HAFNI report on finalized components on task: GAMBLING |
Totally 2 copes finalized |
Subject: 1 |
GLM result cope: 01, threshold: 3.5
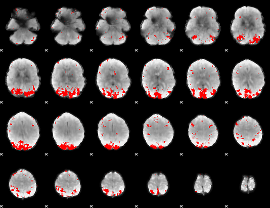 |
GLM result cope: 02, threshold: 3.5
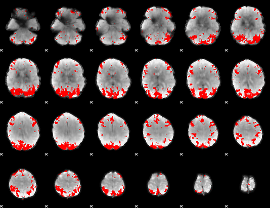 |
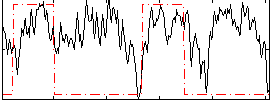
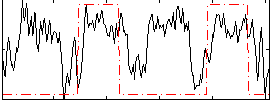
HAFNI result component: 184
Overlap rate: 0.42218
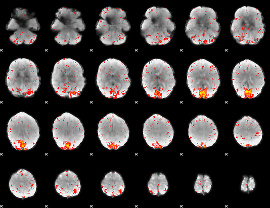 |
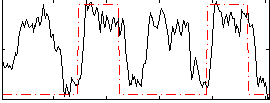
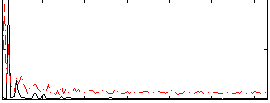
HAFNI result component: 396
Overlap rate: 0.45184
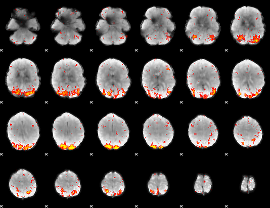 |
Correlation: 0.19985
  |
Correlation: 0.41664
  |
Subject: 2 |
GLM result cope: 01, threshold: 3.5
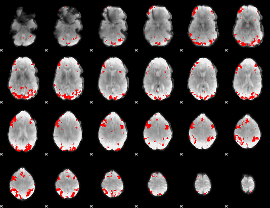 |
GLM result cope: 02, threshold: 3
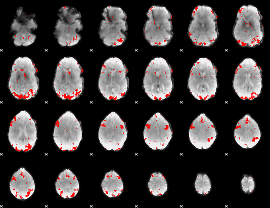 |
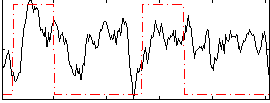
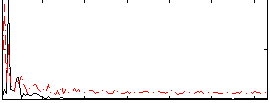
HAFNI result component: 142
Overlap rate: 0.39179
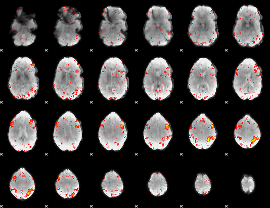 |
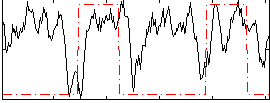
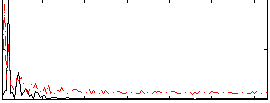
HAFNI result component: 141
Overlap rate: 0.50811
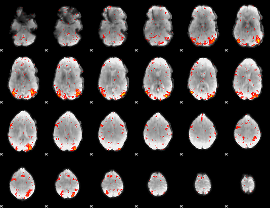 |
Correlation: 0.26288
  |
Correlation: 0.096982
  |
Subject: 3 |
GLM result cope: 01, threshold: 3
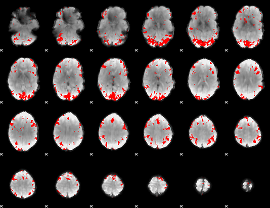 |
GLM result cope: 02, threshold: 4
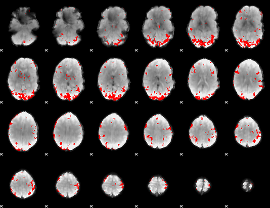 |
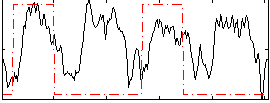
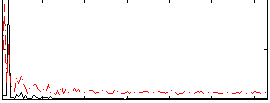
HAFNI result component: 285
Overlap rate: 0.49866
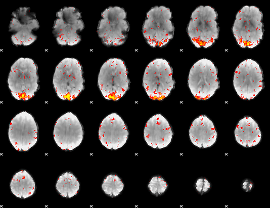 |
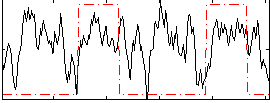
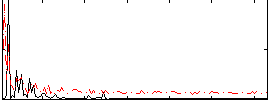
HAFNI result component: 319
Overlap rate: 0.29814
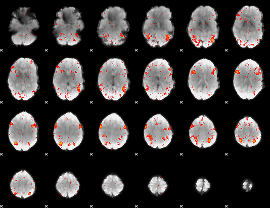 |
Correlation: 0.2204
  |
Correlation: 0.21811
  |
Subject: 4 |
GLM result cope: 01, threshold: 3.5
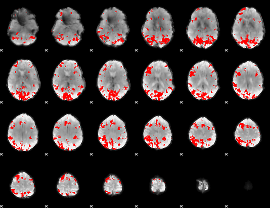 |
GLM result cope: 02, threshold: 3.5
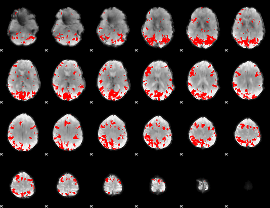 |
HAFNI result component: 211
Overlap rate: 0.34004
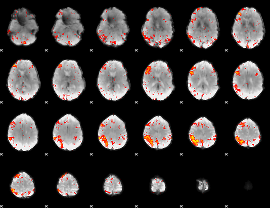 |
HAFNI result component: 211
Overlap rate: 0.31063
 |
Correlation: 0.12698
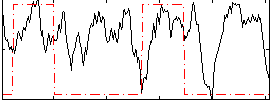  |
Correlation: -0.016102
  |
Subject: 5 |
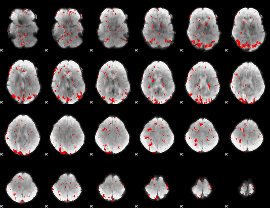
GLM result cope: 01, threshold: 3.5
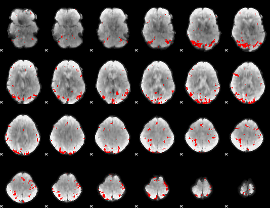 |
GLM result cope: 02, threshold: 2
 |
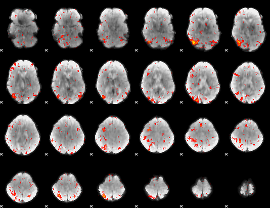
HAFNI result component: 318
Overlap rate: 0.43509
 |
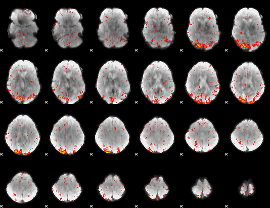
HAFNI result component: 050
Overlap rate: 0.77386
 |
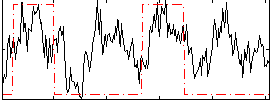
Correlation: 0.40882
  |
Correlation: 0.26049
  |
Subject: 6 |
GLM result cope: 01, threshold: 3
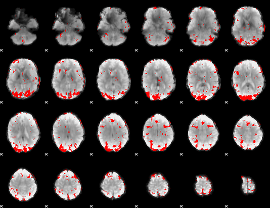 |
GLM result cope: 02, threshold: 4
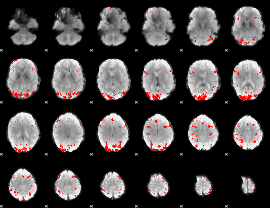 |
HAFNI result component: 095
Overlap rate: 0.5508
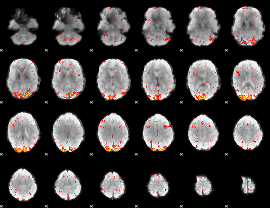 |
HAFNI result component: 264
Overlap rate: 0.35167
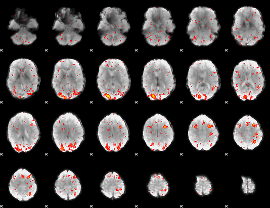 |
Correlation: 0.36704
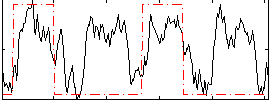 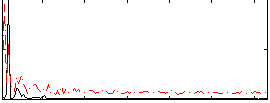 |
Correlation: 0.29399
  |
Subject: 7 |
GLM result cope: 01, threshold: 3.5
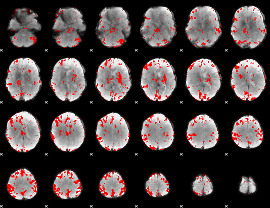 |
GLM result cope: 02, threshold: 3
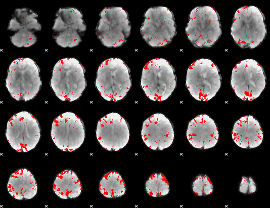 |
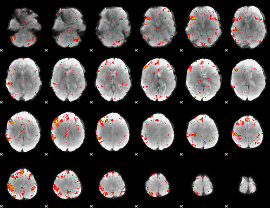
HAFNI result component: 397
Overlap rate: 0.25774
 |
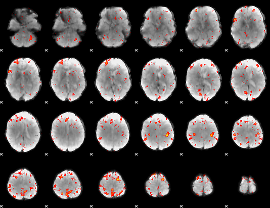
HAFNI result component: 024
Overlap rate: 0.48595
 |
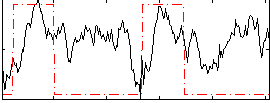
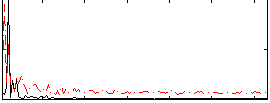
Correlation: 0.2632
  |
Correlation: 0.06917
  |
Subject: 8 |
GLM result cope: 01, threshold: 3
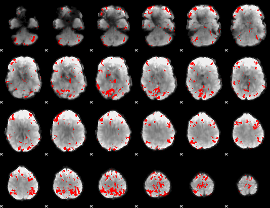 |
GLM result cope: 02, threshold: 3.5
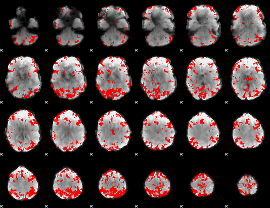 |
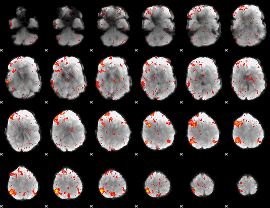
HAFNI result component: 105
Overlap rate: 0.31684
 |
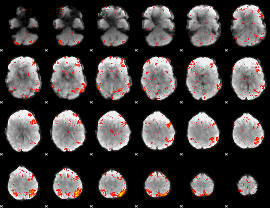
HAFNI result component: 137
Overlap rate: 0.29515
 |
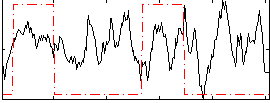
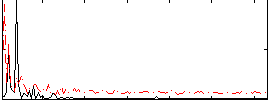
Correlation: 0.13283
  |
Correlation: 0.36644
  |
Subject: 9 |
GLM result cope: 01, threshold: 3.5
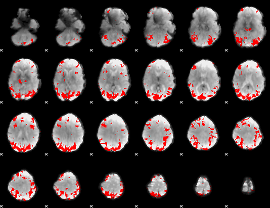 |
GLM result cope: 02, threshold: 3.5
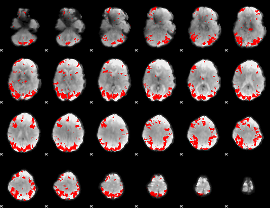 |
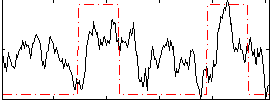
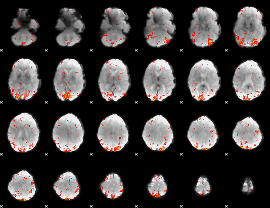
HAFNI result component: 228
Overlap rate: 0.38054
 |
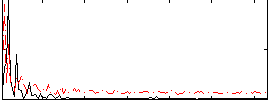
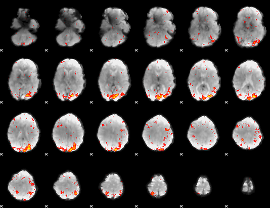
HAFNI result component: 066
Overlap rate: 0.28007
 |
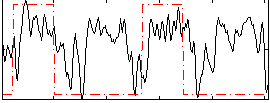
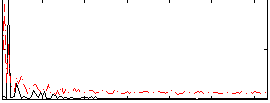
Correlation: 0.29775
  |
Correlation: 0.27526
  |
Subject: 10 |
GLM result cope: 01, threshold: 3.5
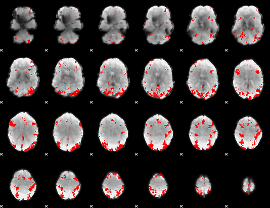 |
GLM result cope: 02, threshold: 4
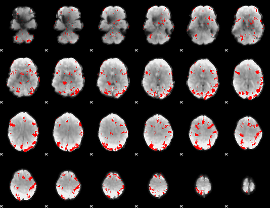 |
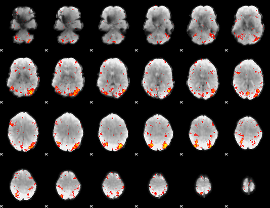
HAFNI result component: 069
Overlap rate: 0.50888
 |
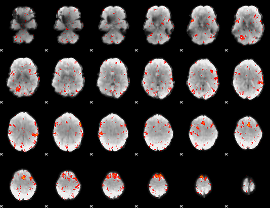
HAFNI result component: 349
Overlap rate: 0.27575
 |
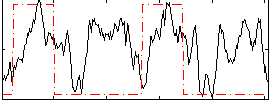
Correlation: 0.27815
 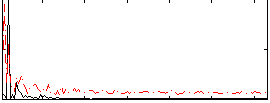 |
Correlation: 0.17968
  |
Subject: 11 |
GLM result cope: 01, threshold: 3.5
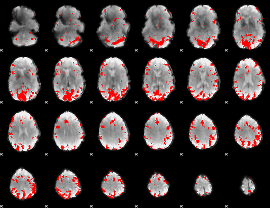 |
GLM result cope: 02, threshold: 3.5
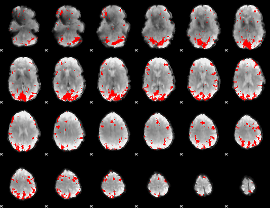 |
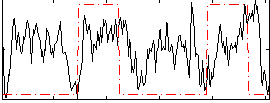
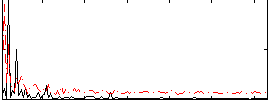
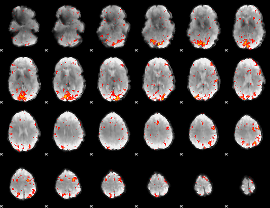
HAFNI result component: 114
Overlap rate: 0.40245
 |
HAFNI result component: 114
Overlap rate: 0.47323
 |
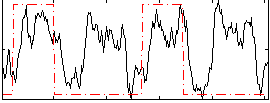
Correlation: 0.31873
  |
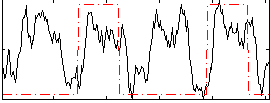
Correlation: 0.27085
  |
Subject: 12 |
GLM result cope: 01, threshold: 3
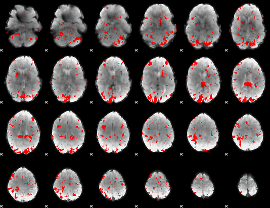 |
GLM result cope: 02, threshold: 3.5
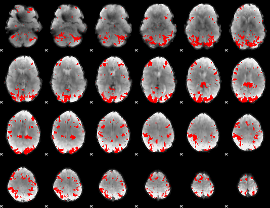 |
HAFNI result component: 130
Overlap rate: 0.38399
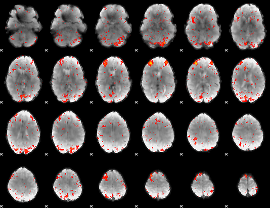 |
HAFNI result component: 310
Overlap rate: 0.23959
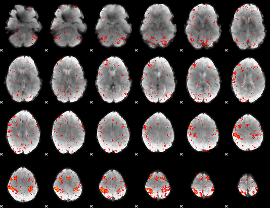 |
Correlation: 0.43602
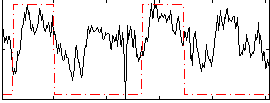 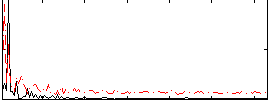 |
Correlation: 0.053669
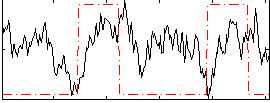  |
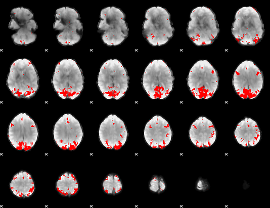
Subject: 13 |
GLM result cope: 01, threshold: 4
 |
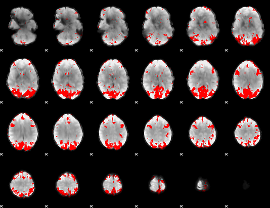
GLM result cope: 02, threshold: 3.5
 |
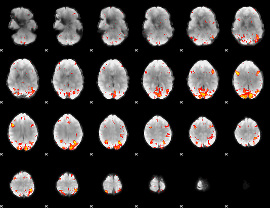
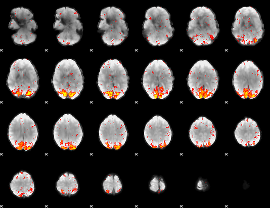
HAFNI result component: 363
Overlap rate: 0.5209
 |
HAFNI result component: 123
Overlap rate: 0.42801
 |
Correlation: 0.34509
  |
Correlation: 0.3807
  |
Subject: 14 |
GLM result cope: 01, threshold: 3.5
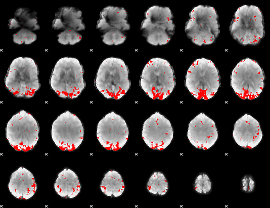 |
GLM result cope: 02, threshold: 3.5
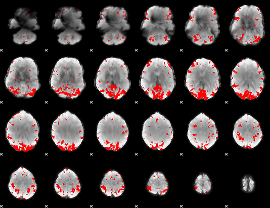 |
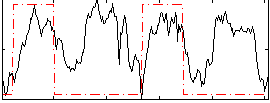
HAFNI result component: 157
Overlap rate: 0.59109
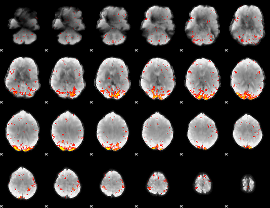 |
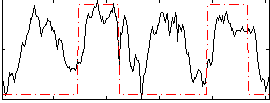
HAFNI result component: 157
Overlap rate: 0.41741
 |
Correlation: 0.2513
  |
Correlation: 0.41637
  |
Subject: 15 |
GLM result cope: 01, threshold: 4
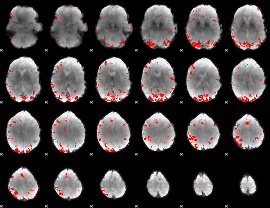 |
GLM result cope: 02, threshold: 3
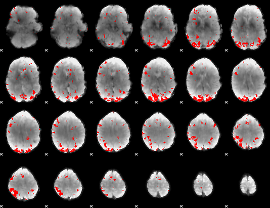 |
HAFNI result component: 137
Overlap rate: 0.39274
 |
HAFNI result component: 399
Overlap rate: 0.49382
 |
Correlation: 0.30495
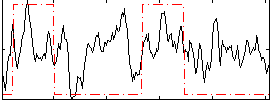 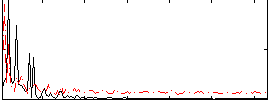 |
Correlation: 0.21271
  |
Subject: 16 |
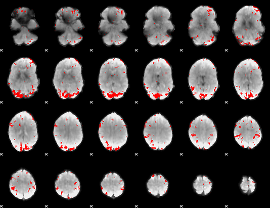
GLM result cope: 01, threshold: 3.5
 |
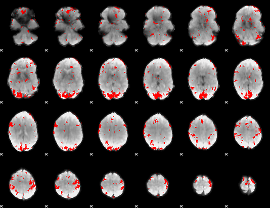
GLM result cope: 02, threshold: 3
 |
HAFNI result component: 002
Overlap rate: 0.45221
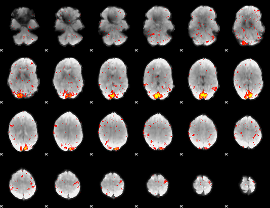 |
HAFNI result component: 002
Overlap rate: 0.50415
 |
Correlation: 0.16661
  |
Correlation: 0.36946
  |
Subject: 17 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 18 |
GLM result cope: 01, threshold: 3.5
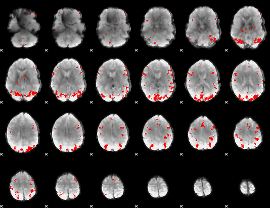 |
GLM result cope: 02, threshold: 3
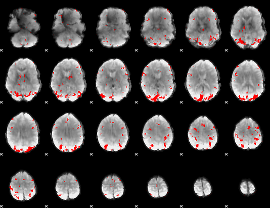 |
HAFNI result component: 386
Overlap rate: 0.55346
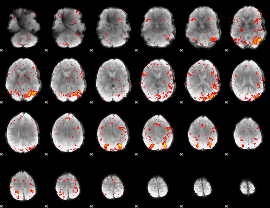 |
HAFNI result component: 222
Overlap rate: 0.55789
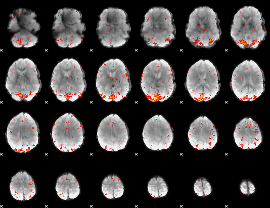 |
Correlation: 0.20145
  |
Correlation: 0.38021
  |
Subject: 19 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
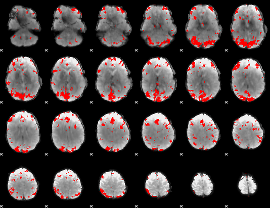
Subject: 20 |
GLM result cope: 01, threshold: 4
 |
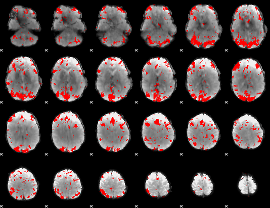
GLM result cope: 02, threshold: 3.5
 |
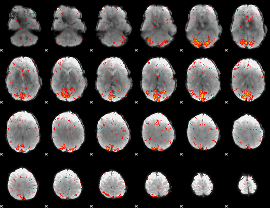
HAFNI result component: 335
Overlap rate: 0.19536
 |
HAFNI result component: 188
Overlap rate: 0.20737
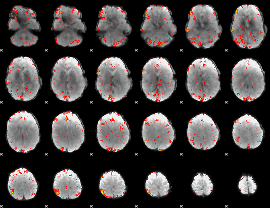 |
Correlation: 0.36004
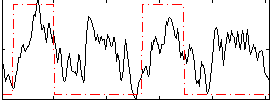  |
Correlation: 0.1815
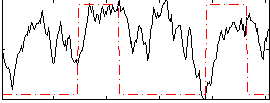  |
Subject: 21 |
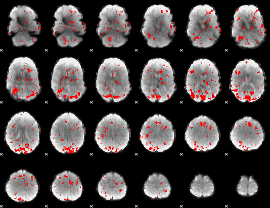
GLM result cope: 01, threshold: 2
 |
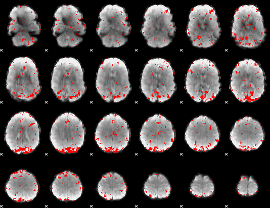
GLM result cope: 02, threshold: 2.5
 |
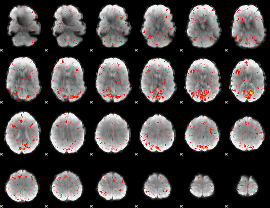
HAFNI result component: 030
Overlap rate: 0.51542
 |
HAFNI result component: 235
Overlap rate: 0.52659
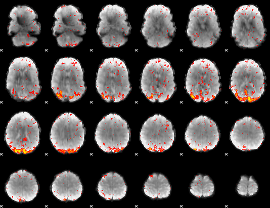 |
Correlation: 0.27654
  |
Correlation: 0.1383
  |
Subject: 22 |
GLM result cope: 01, threshold: 3
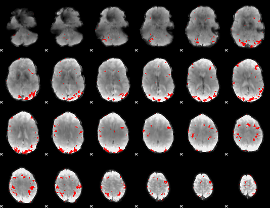 |
GLM result cope: 02, threshold: 2
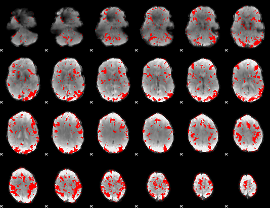 |
HAFNI result component: 055
Overlap rate: 0.023728
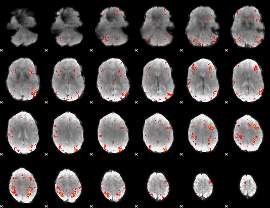 |
HAFNI result component: 055
Overlap rate: 0.027722
 |
Correlation: 0.22268
  |
Correlation: 0.16016
  |
Subject: 23 |
GLM result cope: 01, threshold: 3.5
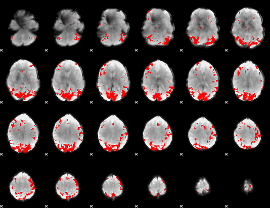 |
GLM result cope: 02, threshold: 4
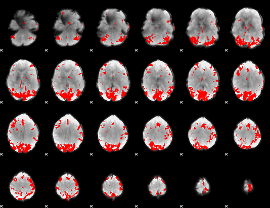 |
HAFNI result component: 205
Overlap rate: 0.45423
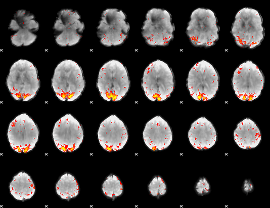 |
HAFNI result component: 208
Overlap rate: 0.28747
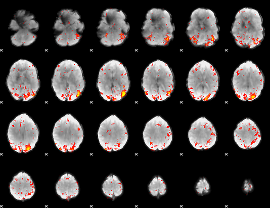 |
Correlation: 0.29208
  |
Correlation: 0.28509
  |
Subject: 24 |
GLM result cope: 01, threshold: 3.5
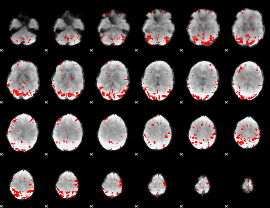 |
GLM result cope: 02, threshold: 4
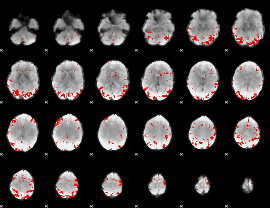 |
HAFNI result component: 386
Overlap rate: 0.37538
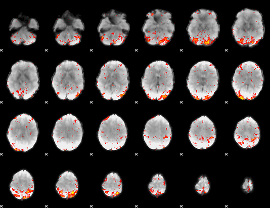 |
HAFNI result component: 346
Overlap rate: 0.33706
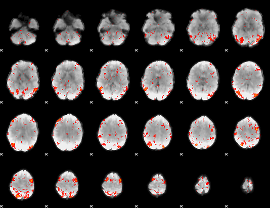 |
Correlation: 0.2647
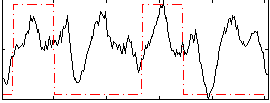 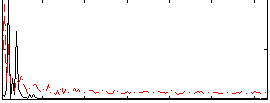 |
Correlation: 0.029104
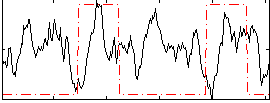 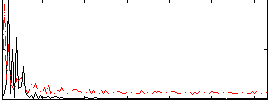 |
Subject: 25 |
GLM result cope: 01, threshold: 3
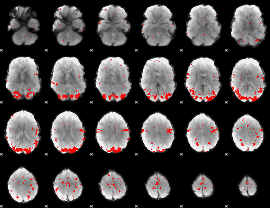 |
GLM result cope: 02, threshold: 3
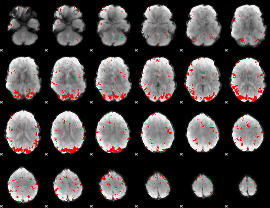 |
HAFNI result component: 299
Overlap rate: 0.62352
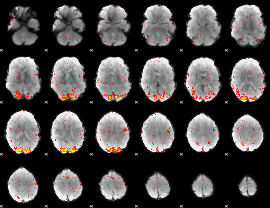 |
HAFNI result component: 299
Overlap rate: 0.61449
 |
Correlation: 0.27559
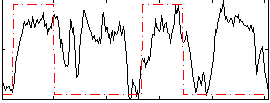  |
Correlation: 0.2916
  |
Subject: 26 |
GLM result cope: 01, threshold: 3.5
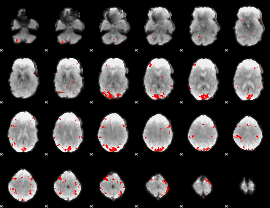 |
GLM result cope: 02, threshold: 3
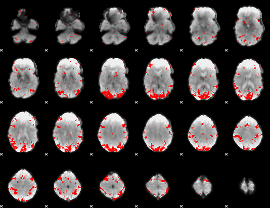 |
HAFNI result component: 204
Overlap rate: 0.4642
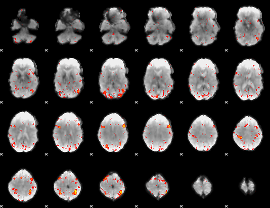 |
HAFNI result component: 204
Overlap rate: 0.4534
 |
Correlation: 0.15821
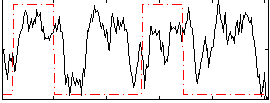 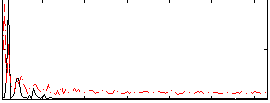 |
Correlation: 0.26739
  |
Subject: 27 |
GLM result cope: 01, threshold: 3
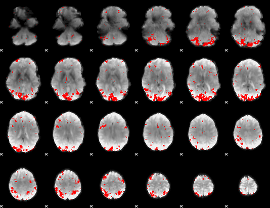 |
GLM result cope: 02, threshold: 3.5
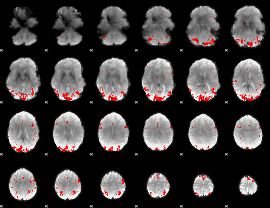 |
HAFNI result component: 349
Overlap rate: 0.57382
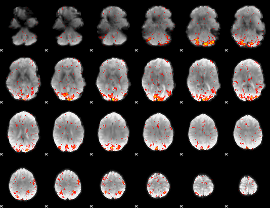 |
HAFNI result component: 349
Overlap rate: 0.56749
 |
Correlation: 0.23514
  |
Correlation: 0.23651
  |
Subject: 28 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 29 |
GLM result cope: 01, threshold: 3.5
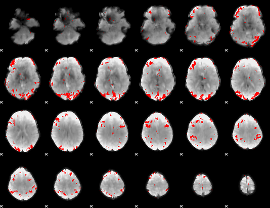 |
GLM result cope: 02, threshold: 3.5
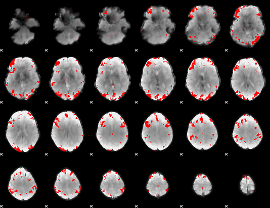 |
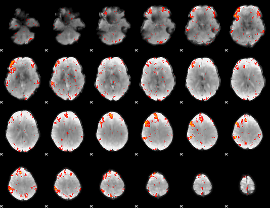
HAFNI result component: 372
Overlap rate: 0.25692
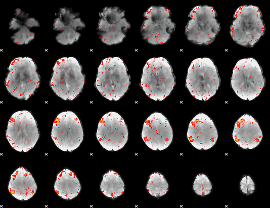 |
HAFNI result component: 335
Overlap rate: 0.40936
 |
Correlation: 0.20041
  |
Correlation: 0.16214
  |
Subject: 30 |
GLM result cope: 01, threshold: 3.5
 |
GLM result cope: 02, threshold: 3.5
 |
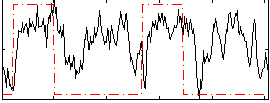
HAFNI result component: 349
Overlap rate: 0.5707
 |
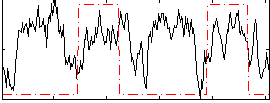
HAFNI result component: 349
Overlap rate: 0.39109
 |
Correlation: 0.31655
  |
Correlation: 0.11445
  |
Subject: 31 |
GLM result cope: 01, threshold: 2.5
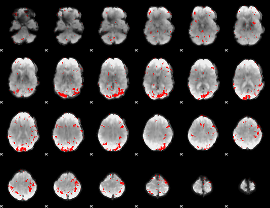 |
GLM result cope: 02, threshold: 3
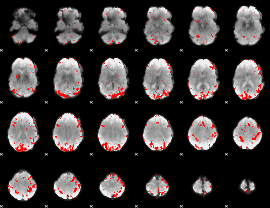 |
HAFNI result component: 310
Overlap rate: 0.69709
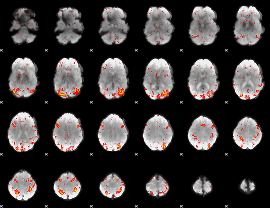 |
HAFNI result component: 310
Overlap rate: 0.56312
 |
Correlation: 0.21822
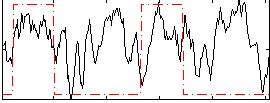  |
Correlation: 0.23229
  |
Subject: 32 |
GLM result cope: 01, threshold: 3
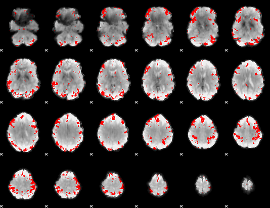 |
GLM result cope: 02, threshold: 3
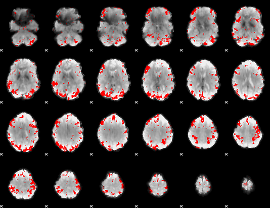 |
HAFNI result component: 396
Overlap rate: 0.45891
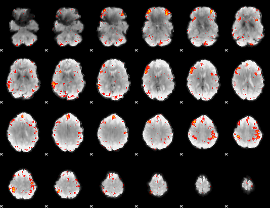 |
HAFNI result component: 396
Overlap rate: 0.41061
 |
Correlation: 0.38003
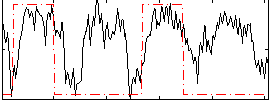 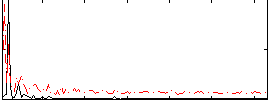 |
Correlation: 0.22266
  |
Subject: 33 |
GLM result cope: 01, threshold: 3
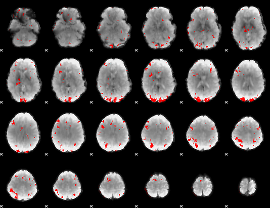 |
GLM result cope: 02, threshold: 3
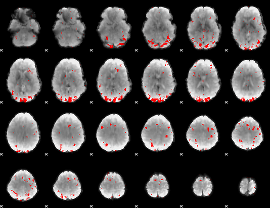 |
HAFNI result component: 017
Overlap rate: 0.336
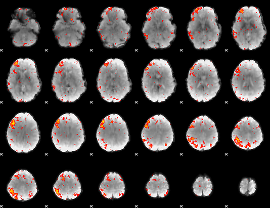 |
HAFNI result component: 278
Overlap rate: 0.6285
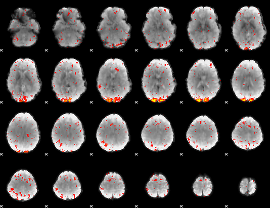 |
Correlation: 0.076102
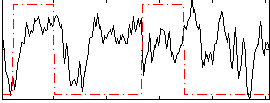 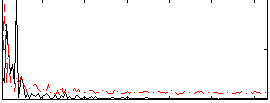 |
Correlation: 0.31278
  |
Subject: 34 |
GLM result cope: 01, threshold: 3.5
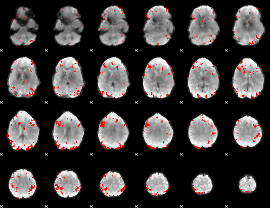 |
GLM result cope: 02, threshold: 3.5
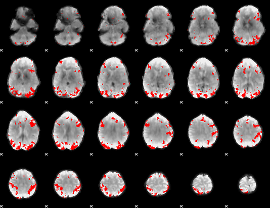 |
HAFNI result component: 003
Overlap rate: 0.3561
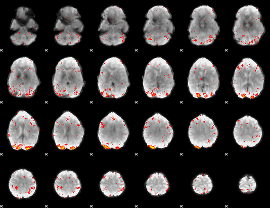 |
HAFNI result component: 158
Overlap rate: 0.4159
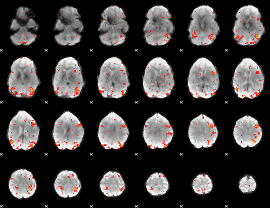 |
Correlation: 0.23907
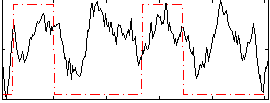 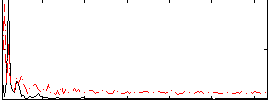 |
Correlation: 0.24418
  |
Subject: 35 |
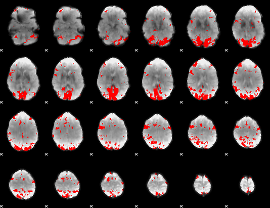
GLM result cope: 01, threshold: 3
 |
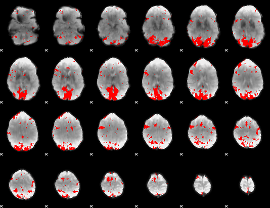
GLM result cope: 02, threshold: 3
 |
HAFNI result component: 099
Overlap rate: 0.47694
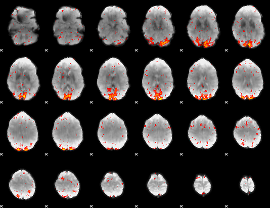 |
HAFNI result component: 086
Overlap rate: 0.38738
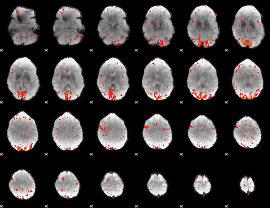 |
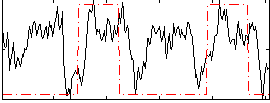
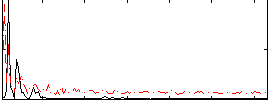
Correlation: 0.29866
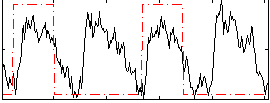 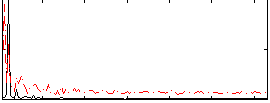 |
Correlation: 0.20352
  |
Subject: 36 |
GLM result cope: 01, threshold: 3.5
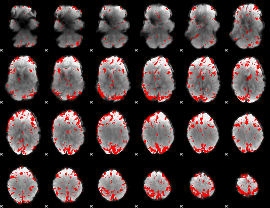 |
GLM result cope: 02, threshold: 3.5
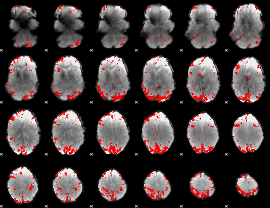 |
HAFNI result component: 261
Overlap rate: 0.42702
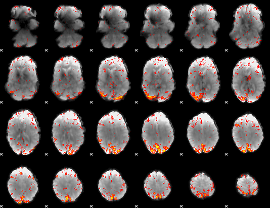 |
HAFNI result component: 039
Overlap rate: 0.36548
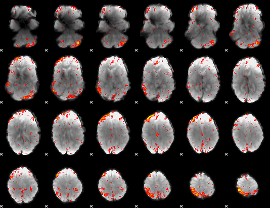 |
Correlation: 0.35512
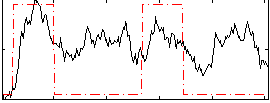 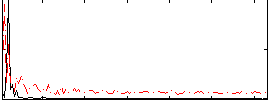 |
Correlation: 0.048348
  |
Subject: 37 |
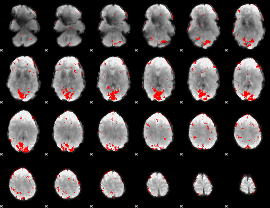
GLM result cope: 01, threshold: 3
 |
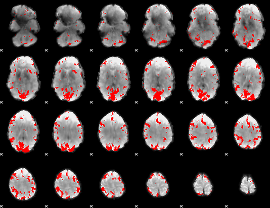
GLM result cope: 02, threshold: 3
 |
HAFNI result component: 188
Overlap rate: 0.45309
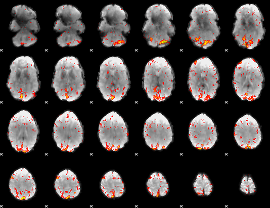 |
HAFNI result component: 225
Overlap rate: 0.33432
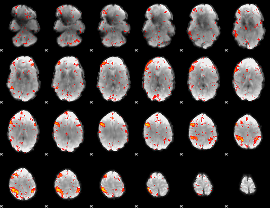 |
Correlation: 0.24989
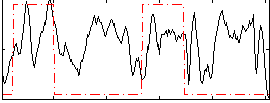 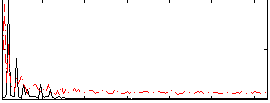 |
Correlation: 0.26798
  |
Subject: 38 |
GLM result cope: 01, threshold: 3
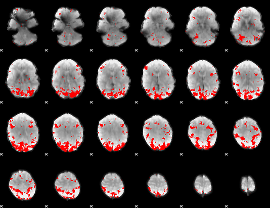 |
GLM result cope: 02, threshold: 3
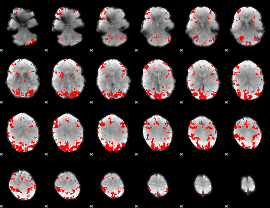 |
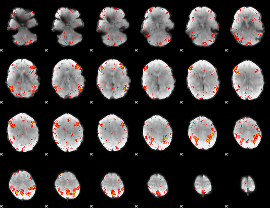
HAFNI result component: 320
Overlap rate: 0.24047
 |
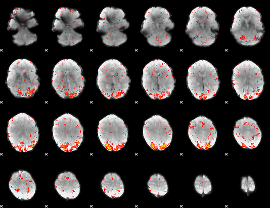
HAFNI result component: 101
Overlap rate: 0.45287
 |
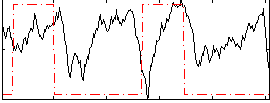
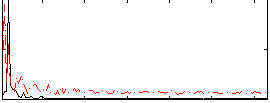
Correlation: 0.046537
  |
Correlation: 0.29532
  |
Subject: 39 |
GLM result cope: 01, threshold: 2.5
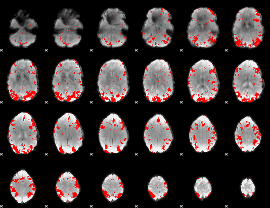 |
GLM result cope: 02, threshold: 3
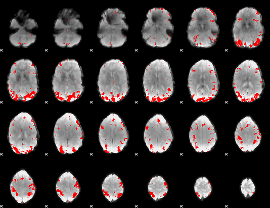 |
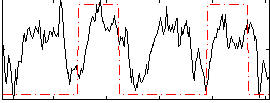
HAFNI result component: 304
Overlap rate: 0.41607
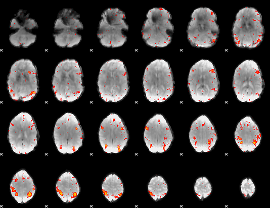 |
HAFNI result component: 070
Overlap rate: 0.46555
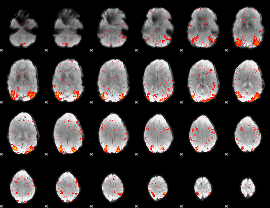 |
Correlation: 0.20602
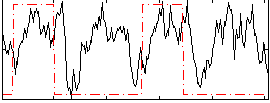 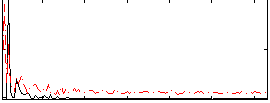 |
Correlation: 0.11847
  |
Subject: 40 |
GLM result cope: 01, threshold: 3.5
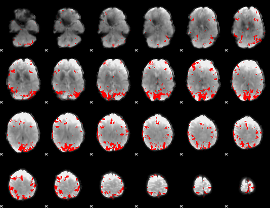 |
GLM result cope: 02, threshold: 3.5
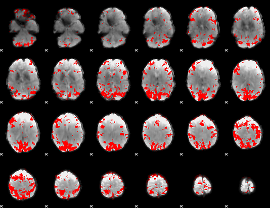 |
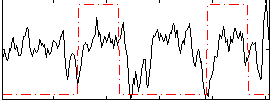
HAFNI result component: 387
Overlap rate: 0.25959
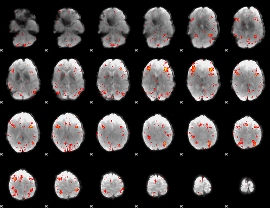 |
HAFNI result component: 291
Overlap rate: 0.25191
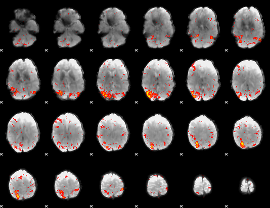 |
Correlation: 0.1504
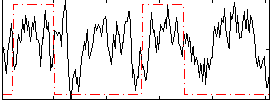 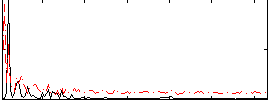 |
Correlation: 0.24237
  |
Subject: 41 |
GLM result cope: 01, threshold: 3
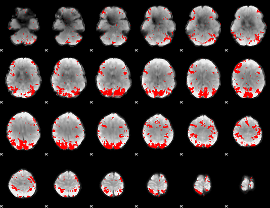 |
GLM result cope: 02, threshold: 3
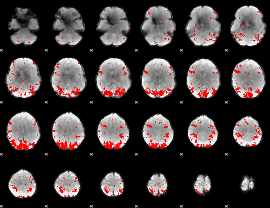 |
HAFNI result component: 247
Overlap rate: 0.23923
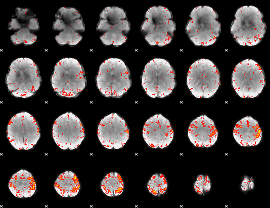 |
HAFNI result component: 327
Overlap rate: 0.381
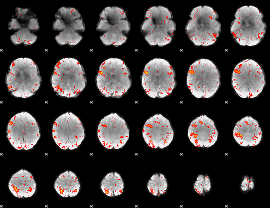 |
Correlation: 0.17347
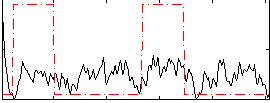 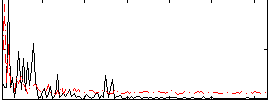 |
Correlation: 0.22395
  |
Subject: 42 |
GLM result cope: 01, threshold: 3
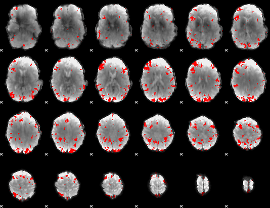 |
GLM result cope: 02, threshold: 3
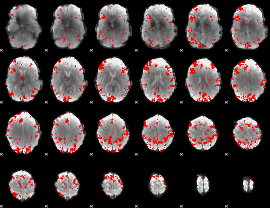 |
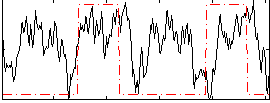
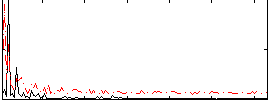
HAFNI result component: 073
Overlap rate: 0.30732
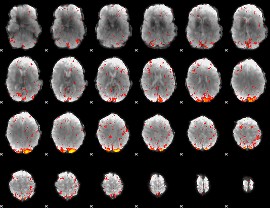 |
HAFNI result component: 362
Overlap rate: 0.45903
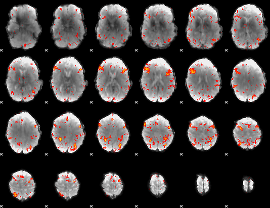 |
Correlation: 0.40361
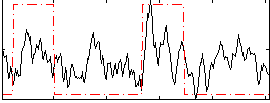 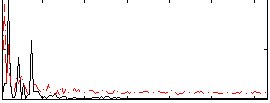 |
Correlation: 0.13815
  |
Subject: 43 |
GLM result cope: 01, threshold: 3
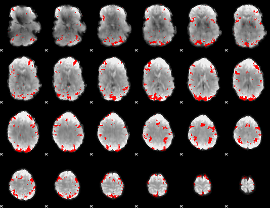 |
GLM result cope: 02, threshold: 3
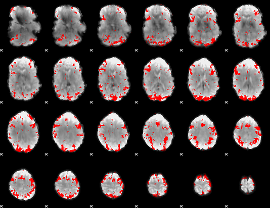 |
HAFNI result component: 258
Overlap rate: 0.27758
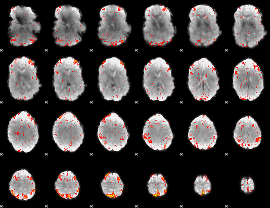 |
HAFNI result component: 096
Overlap rate: 0.46697
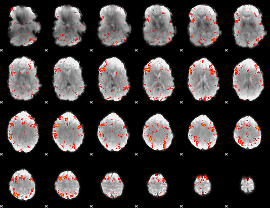 |
Correlation: 0.2217
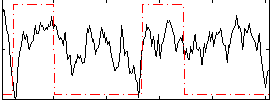 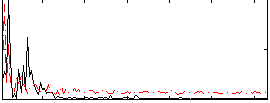 |
Correlation: 0.088126
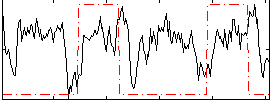  |
Subject: 44 |
GLM result cope: 01, threshold: 3
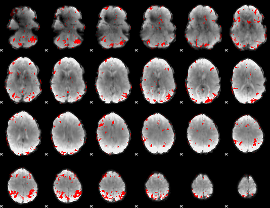 |
GLM result cope: 02, threshold: 2.5
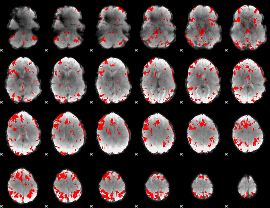 |
HAFNI result component: 184
Overlap rate: 0.23991
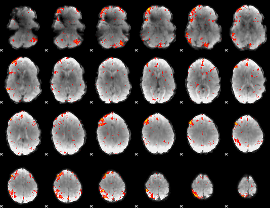 |
HAFNI result component: 170
Overlap rate: 0.38785
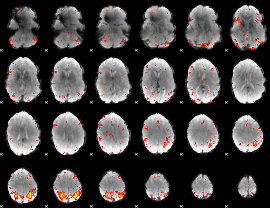 |
Correlation: 0.16383
  |
Correlation: 0.3348
  |
Subject: 45 |
GLM result cope: 01, threshold: 3
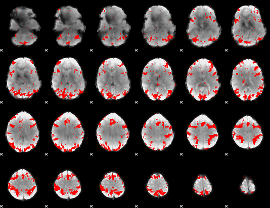 |
GLM result cope: 02, threshold: 3
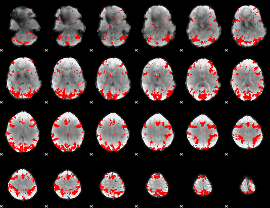 |
HAFNI result component: 200
Overlap rate: 0.38702
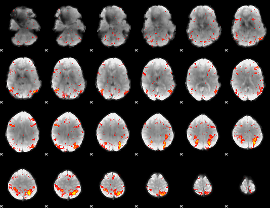 |
HAFNI result component: 370
Overlap rate: 0.25057
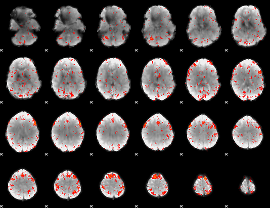 |
Correlation: 0.18492
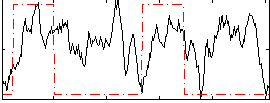 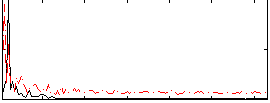 |
Correlation: 0.26422
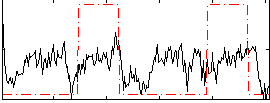 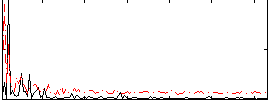 |
Subject: 46 |
GLM result cope: 01, threshold: 3
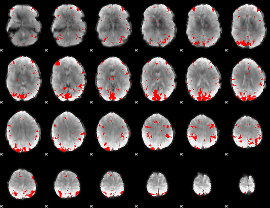 |
GLM result cope: 02, threshold: 3
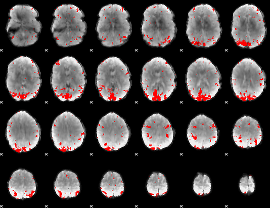 |
HAFNI result component: 194
Overlap rate: 0.4384
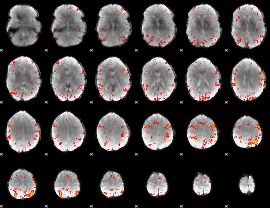 |
HAFNI result component: 106
Overlap rate: 0.49035
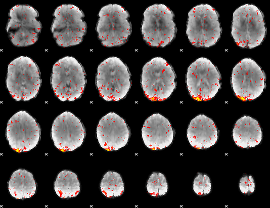 |
Correlation: 0.25718
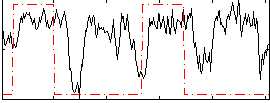  |
Correlation: 0.30534
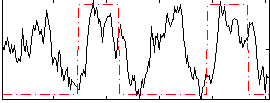  |
Subject: 47 |
GLM result cope: 01, threshold: 2.5
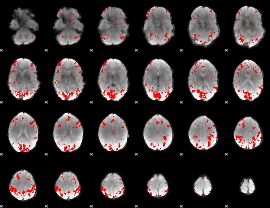 |
GLM result cope: 02, threshold: 3
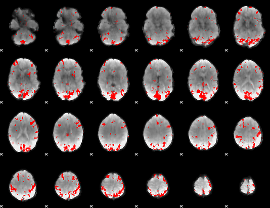 |
HAFNI result component: 295
Overlap rate: 0.28481
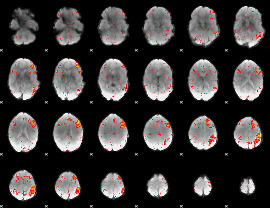 |
HAFNI result component: 044
Overlap rate: 0.43575
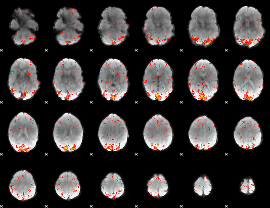 |
Correlation: 0.32604
  |
Correlation: 0.2648
  |
Subject: 48 |
GLM result cope: 01, threshold: 3
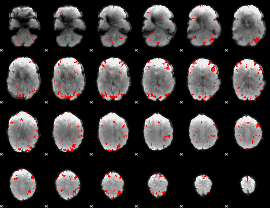 |
GLM result cope: 02, threshold: 3
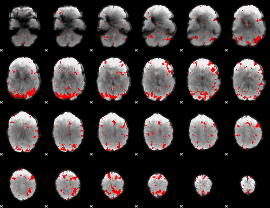 |
HAFNI result component: 092
Overlap rate: 0.53777
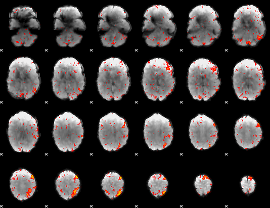 |
HAFNI result component: 352
Overlap rate: 0.43594
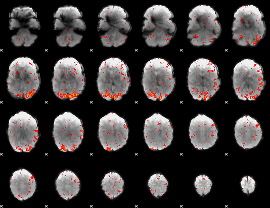 |
Correlation: 0.26927
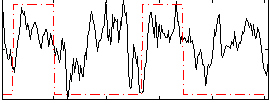 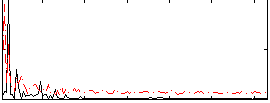 |
Correlation: 0.33526
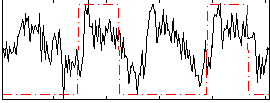  |
Subject: 49 |
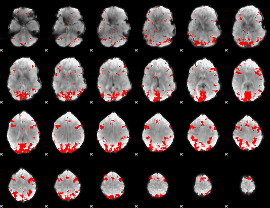
GLM result cope: 01, threshold: 3
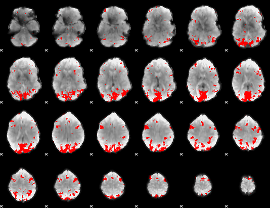 |
GLM result cope: 02, threshold: 3
 |
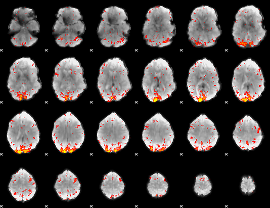
HAFNI result component: 286
Overlap rate: 0.59881
 |
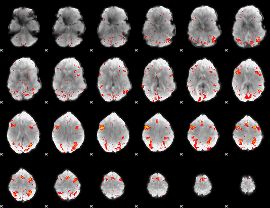
HAFNI result component: 250
Overlap rate: 0.45609
 |
Correlation: 0.26327
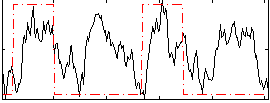  |
Correlation: 0.19293
  |
Subject: 50 |
GLM result cope: 01, threshold: 3
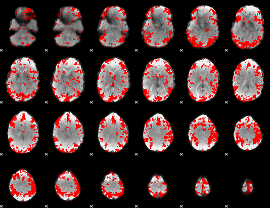 |
GLM result cope: 02, threshold: 3
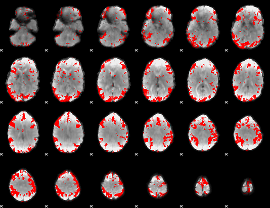 |
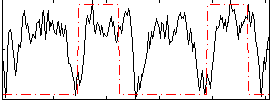
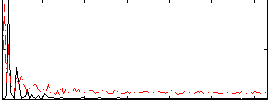
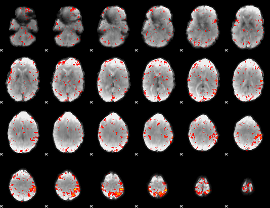
HAFNI result component: 360
Overlap rate: 0.21192
 |
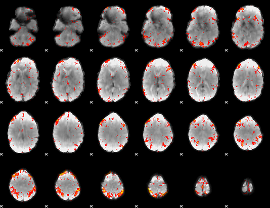
HAFNI result component: 123
Overlap rate: 0.38281
 |
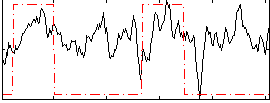
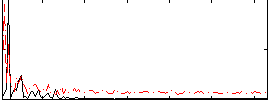
Correlation: 0.3412
  |
Correlation: 0.12406
  |
Subject: 51 |
GLM result cope: 01, threshold: 3
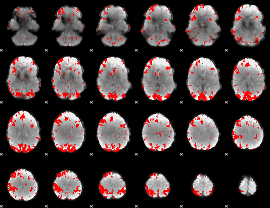 |
GLM result cope: 02, threshold: 3
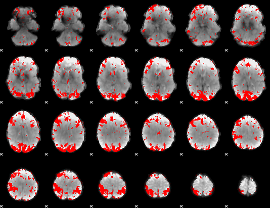 |
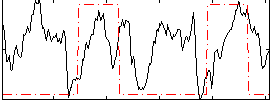
HAFNI result component: 021
Overlap rate: 0.25184
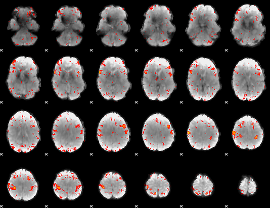 |
HAFNI result component: 220
Overlap rate: 0.36414
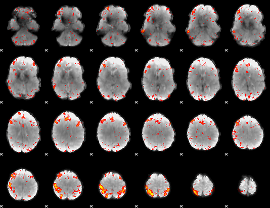 |
Correlation: 0.25875
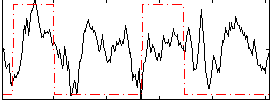 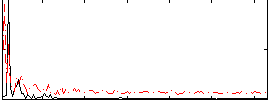 |
Correlation: 0.12419
  |
Subject: 52 |
GLM result cope: 01, threshold: 3
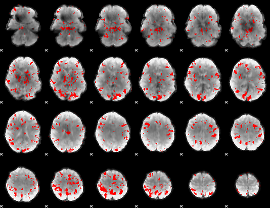 |
GLM result cope: 02, threshold: 3
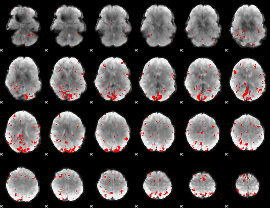 |
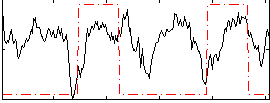
HAFNI result component: 054
Overlap rate: 0.30883
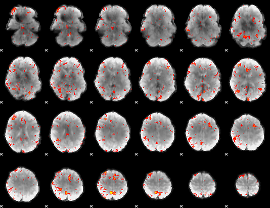 |
HAFNI result component: 293
Overlap rate: 0.71741
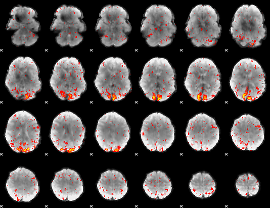 |
Correlation: 0.21374
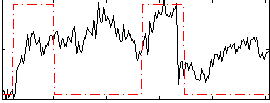 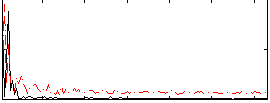 |
Correlation: 0.33001
  |
Subject: 53 |
GLM result cope: 01, threshold: 3
 |
GLM result cope: 02, threshold: 3
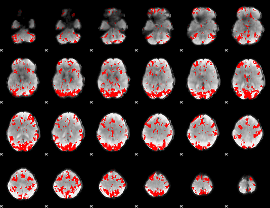 |
HAFNI result component: 130
Overlap rate: 0.31715
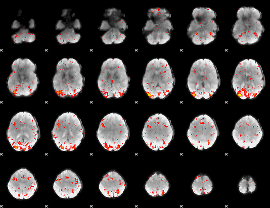 |
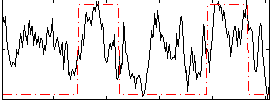
HAFNI result component: 294
Overlap rate: 0.23029
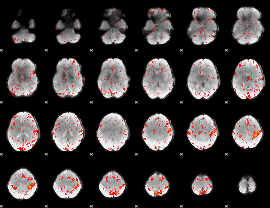 |
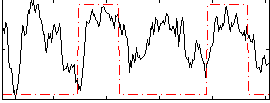
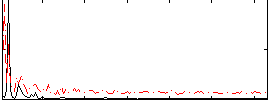
Correlation: 0.3101
  |
Correlation: 0.41717
  |
Subject: 54 |
GLM result cope: 01, threshold: 3
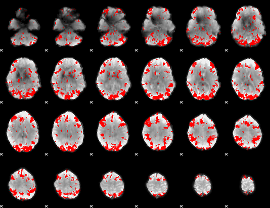 |
GLM result cope: 02, threshold: 3
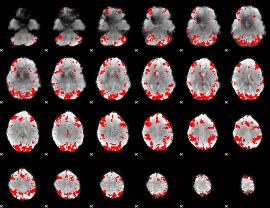 |
HAFNI result component: 208
Overlap rate: 0.28769
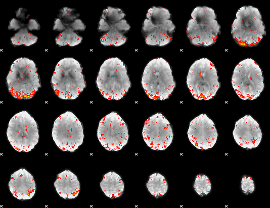 |
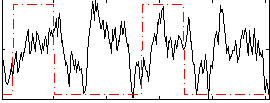
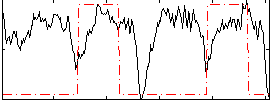
HAFNI result component: 286
Overlap rate: 0.31027
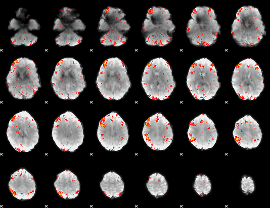 |
Correlation: 0.12985
 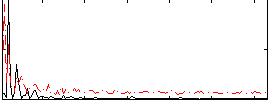 |
Correlation: 0.13367
  |
Subject: 55 |
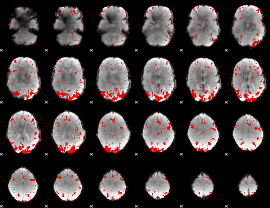
GLM result cope: 01, threshold: 2.5
 |
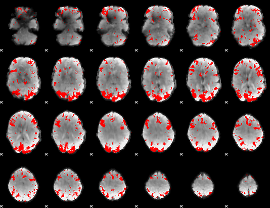
GLM result cope: 02, threshold: 3
 |
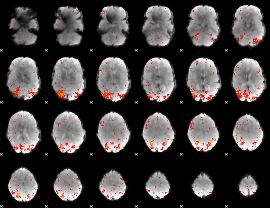
HAFNI result component: 076
Overlap rate: 0.31297
 |
HAFNI result component: 241
Overlap rate: 0.34487
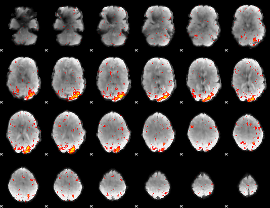 |
Correlation: 0.14054
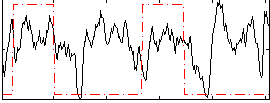 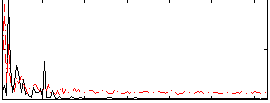 |
Correlation: 0.24449
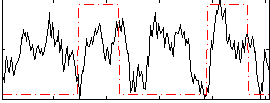  |
Subject: 56 |
GLM result cope: 01, threshold: 3
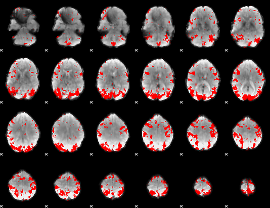 |
GLM result cope: 02, threshold: 3
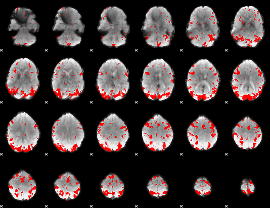 |
HAFNI result component: 395
Overlap rate: 0.39818
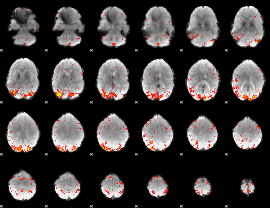 |
HAFNI result component: 359
Overlap rate: 0.4391
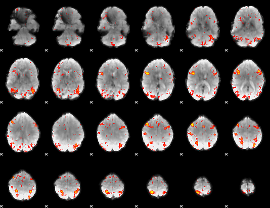 |
Correlation: 0.42012
  |
Correlation: 0.20598
  |
Subject: 57 |
GLM result cope: 01, threshold: 3
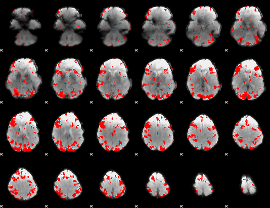 |
GLM result cope: 02, threshold: 3
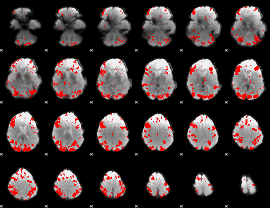 |
HAFNI result component: 334
Overlap rate: 0.23026
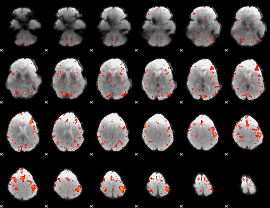 |
HAFNI result component: 137
Overlap rate: 0.27962
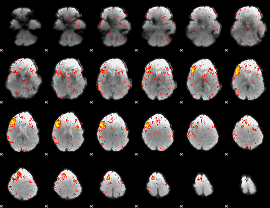 |
Correlation: 0.20514
  |
Correlation: 0.22654
  |
Subject: 58 |
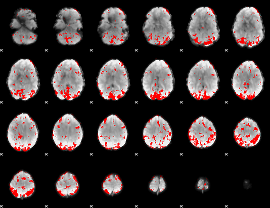
GLM result cope: 01, threshold: 2.5
 |
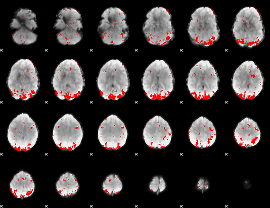
GLM result cope: 02, threshold: 3
 |
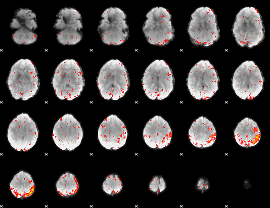
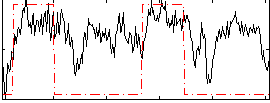
HAFNI result component: 049
Overlap rate: 0.36221
 |
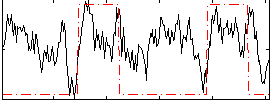
HAFNI result component: 028
Overlap rate: 0.32723
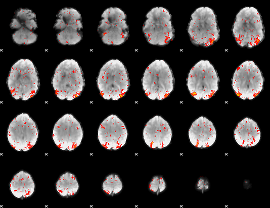 |
Correlation: 0.47646
  |
Correlation: 0.3315
  |
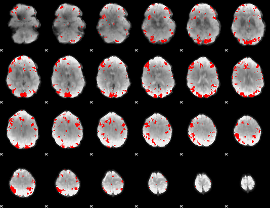
Subject: 59 |
GLM result cope: 01, threshold: 3
 |
GLM result cope: 02, threshold: 2
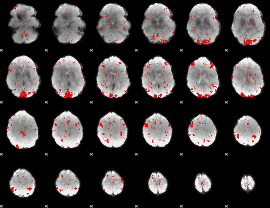 |
HAFNI result component: 385
Overlap rate: 0.20411
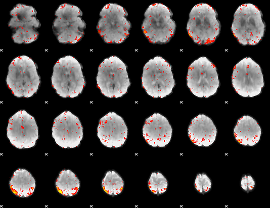 |
HAFNI result component: 069
Overlap rate: 0.40055
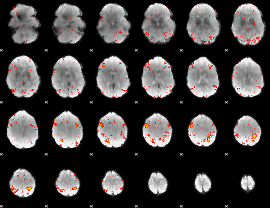 |
Correlation: 0.39291
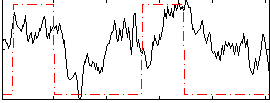  |
Correlation: 0.048325
  |
Subject: 60 |
GLM result cope: 01, threshold: 2.5
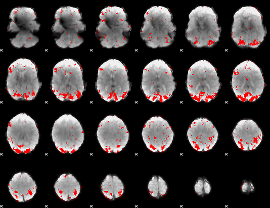 |
GLM result cope: 02, threshold: 3
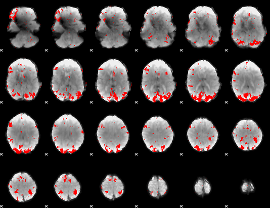 |
HAFNI result component: 341
Overlap rate: 0.58385
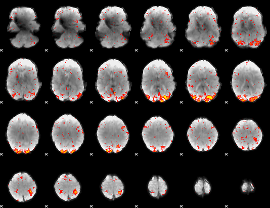 |
HAFNI result component: 067
Overlap rate: 0.32902
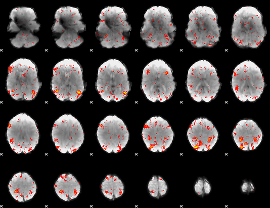 |
Correlation: 0.25498
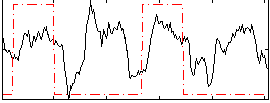  |
Correlation: -0.0886
  |
Subject: 61 |
GLM result cope: 01, threshold: 3
 |
GLM result cope: 02, threshold: 3
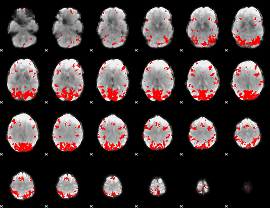 |
HAFNI result component: 396
Overlap rate: 0.50221
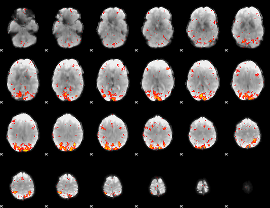 |
HAFNI result component: 298
Overlap rate: 0.29976
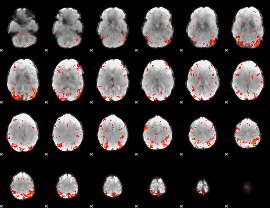 |
Correlation: 0.24762
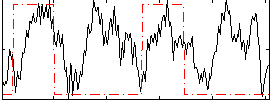  |
Correlation: 0.040467
  |
Subject: 62 |
GLM result cope: 01, threshold: 3
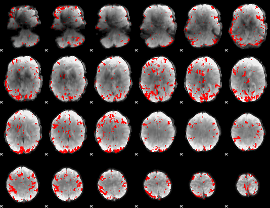 |
GLM result cope: 02, threshold: 3
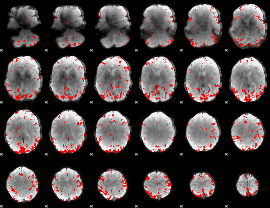 |
HAFNI result component: 109
Overlap rate: 0.33845
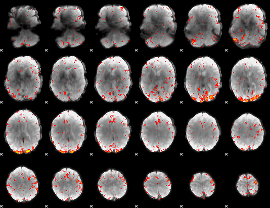 |
HAFNI result component: 374
Overlap rate: 0.34502
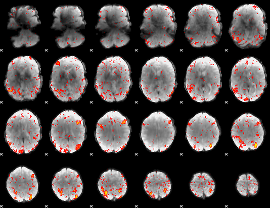 |
Correlation: 0.3567
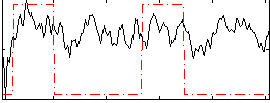 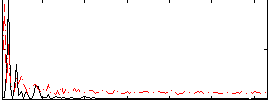 |
Correlation: 0.13108
  |
Subject: 63 |
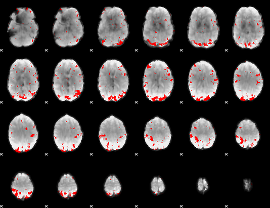
GLM result cope: 01, threshold: 3
 |
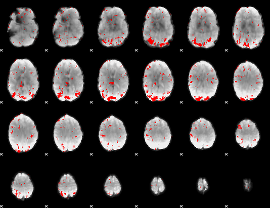
GLM result cope: 02, threshold: 2.5
 |
HAFNI result component: 247
Overlap rate: 0.33552
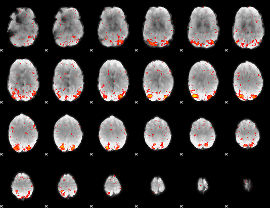 |
HAFNI result component: 308
Overlap rate: 0.67849
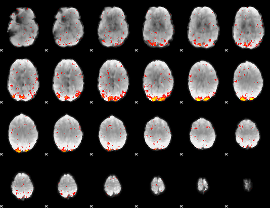 |
Correlation: 0.3544
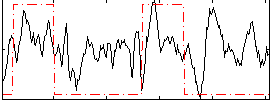 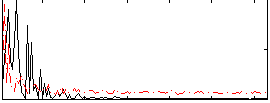 |
Correlation: 0.24828
  |
Subject: 64 |
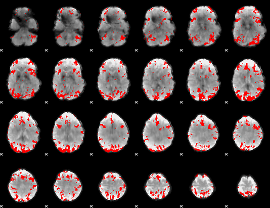
GLM result cope: 01, threshold: 3
 |
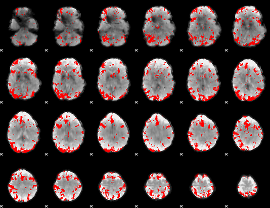
GLM result cope: 02, threshold: 3
 |
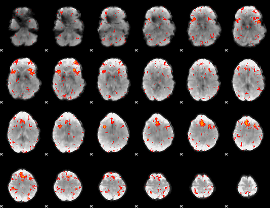
HAFNI result component: 278
Overlap rate: 0.33212
 |
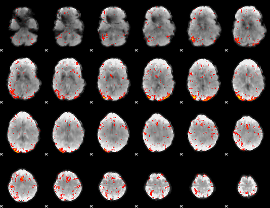
HAFNI result component: 397
Overlap rate: 0.36026
 |
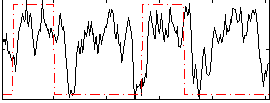
Correlation: 0.23549
  |
Correlation: 0.32174
  |
Subject: 65 |
GLM result cope: 01, threshold: 2.5
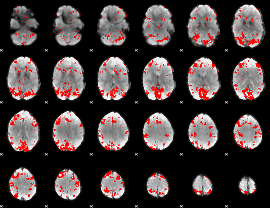 |
GLM result cope: 02, threshold: 2.5
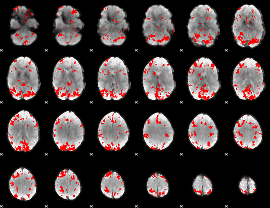 |
HAFNI result component: 263
Overlap rate: 0.37506
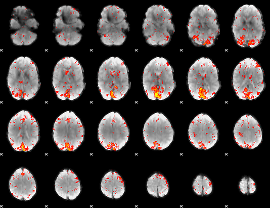 |
HAFNI result component: 394
Overlap rate: 0.33019
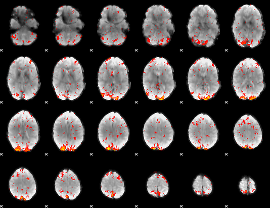 |
Correlation: 0.26962
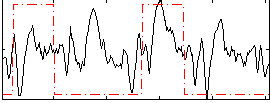 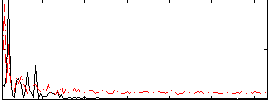 |
Correlation: 0.2164
  |
Subject: 66 |
GLM result cope: 01, threshold: 3
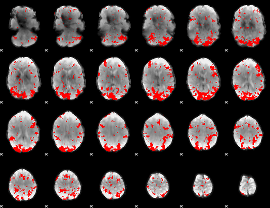 |
GLM result cope: 02, threshold: 3
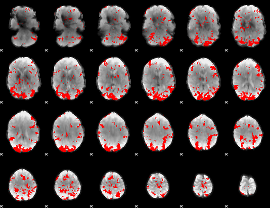 |
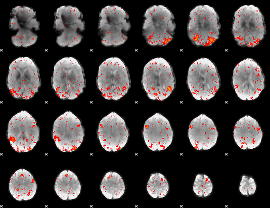
HAFNI result component: 172
Overlap rate: 0.31976
 |
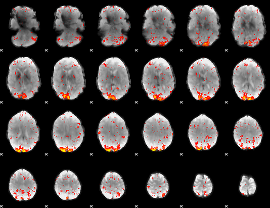
HAFNI result component: 252
Overlap rate: 0.47897
 |
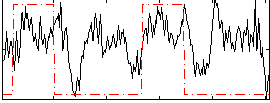
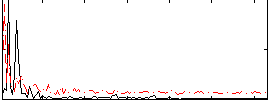
Correlation: 0.32363
  |
Correlation: 0.24451
  |
Subject: 67 |
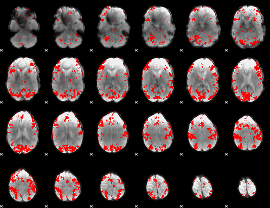
GLM result cope: 01, threshold: 3
 |
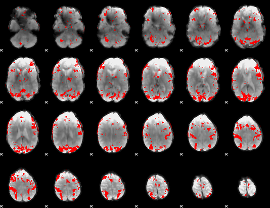
GLM result cope: 02, threshold: 3
 |
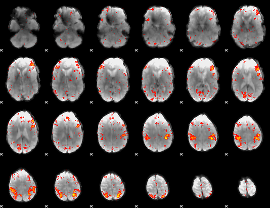
HAFNI result component: 360
Overlap rate: 0.32862
 |
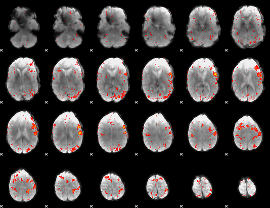
HAFNI result component: 211
Overlap rate: 0.36513
 |
Correlation: 0.056337
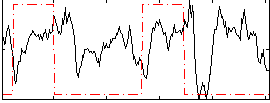 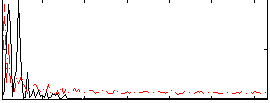 |
Correlation: 0.24676
  |
Subject: 68 |
GLM result cope: 01, threshold: 2.5
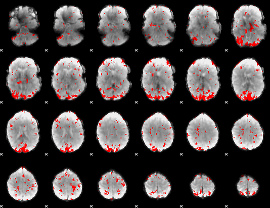 |
GLM result cope: 02, threshold: 3
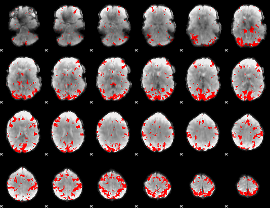 |
HAFNI result component: 194
Overlap rate: 0.38814
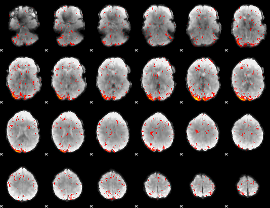 |
HAFNI result component: 285
Overlap rate: 0.44416
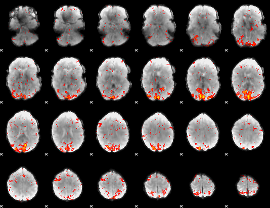 |
Correlation: 0.21285
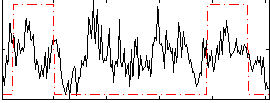 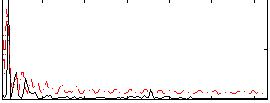 |
Correlation: 0.36567
  |































































































































































































































































































































































































































































































































































