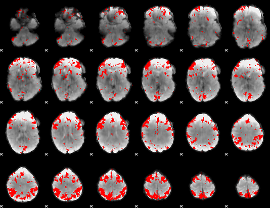 |
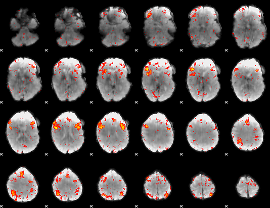
HAFNI result component: 301
Overlap rate: 0.29105
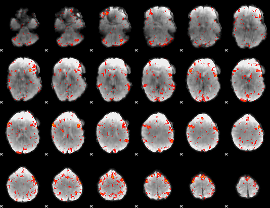 |
HAFNI result component: 141
Overlap rate: 0.31205
 |
Correlation: -0.42389
  |
Correlation: 0.65749
  |































































































































































































































































































































































































































































































































































