HAFNI report on finalized components on task: EMOTION |
Totally 3 copes finalized |
Subject: 1 |
GLM result cope: 01, threshold: 2.5
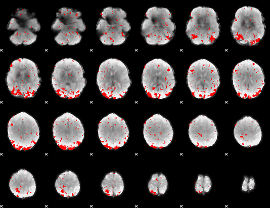 |
GLM result cope: 02, threshold: 3
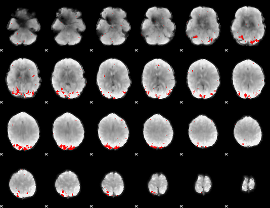 |
GLM result cope: 03, threshold: 3.5
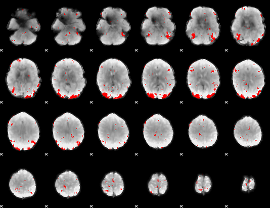 |
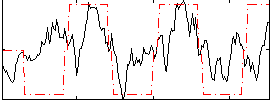
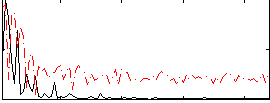
HAFNI result component: 312
Overlap rate: 0.50672
 |
HAFNI result component: 255
Overlap rate: 0.67768
 |
HAFNI result component: 187
Overlap rate: 0.57932
 |
Correlation: 0.61887
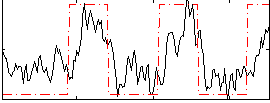 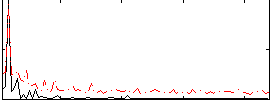 |
Correlation: 0.20321
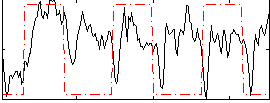  |
Correlation: 0.73571
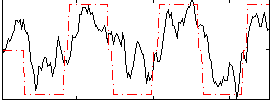 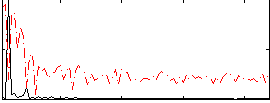 |
Subject: 2 |
GLM result cope: 01, threshold: 3
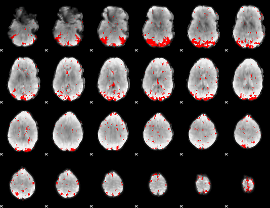 |
GLM result cope: 02, threshold: 2.5
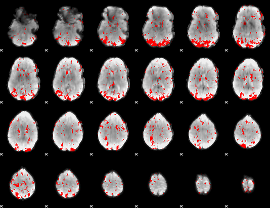 |
GLM result cope: 03, threshold: 3.5
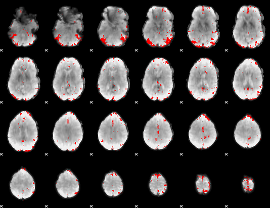 |
HAFNI result component: 100
Overlap rate: 0.50503
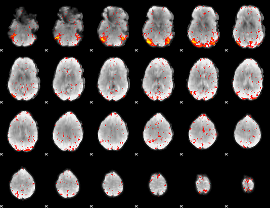 |
HAFNI result component: 095
Overlap rate: 0.6124
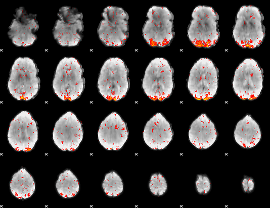 |
HAFNI result component: 100
Overlap rate: 0.49277
 |
Correlation: 0.57097
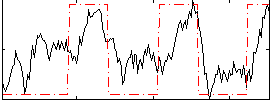 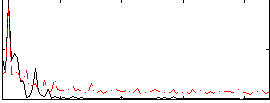 |
Correlation: 0.094151
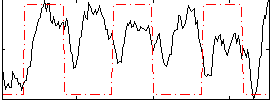  |
Correlation: 0.52781
  |
Subject: 3 |
GLM result cope: 01, threshold: 2.5
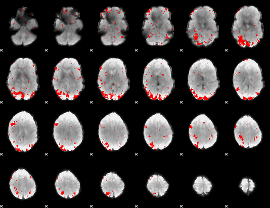 |
GLM result cope: 02, threshold: 2.5
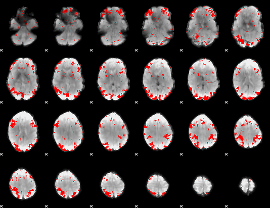 |
GLM result cope: 03, threshold: 2.5
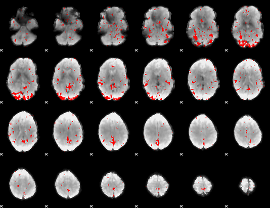 |
HAFNI result component: 159
Overlap rate: 0.55347
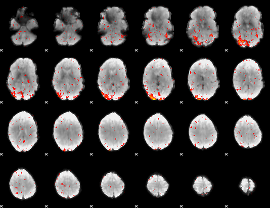 |
HAFNI result component: 139
Overlap rate: 0.22842
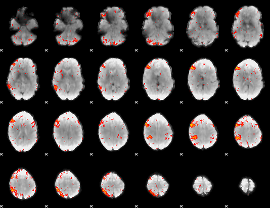 |
HAFNI result component: 159
Overlap rate: 0.49437
 |
Correlation: 0.48261
  |
Correlation: 0.11051
  |
Correlation: 0.42776
  |
Subject: 4 |
GLM result cope: 01, threshold: 3
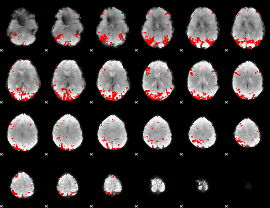 |
GLM result cope: 02, threshold: 3.5
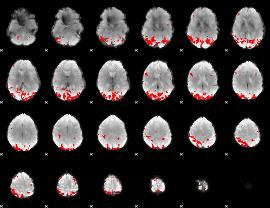 |
GLM result cope: 03, threshold: 3
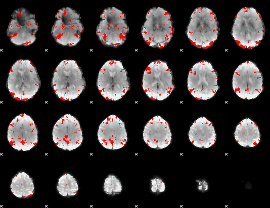 |
HAFNI result component: 089
Overlap rate: 0.43522
 |
HAFNI result component: 332
Overlap rate: 0.35639
 |
HAFNI result component: 089
Overlap rate: 0.48716
 |
Correlation: 0.57654
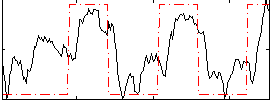 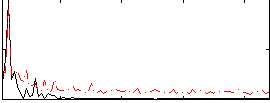 |
Correlation: 0.44855
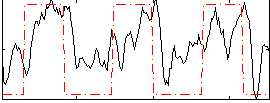  |
Correlation: 0.56809
  |
Subject: 5 |
GLM result cope: 01, threshold: 3
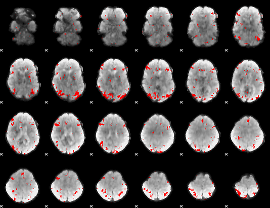 |
GLM result cope: 02, threshold: 2
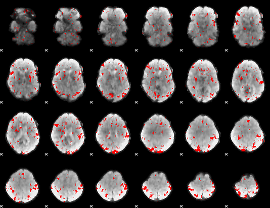 |
GLM result cope: 03, threshold: 2.5
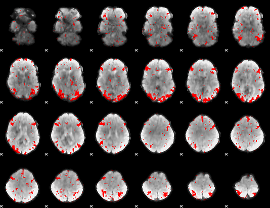 |
HAFNI result component: 021
Overlap rate: 0.65978
 |
HAFNI result component: 216
Overlap rate: 0.6294
 |
HAFNI result component: 021
Overlap rate: 0.72296
 |
Correlation: 0.5493
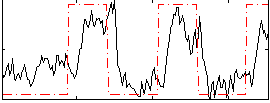 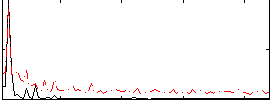 |
Correlation: 0.037515
  |
Correlation: 0.58268
  |
Subject: 6 |
GLM result cope: 01, threshold: 2.5
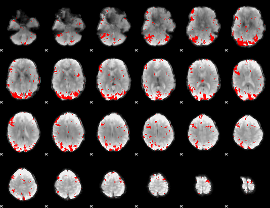 |
GLM result cope: 02, threshold: 2.5
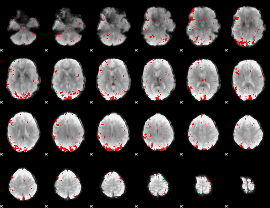 |
GLM result cope: 03, threshold: 3
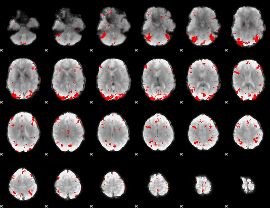 |
HAFNI result component: 381
Overlap rate: 0.43422
 |
HAFNI result component: 169
Overlap rate: 0.67053
 |
HAFNI result component: 381
Overlap rate: 0.689
 |
Correlation: 0.55287
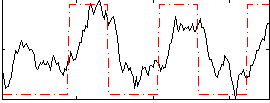 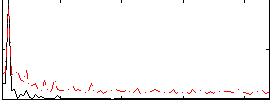 |
Correlation: 0.43044
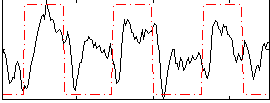 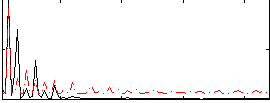 |
Correlation: 0.54973
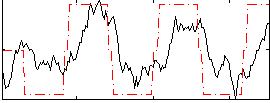  |
Subject: 7 |
GLM result cope: 01, threshold: 3
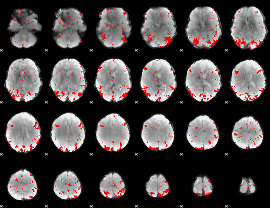 |
GLM result cope: 02, threshold: 2
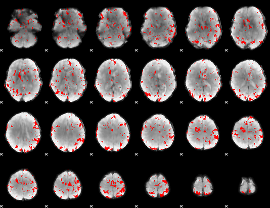 |
GLM result cope: 03, threshold: 3.5
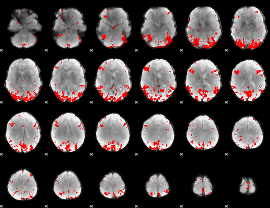 |
HAFNI result component: 329
Overlap rate: 0.46399
 |
HAFNI result component: 257
Overlap rate: 0.35188
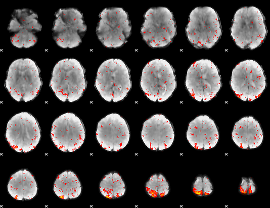 |
HAFNI result component: 393
Overlap rate: 0.30541
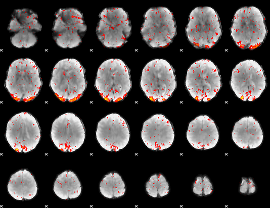 |
Correlation: 0.32479
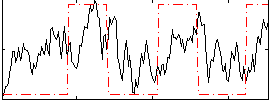 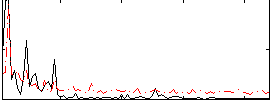 |
Correlation: 0.26631
  |
Correlation: 0.56626
  |
Subject: 8 |
GLM result cope: 01, threshold: 1.5
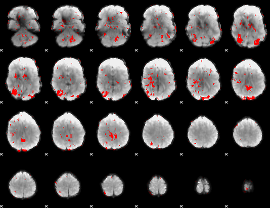 |
GLM result cope: 02, threshold: 1.5
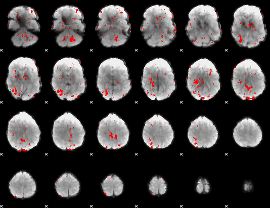 |
GLM result cope: 03, threshold: 3
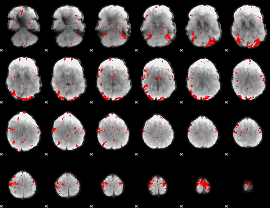 |
HAFNI result component: 365
Overlap rate: 0.8384
 |
HAFNI result component: 126
Overlap rate: 0.13809
 |
HAFNI result component: 365
Overlap rate: 0.58426
 |
Correlation: 0.54408
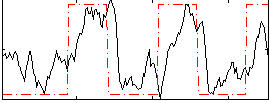  |
Correlation: 0.22557
  |
Correlation: 0.59843
  |
Subject: 9 |
GLM result cope: 01, threshold: 2.5
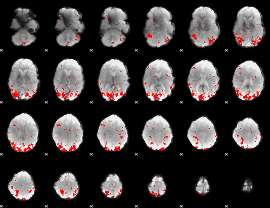 |
GLM result cope: 02, threshold: 2.5
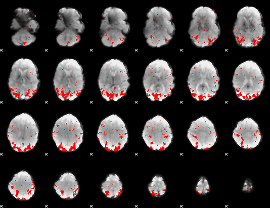 |
GLM result cope: 03, threshold: 2
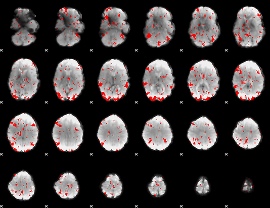 |
HAFNI result component: 053
Overlap rate: 0.52894
 |
HAFNI result component: 265
Overlap rate: 0.61948
 |
HAFNI result component: 058
Overlap rate: 0.64113
 |
Correlation: 0.40497
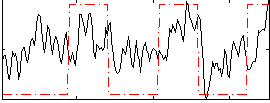 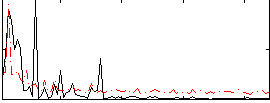 |
Correlation: 0.2606
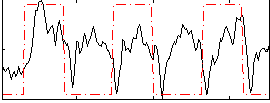 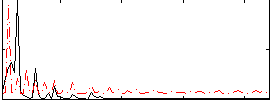 |
Correlation: 0.52616
  |
Subject: 10 |
GLM result cope: 01, threshold: 2
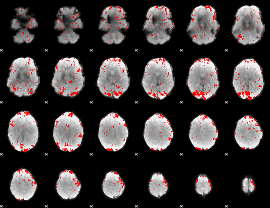 |
GLM result cope: 02, threshold: 2.5
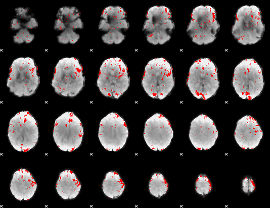 |
GLM result cope: 03, threshold: 3
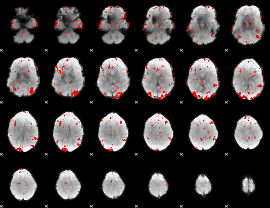 |
HAFNI result component: 030
Overlap rate: 0.55391
 |
HAFNI result component: 357
Overlap rate: 0.66398
 |
HAFNI result component: 030
Overlap rate: 0.40489
 |
Correlation: 0.48987
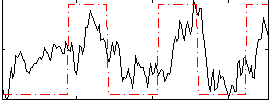 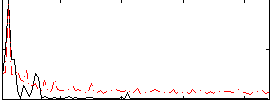 |
Correlation: 0.25366
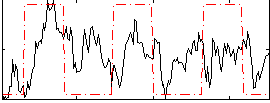  |
Correlation: 0.48754
  |
Subject: 11 |
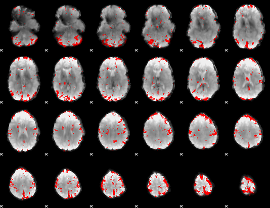
GLM result cope: 01, threshold: 3
 |
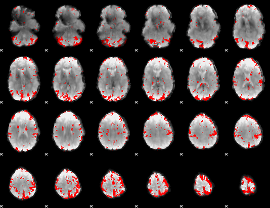
GLM result cope: 02, threshold: 3
 |
GLM result cope: 03, threshold: 2
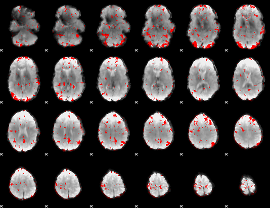 |
HAFNI result component: 344
Overlap rate: 0.2153
 |
HAFNI result component: 215
Overlap rate: 0.5257
 |
HAFNI result component: 191
Overlap rate: 0.47666
 |
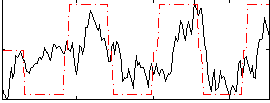
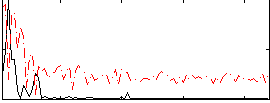
Correlation: 0.59866
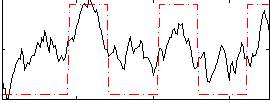 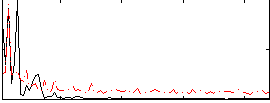 |
Correlation: 0.27356
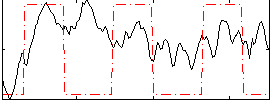  |
Correlation: 0.5746
  |
Subject: 12 |
GLM result cope: 01, threshold: 2
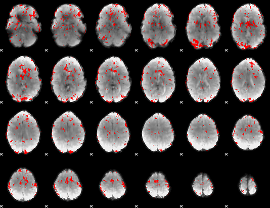 |
GLM result cope: 02, threshold: 2
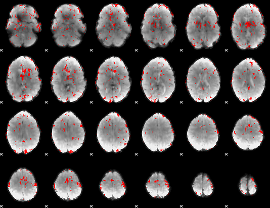 |
GLM result cope: 03, threshold: 2.5
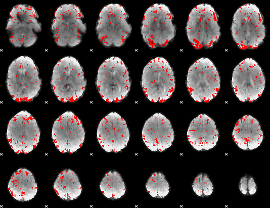 |
HAFNI result component: 075
Overlap rate: 0.43758
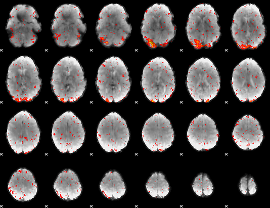 |
HAFNI result component: 226
Overlap rate: 0.25258
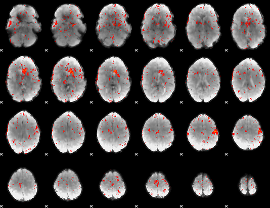 |
HAFNI result component: 075
Overlap rate: 0.55556
 |
Correlation: 0.52301
  |
Correlation: 0.27465
  |
Correlation: 0.51911
  |
Subject: 13 |
GLM result cope: 01, threshold: 2
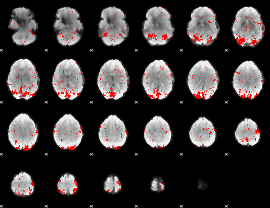 |
GLM result cope: 02, threshold: 2.5
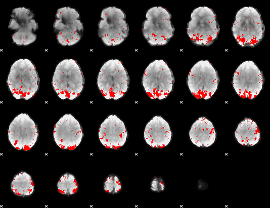 |
GLM result cope: 03, threshold: 2
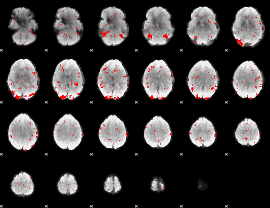 |
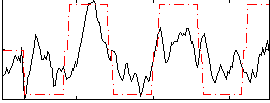
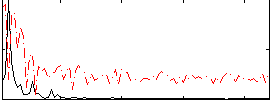
HAFNI result component: 257
Overlap rate: 0.62734
 |
HAFNI result component: 282
Overlap rate: 0.71148
 |
HAFNI result component: 084
Overlap rate: 0.74548
 |
Correlation: 0.24905
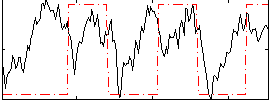 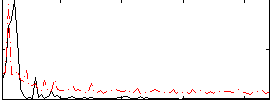 |
Correlation: 0.41062
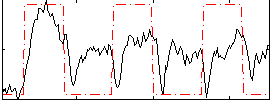  |
Correlation: 0.63957
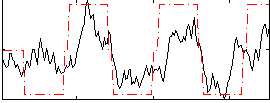 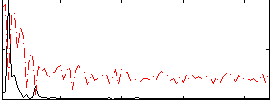 |
Subject: 14 |
GLM result cope: 01, threshold: 3
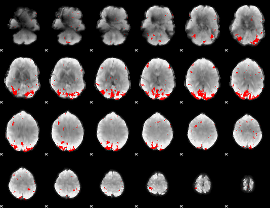 |
GLM result cope: 02, threshold: 2.5
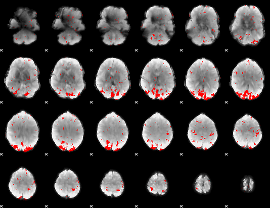 |
GLM result cope: 03, threshold: 3
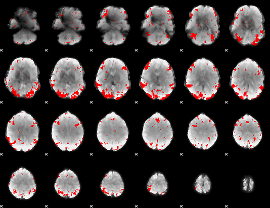 |
HAFNI result component: 026
Overlap rate: 0.6907
 |
HAFNI result component: 350
Overlap rate: 0.61808
 |
HAFNI result component: 374
Overlap rate: 0.43715
 |
Correlation: 0.35705
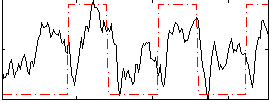 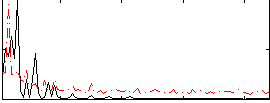 |
Correlation: 0.41166
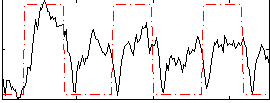 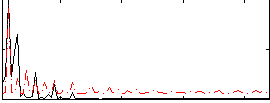 |
Correlation: 0.47676
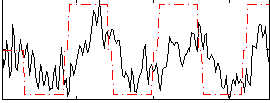 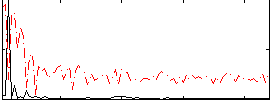 |
Subject: 15 |
GLM result cope: 01, threshold: 2.5
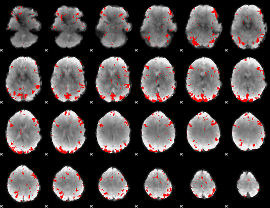 |
GLM result cope: 02, threshold: 1.5
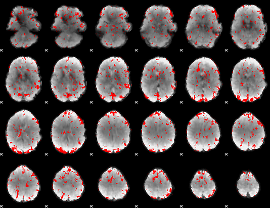 |
GLM result cope: 03, threshold: 3
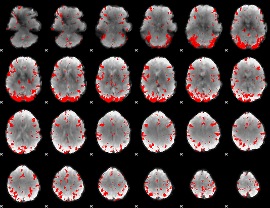 |
HAFNI result component: 030
Overlap rate: 0.39808
 |
HAFNI result component: 064
Overlap rate: 0.65993
 |
HAFNI result component: 029
Overlap rate: 0.53069
 |
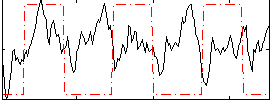
Correlation: 0.63768
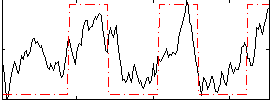 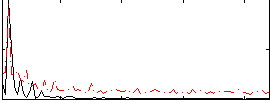 |
Correlation: 0.016048
 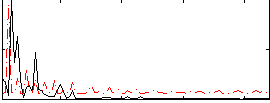 |
Correlation: 0.60925
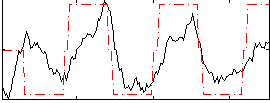 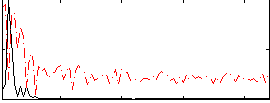 |
Subject: 16 |
GLM result cope: 01, threshold: 3
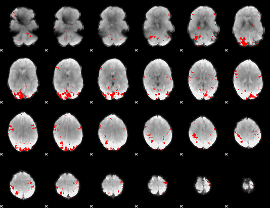 |
GLM result cope: 02, threshold: 2.5
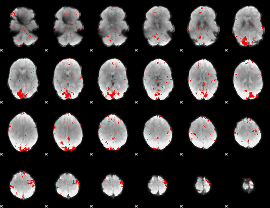 |
GLM result cope: 03, threshold: 3
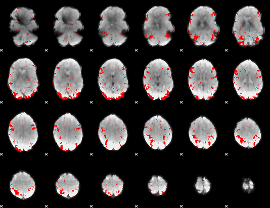 |
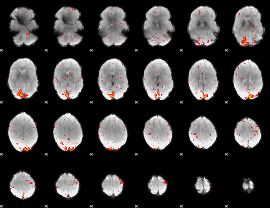
HAFNI result component: 083
Overlap rate: 0.36552
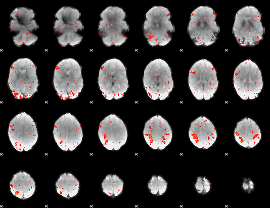 |
HAFNI result component: 045
Overlap rate: 0.83345
 |
HAFNI result component: 141
Overlap rate: 0.46178
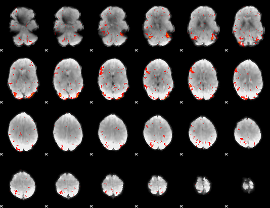 |
Correlation: 0.30827
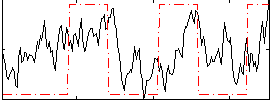 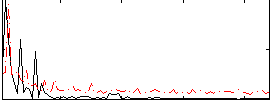 |
Correlation: 0.32708
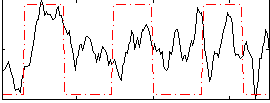  |
Correlation: 0.55757
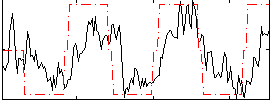 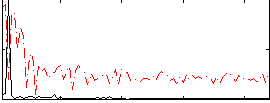 |
Subject: 17 |
GLM result cope: 01, threshold: 0
 |
GLM result cope: 02, threshold: 0
 |
GLM result cope: 03, threshold: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
HAFNI result component: 000
Overlap rate: 0
 |
Correlation: NaN
  |
Correlation: NaN
  |
Correlation: NaN
  |
Subject: 18 |
GLM result cope: 01, threshold: 3
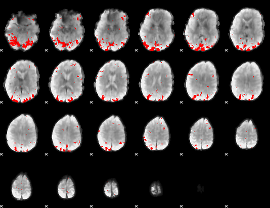 |
GLM result cope: 02, threshold: 2.5
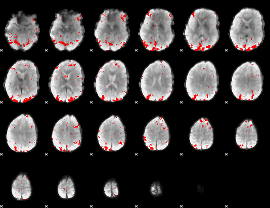 |
GLM result cope: 03, threshold: 3.5
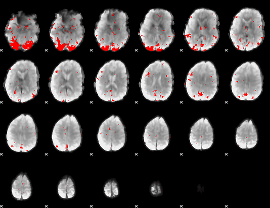 |
HAFNI result component: 371
Overlap rate: 0.4262
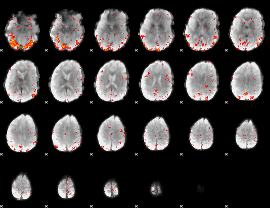 |
HAFNI result component: 209
Overlap rate: 0.55058
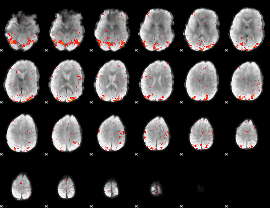 |
HAFNI result component: 371
Overlap rate: 0.56369
 |
Correlation: 0.66399
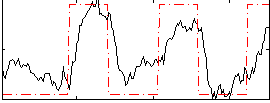 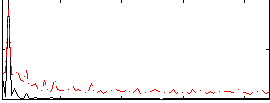 |
Correlation: -0.060801
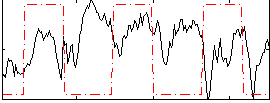  |
Correlation: 0.7021
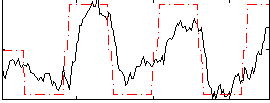  |
Subject: 19 |
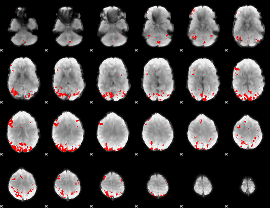
GLM result cope: 01, threshold: 3.5
 |
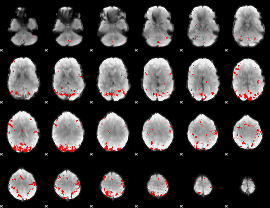
GLM result cope: 02, threshold: 3.5
 |
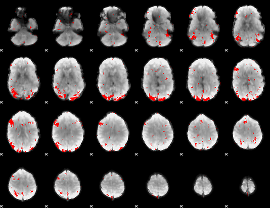
GLM result cope: 03, threshold: 2.5
 |
HAFNI result component: 337
Overlap rate: 0.37594
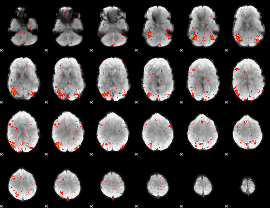 |
HAFNI result component: 262
Overlap rate: 0.44094
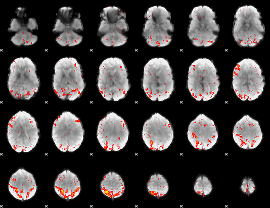 |
HAFNI result component: 043
Overlap rate: 0.66818
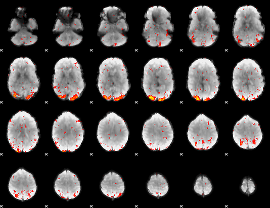 |
Correlation: 0.43709
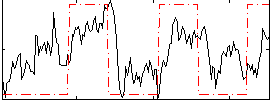 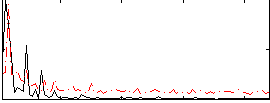 |
Correlation: 0.085114
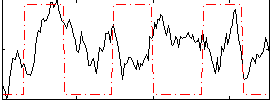 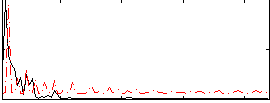 |
Correlation: 0.28371
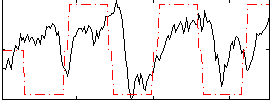  |
Subject: 20 |
GLM result cope: 01, threshold: 3
 |
GLM result cope: 02, threshold: 2.5
 |
GLM result cope: 03, threshold: 3
 |
HAFNI result component: 224
Overlap rate: 0.68035
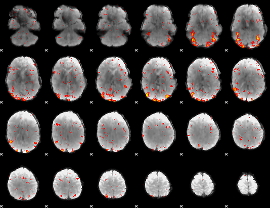 |
HAFNI result component: 103
Overlap rate: 0.666
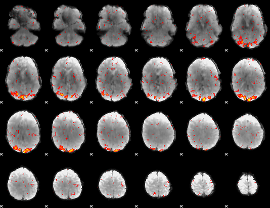 |
HAFNI result component: 224
Overlap rate: 0.59826
 |
Correlation: 0.61023
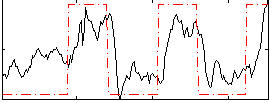 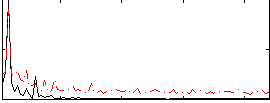 |
Correlation: 0.34939
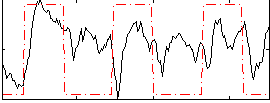  |
Correlation: 0.60106
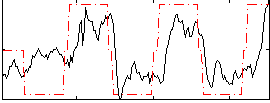 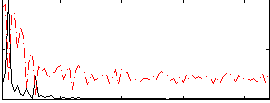 |
Subject: 21 |
GLM result cope: 01, threshold: 2.5
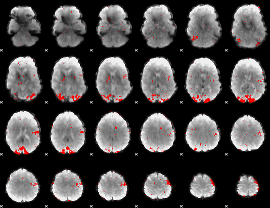 |
GLM result cope: 02, threshold: 2
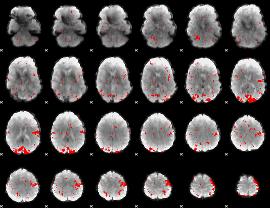 |
GLM result cope: 03, threshold: 2.5
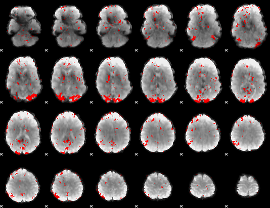 |
HAFNI result component: 236
Overlap rate: 0.69822
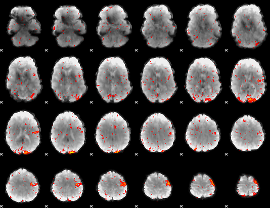 |
HAFNI result component: 236
Overlap rate: 0.93444
 |
HAFNI result component: 188
Overlap rate: 0.56276
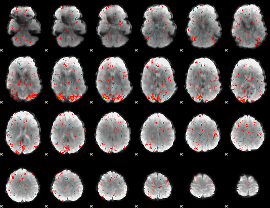 |
Correlation: 0.02886
  |
Correlation: 0.12793
  |
Correlation: 0.50409
  |
Subject: 22 |
GLM result cope: 01, threshold: 2.5
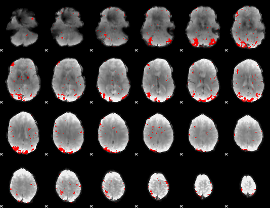 |
GLM result cope: 02, threshold: 2
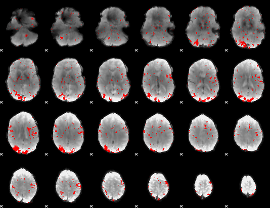 |
GLM result cope: 03, threshold: 2.5
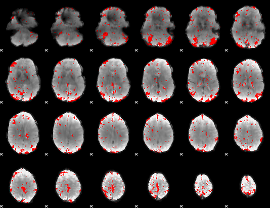 |
HAFNI result component: 259
Overlap rate: 0.38842
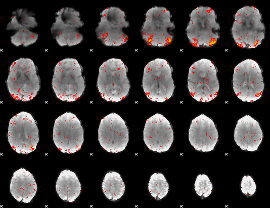 |
HAFNI result component: 193
Overlap rate: 0.69928
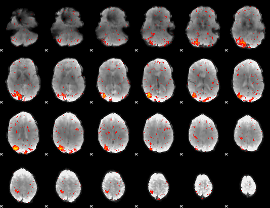 |
HAFNI result component: 259
Overlap rate: 0.63253
 |
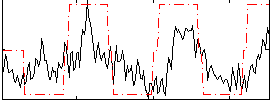
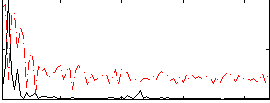
Correlation: 0.64462
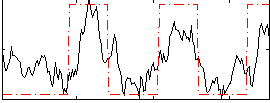  |
Correlation: 0.10786
  |
Correlation: 0.65696
  |
Subject: 23 |
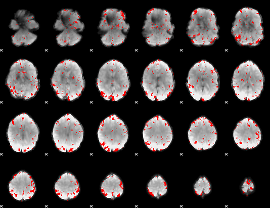
GLM result cope: 01, threshold: 2.5
 |
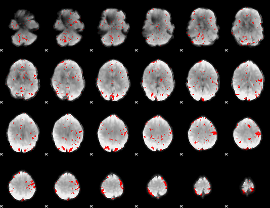
GLM result cope: 02, threshold: 2
 |
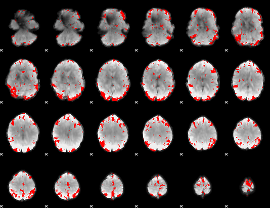
GLM result cope: 03, threshold: 3.5
 |
HAFNI result component: 391
Overlap rate: 0.38609
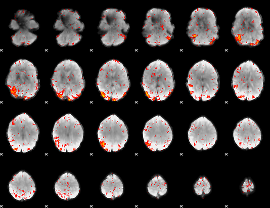 |
HAFNI result component: 191
Overlap rate: 0.31662
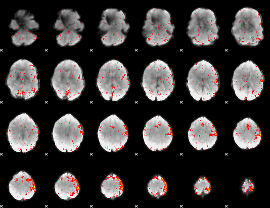 |
HAFNI result component: 391
Overlap rate: 0.44741
 |
Correlation: 0.6529
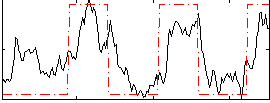 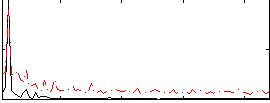 |
Correlation: 0.1474
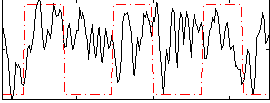 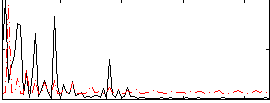 |
Correlation: 0.67503
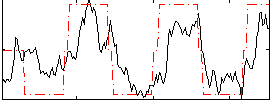 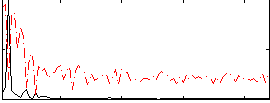 |
Subject: 24 |
GLM result cope: 01, threshold: 2
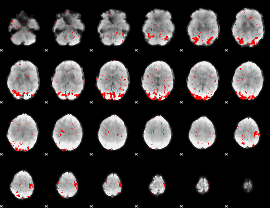 |
GLM result cope: 02, threshold: 2
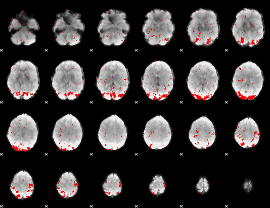 |
GLM result cope: 03, threshold: 2.5
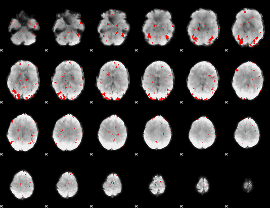 |
HAFNI result component: 179
Overlap rate: 0.63065
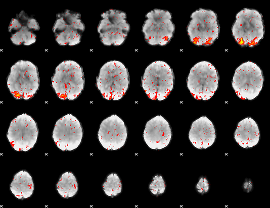 |
HAFNI result component: 235
Overlap rate: 0.81173
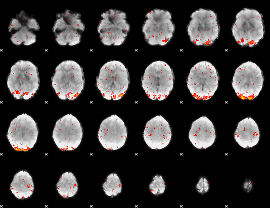 |
HAFNI result component: 179
Overlap rate: 0.54987
 |
Correlation: 0.518
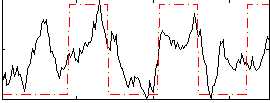 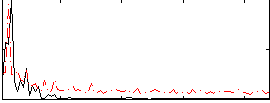 |
Correlation: 0.19181
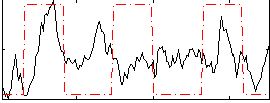  |
Correlation: 0.46881
  |
Subject: 25 |
GLM result cope: 01, threshold: 2.5
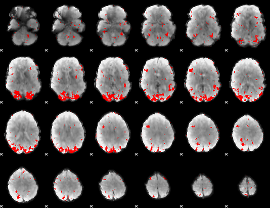 |
GLM result cope: 02, threshold: 2.5
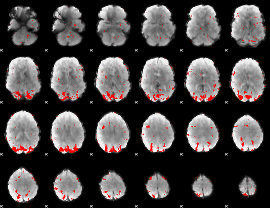 |
GLM result cope: 03, threshold: 2.5
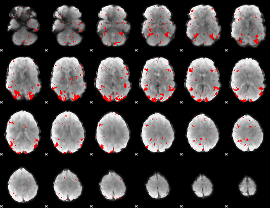 |
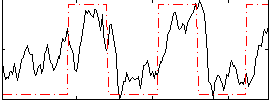
HAFNI result component: 214
Overlap rate: 0.56809
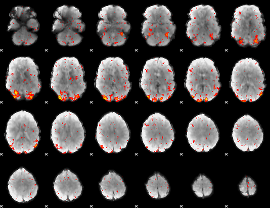 |
HAFNI result component: 239
Overlap rate: 0.68999
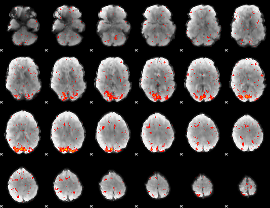 |
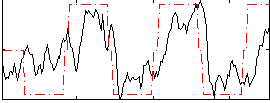
HAFNI result component: 214
Overlap rate: 0.85784
 |
Correlation: 0.55249
 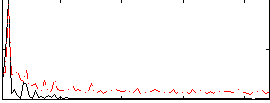 |
Correlation: 0.12719
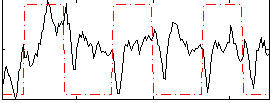  |
Correlation: 0.56179
  |
Subject: 26 |
GLM result cope: 01, threshold: 3
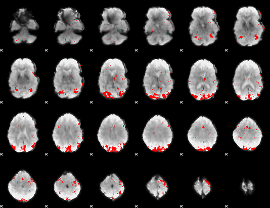 |
GLM result cope: 02, threshold: 3
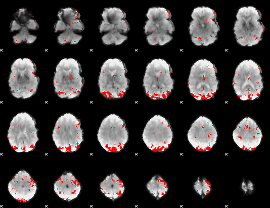 |
GLM result cope: 03, threshold: 2
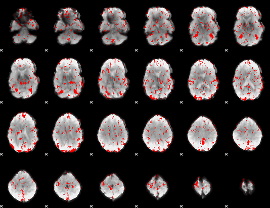 |
HAFNI result component: 246
Overlap rate: 0.43943
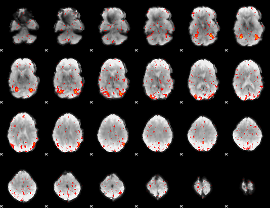 |
HAFNI result component: 060
Overlap rate: 0.60065
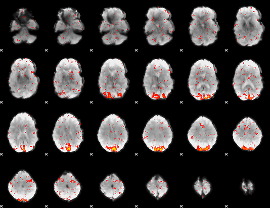 |
HAFNI result component: 246
Overlap rate: 0.68472
 |
Correlation: 0.45232
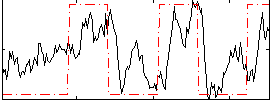 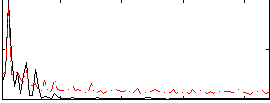 |
Correlation: 0.2106
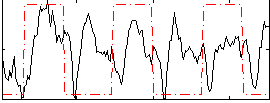 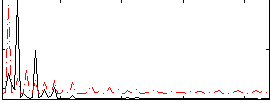 |
Correlation: 0.46228
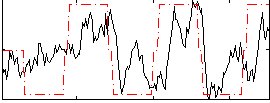 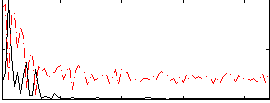 |
Subject: 27 |
GLM result cope: 01, threshold: 2.5
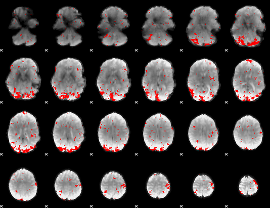 |
GLM result cope: 02, threshold: 2.5
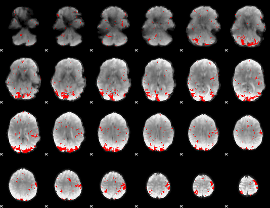 |
GLM result cope: 03, threshold: 2.5
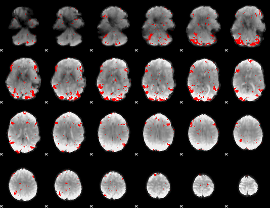 |
HAFNI result component: 287
Overlap rate: 0.54909
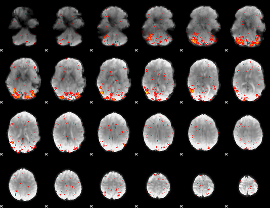 |
HAFNI result component: 256
Overlap rate: 0.68232
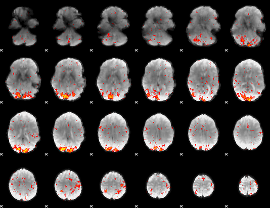 |
HAFNI result component: 337
Overlap rate: 0.57086
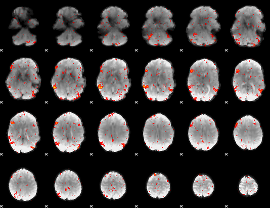 |
Correlation: 0.4858
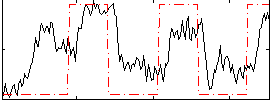 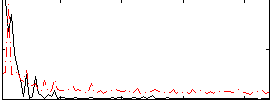 |
Correlation: 0.33523
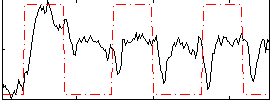 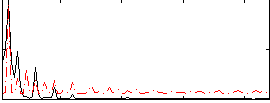 |
Correlation: 0.28976
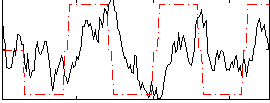 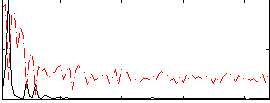 |
Subject: 28 |
GLM result cope: 01, threshold: 2.5
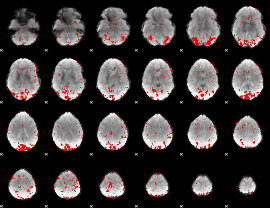 |
GLM result cope: 02, threshold: 2.5
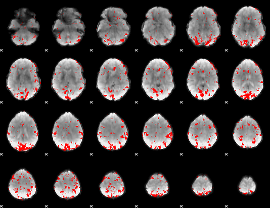 |
GLM result cope: 03, threshold: 2.5
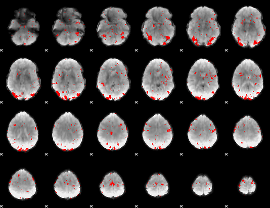 |
HAFNI result component: 048
Overlap rate: 0.65608
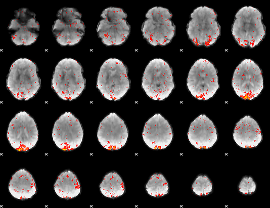 |
HAFNI result component: 390
Overlap rate: 0.38818
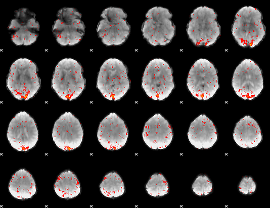 |
HAFNI result component: 185
Overlap rate: 0.81306
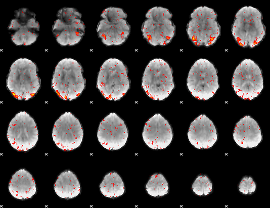 |
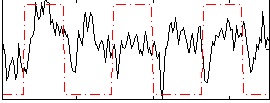
Correlation: 0.36907
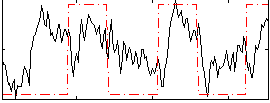 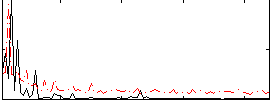 |
Correlation: 0.2045
 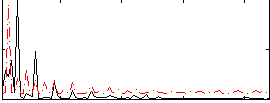 |
Correlation: 0.60584
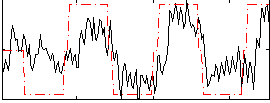 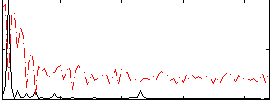 |
Subject: 29 |
GLM result cope: 01, threshold: 2
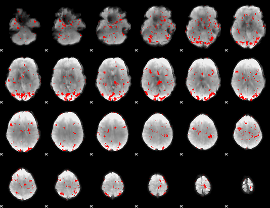 |
GLM result cope: 02, threshold: 2.5
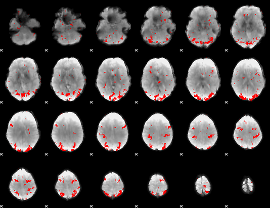 |
GLM result cope: 03, threshold: 1.5
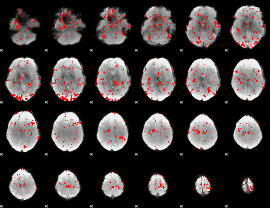 |
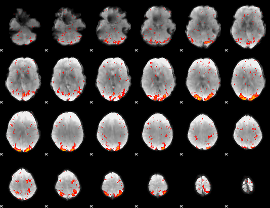
HAFNI result component: 284
Overlap rate: 0.70555
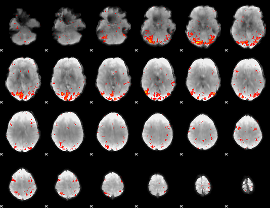 |
HAFNI result component: 034
Overlap rate: 0.84116
 |
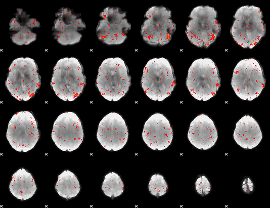
HAFNI result component: 318
Overlap rate: 0.41033
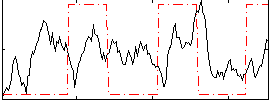
 |
Correlation: 0.11707
 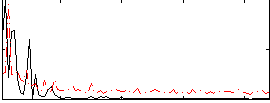 |
Correlation: 0.48503
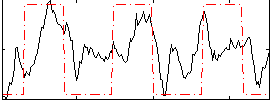 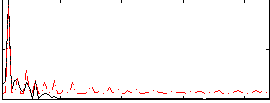 |
Correlation: 0.22954
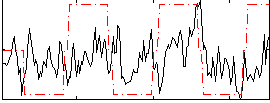 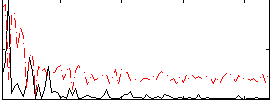 |
Subject: 30 |
GLM result cope: 01, threshold: 3
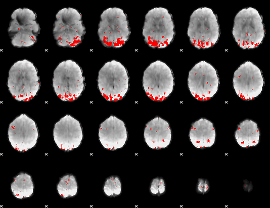 |
GLM result cope: 02, threshold: 2.5
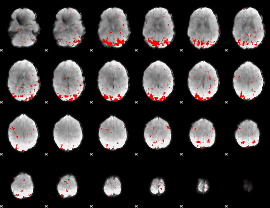 |
GLM result cope: 03, threshold: 2.5
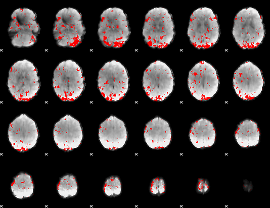 |
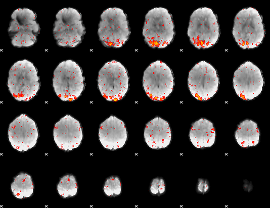
HAFNI result component: 279
Overlap rate: 0.65953
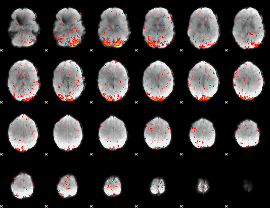 |
HAFNI result component: 178
Overlap rate: 0.7254
 |
HAFNI result component: 048
Overlap rate: 0.58201
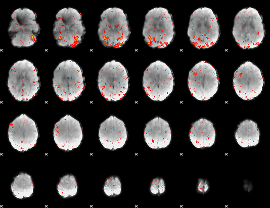 |
Correlation: 0.25402
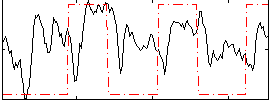 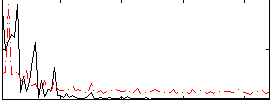 |
Correlation: 0.15888
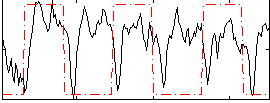 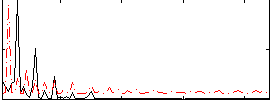 |
Correlation: 0.65319
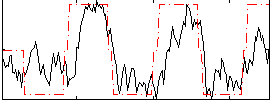 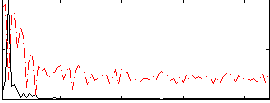 |
Subject: 31 |
GLM result cope: 01, threshold: 2.5
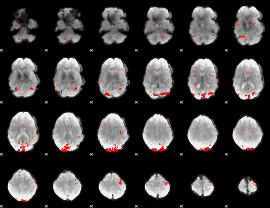 |
GLM result cope: 02, threshold: 1.5
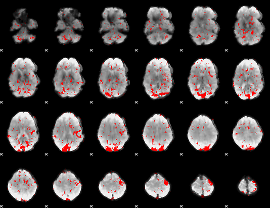 |
GLM result cope: 03, threshold: 2.5
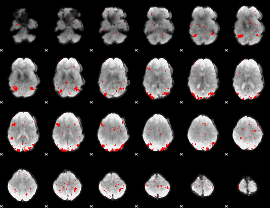 |
HAFNI result component: 136
Overlap rate: 0.67254
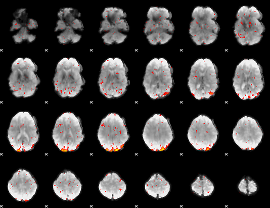 |
HAFNI result component: 024
Overlap rate: 0.75681
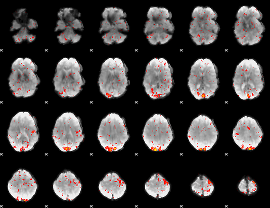 |
HAFNI result component: 379
Overlap rate: 0.63727
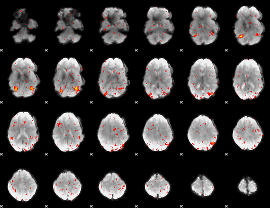 |
Correlation: 0.36041
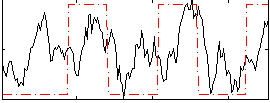 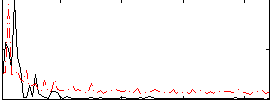 |
Correlation: 0.48375
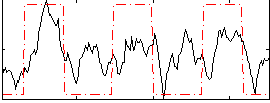 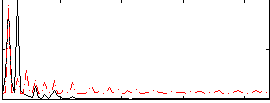 |
Correlation: 0.57112
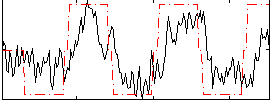 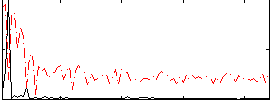 |
Subject: 32 |
GLM result cope: 01, threshold: 3.5
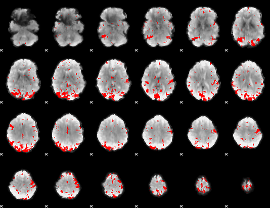 |
GLM result cope: 02, threshold: 3
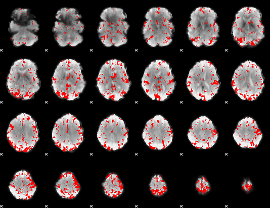 |
GLM result cope: 03, threshold: 2.5
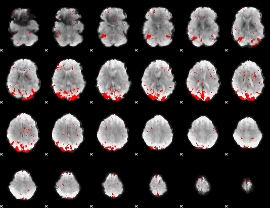 |
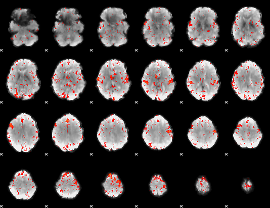
HAFNI result component: 355
Overlap rate: 0.43364
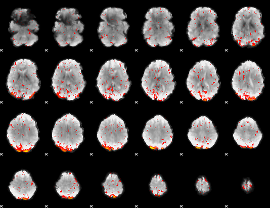 |
HAFNI result component: 069
Overlap rate: 0.3492
 |
HAFNI result component: 310
Overlap rate: 0.56818
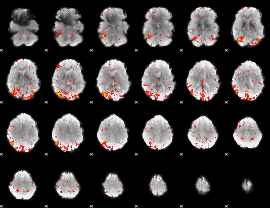 |
Correlation: 0.36533
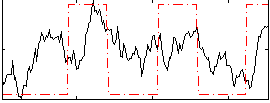 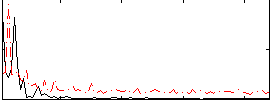 |
Correlation: 0.19874
  |
Correlation: 0.41176
  |
Subject: 33 |
GLM result cope: 01, threshold: 2.5
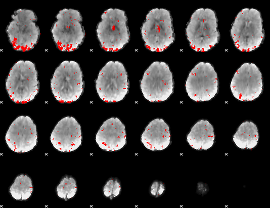 |
GLM result cope: 02, threshold: 2
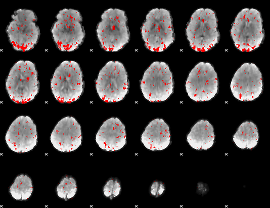 |
GLM result cope: 03, threshold: 2.5
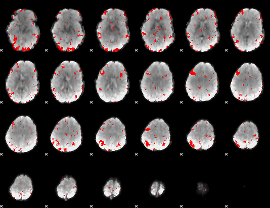 |
HAFNI result component: 297
Overlap rate: 0.73178
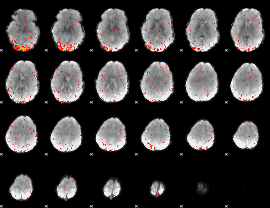 |
HAFNI result component: 275
Overlap rate: 0.62701
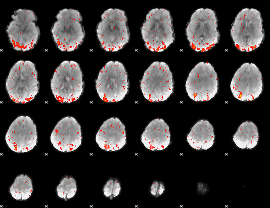 |
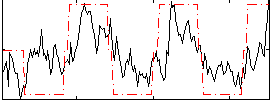
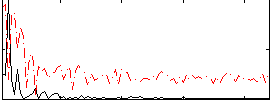
HAFNI result component: 009
Overlap rate: 0.46476
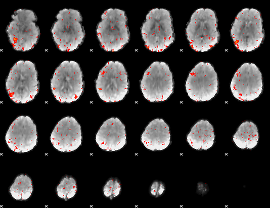 |
Correlation: 0.31691
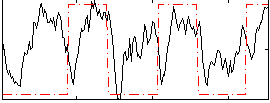 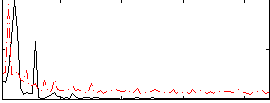 |
Correlation: 0.11544
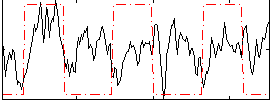  |
Correlation: 0.58669
  |
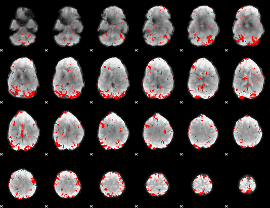
Subject: 34 |
GLM result cope: 01, threshold: 2.5
 |
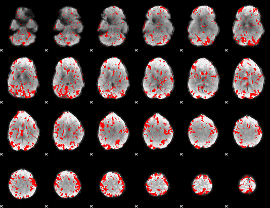
GLM result cope: 02, threshold: 2
 |
GLM result cope: 03, threshold: 2
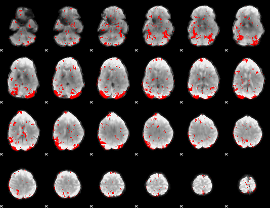 |
HAFNI result component: 018
Overlap rate: 0.66167
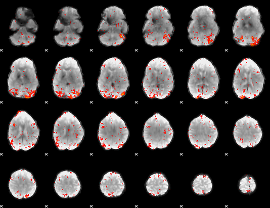 |
HAFNI result component: 162
Overlap rate: 0.28305
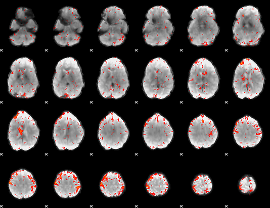 |
HAFNI result component: 380
Overlap rate: 0.75071
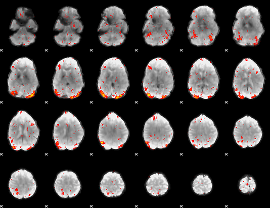 |
Correlation: 0.41119
  |
Correlation: 0.26896
  |
Correlation: 0.35337
  |
Subject: 35 |
GLM result cope: 01, threshold: 2.5
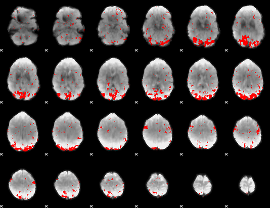 |
GLM result cope: 02, threshold: 3
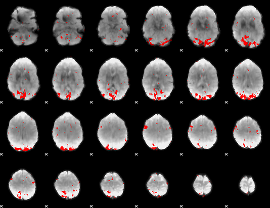 |
GLM result cope: 03, threshold: 3
 |
HAFNI result component: 115
Overlap rate: 0.33791
 |
HAFNI result component: 150
Overlap rate: 0.72861
 |
HAFNI result component: 337
Overlap rate: 0.47446
 |
Correlation: 0.36422
  |
Correlation: 0.16974
  |
Correlation: 0.59464
  |
Subject: 36 |
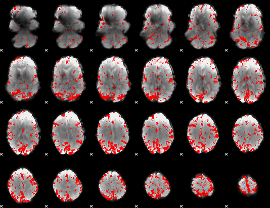
GLM result cope: 01, threshold: 3
 |
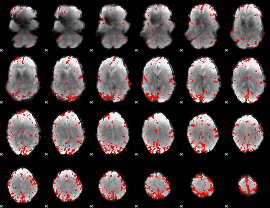
GLM result cope: 02, threshold: 4
 |
GLM result cope: 03, threshold: 3
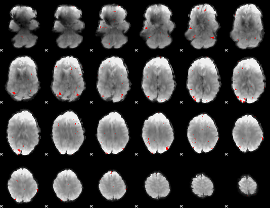 |
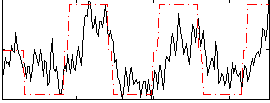
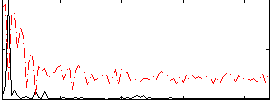
HAFNI result component: 394
Overlap rate: 0.25778
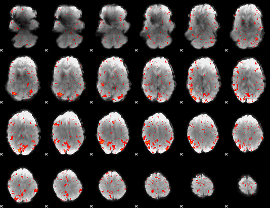 |
HAFNI result component: 244
Overlap rate: 0.23012
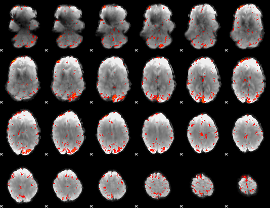 |
HAFNI result component: 017
Overlap rate: 0.38361
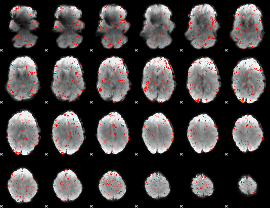 |
Correlation: 0.37554
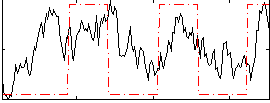 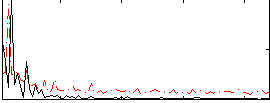 |
Correlation: 0.33705
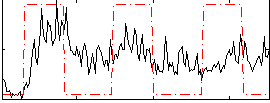 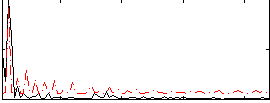 |
Correlation: 0.34496
  |
Subject: 37 |
GLM result cope: 01, threshold: 2.5
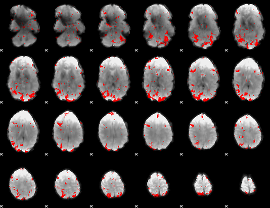 |
GLM result cope: 02, threshold: 2.5
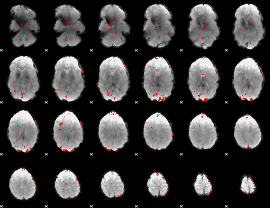 |
GLM result cope: 03, threshold: 4
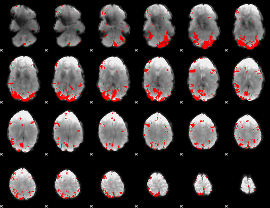 |
HAFNI result component: 191
Overlap rate: 0.48527
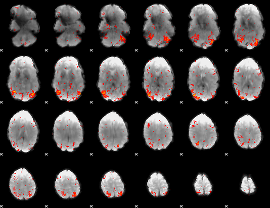 |
HAFNI result component: 182
Overlap rate: 0.743
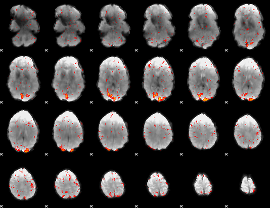 |
HAFNI result component: 377
Overlap rate: 0.40285
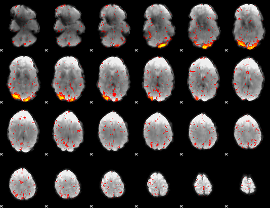 |
Correlation: 0.41238
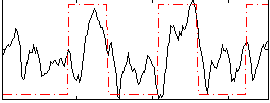 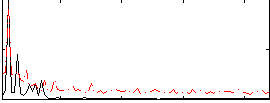 |
Correlation: 0.16619
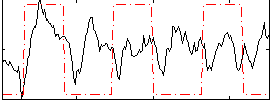 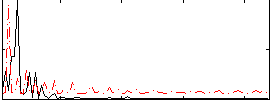 |
Correlation: 0.62017
  |
Subject: 38 |
GLM result cope: 01, threshold: 3
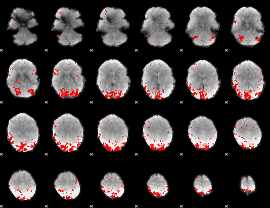 |
GLM result cope: 02, threshold: 3
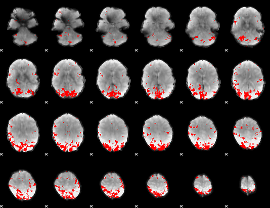 |
GLM result cope: 03, threshold: 3
 |
HAFNI result component: 327
Overlap rate: 0.63214
 |
HAFNI result component: 131
Overlap rate: 0.5377
 |
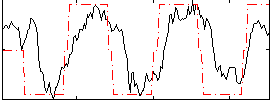
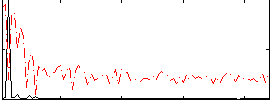
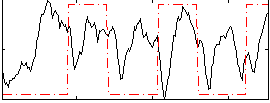
HAFNI result component: 337
Overlap rate: 0.64651
 |
Correlation: 0.11945
 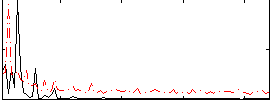 |
Correlation: 0.34884
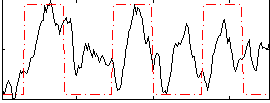  |
Correlation: 0.60881
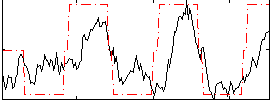  |
Subject: 39 |
GLM result cope: 01, threshold: 3
 |
GLM result cope: 02, threshold: 2
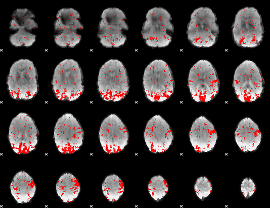 |
GLM result cope: 03, threshold: 3
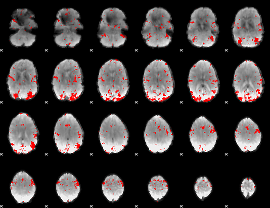 |
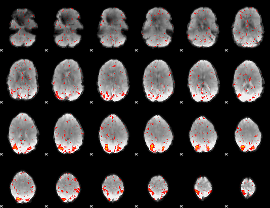
HAFNI result component: 170
Overlap rate: 0.4016
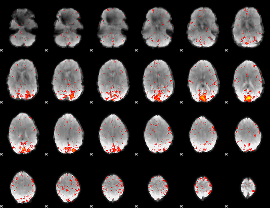 |
HAFNI result component: 203
Overlap rate: 0.20876
 |
HAFNI result component: 103
Overlap rate: 0.71054
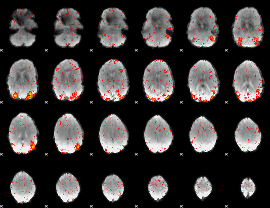 |
Correlation: 0.10799
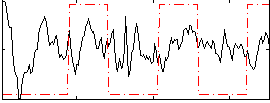 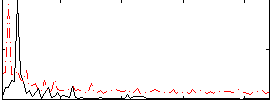 |
Correlation: 0.15118
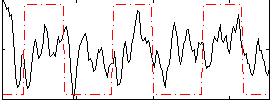  |
Correlation: 0.62085
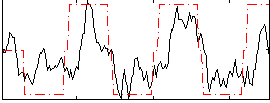 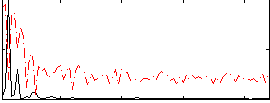 |
Subject: 40 |
GLM result cope: 01, threshold: 2
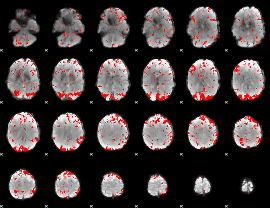 |
GLM result cope: 02, threshold: 2
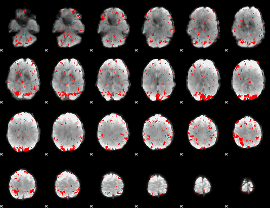 |
GLM result cope: 03, threshold: 2.5
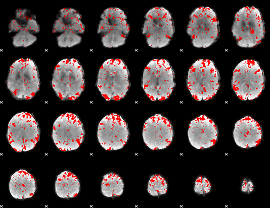 |
HAFNI result component: 013
Overlap rate: 0.71508
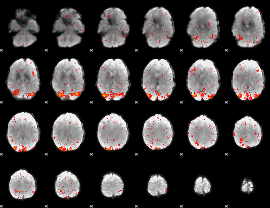 |
HAFNI result component: 287
Overlap rate: 0.69555
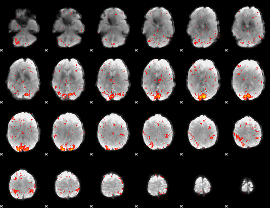 |
HAFNI result component: 390
Overlap rate: 0.36415
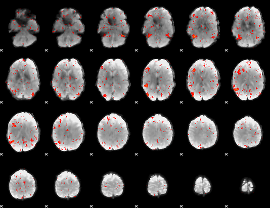 |
Correlation: 0.45045
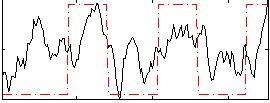 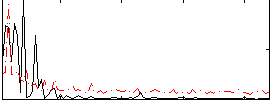 |
Correlation: 0.29808
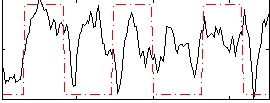  |
Correlation: 0.56736
  |
Subject: 41 |
GLM result cope: 01, threshold: 2
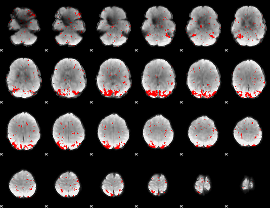 |
GLM result cope: 02, threshold: 2.5
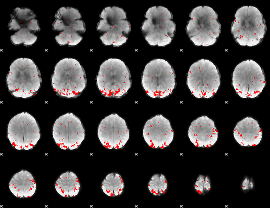 |
GLM result cope: 03, threshold: 2
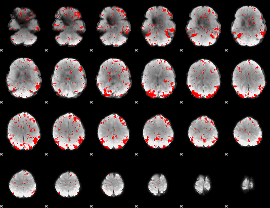 |
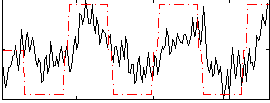
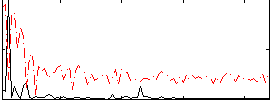
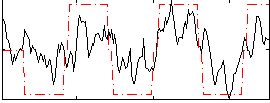
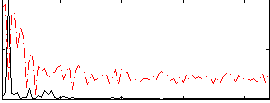
HAFNI result component: 272
Overlap rate: 0.81199
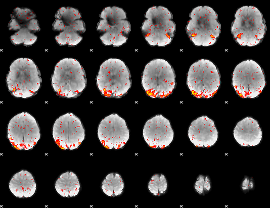 |
HAFNI result component: 246
Overlap rate: 0.6277
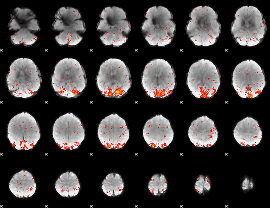 |
HAFNI result component: 355
Overlap rate: 0.52168
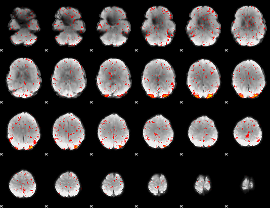 |
Correlation: 0.56328
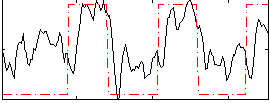 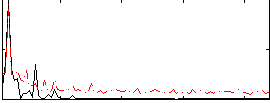 |
Correlation: 0.20048
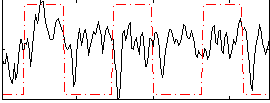 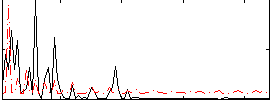 |
Correlation: 0.64532
  |
Subject: 42 |
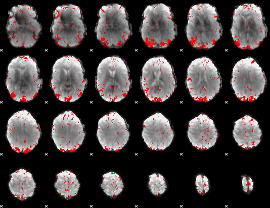
GLM result cope: 01, threshold: 2.5
 |
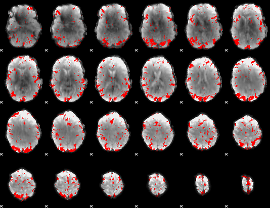
GLM result cope: 02, threshold: 2.5
 |
GLM result cope: 03, threshold: 2.5
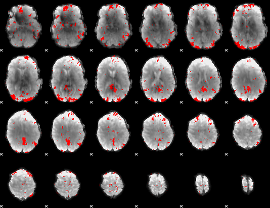 |
HAFNI result component: 055
Overlap rate: 0.34436
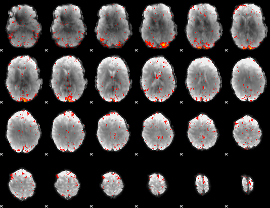 |
HAFNI result component: 110
Overlap rate: 0.33252
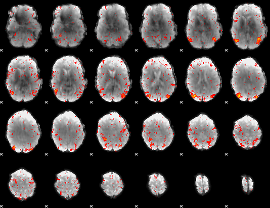 |
HAFNI result component: 173
Overlap rate: 0.54556
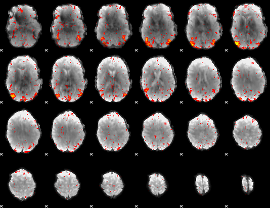 |
Correlation: 0.44672
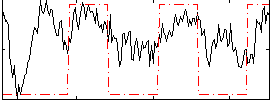 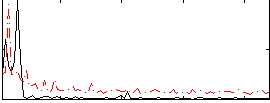 |
Correlation: 0.466
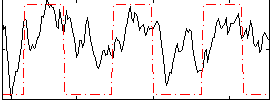  |
Correlation: 0.46239
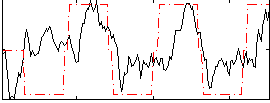  |
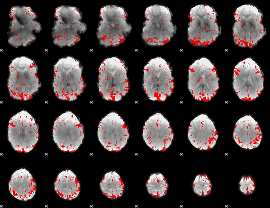
Subject: 43 |
GLM result cope: 01, threshold: 2.5
 |
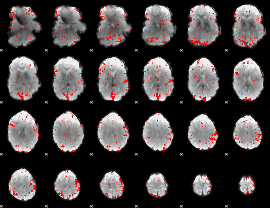
GLM result cope: 02, threshold: 2.5
 |
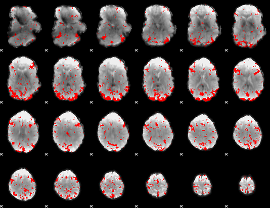
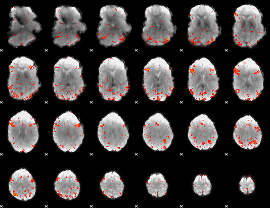
GLM result cope: 03, threshold: 2.5
 |
HAFNI result component: 115
Overlap rate: 0.52058
 |
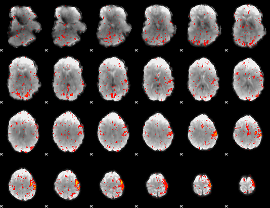
HAFNI result component: 374
Overlap rate: 0.60621
 |
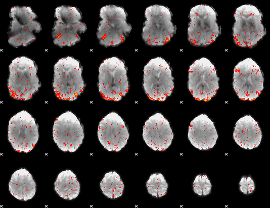
HAFNI result component: 127
Overlap rate: 0.62054
 |
Correlation: 0.41218
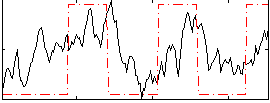 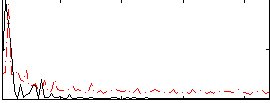 |
Correlation: 0.10992
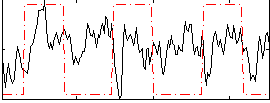 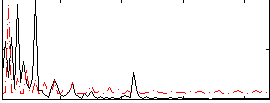 |
Correlation: 0.4525
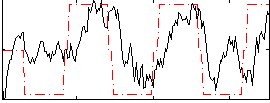  |
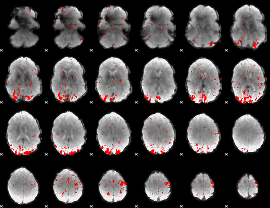
Subject: 44 |
GLM result cope: 01, threshold: 3
 |
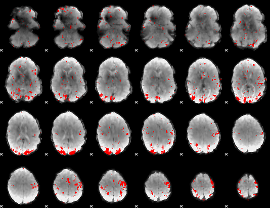
GLM result cope: 02, threshold: 3
 |
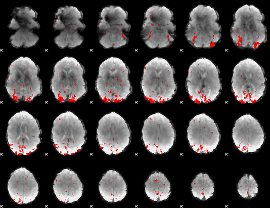
GLM result cope: 03, threshold: 3
 |
HAFNI result component: 062
Overlap rate: 0.39132
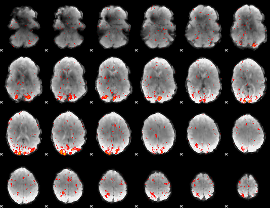 |
HAFNI result component: 285
Overlap rate: 0.35764
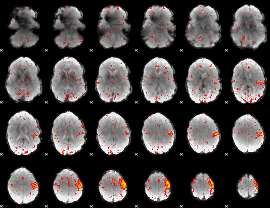 |
HAFNI result component: 030
Overlap rate: 0.72407
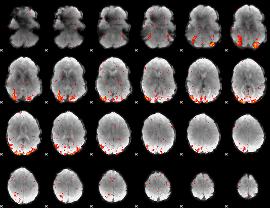 |
Correlation: 0.21817
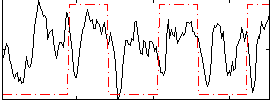 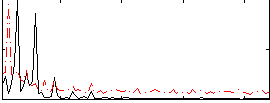 |
Correlation: 0.34069
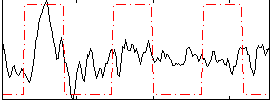  |
Correlation: 0.53236
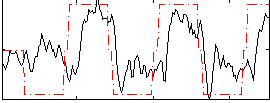 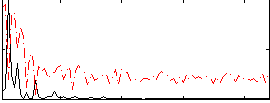 |
Subject: 45 |
GLM result cope: 01, threshold: 3
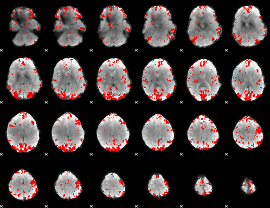 |
GLM result cope: 02, threshold: 2.5
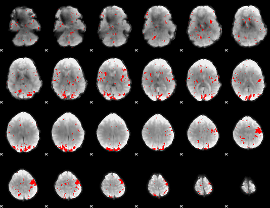 |
GLM result cope: 03, threshold: 3.5
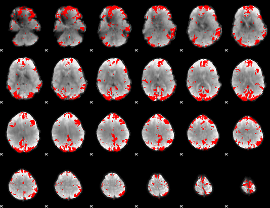 |
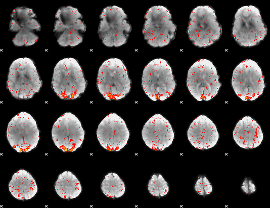
HAFNI result component: 264
Overlap rate: 0.52684
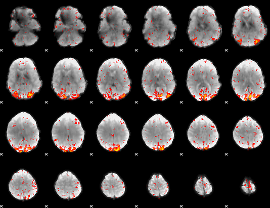 |
HAFNI result component: 101
Overlap rate: 0.58375
 |
HAFNI result component: 361
Overlap rate: 0.22777
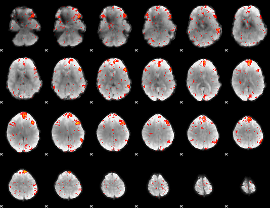 |
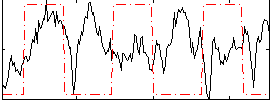
Correlation: 0.54562
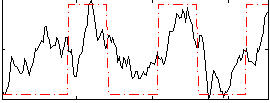 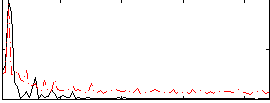 |
Correlation: -0.063309
 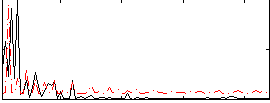 |
Correlation: 0.47107
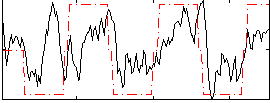 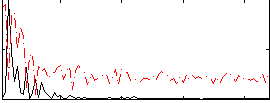 |
Subject: 46 |
GLM result cope: 01, threshold: 2.5
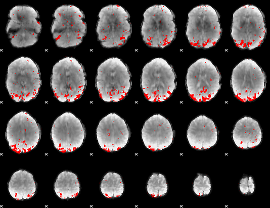 |
GLM result cope: 02, threshold: 2
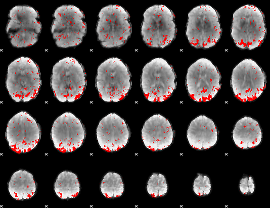 |
GLM result cope: 03, threshold: 2.5
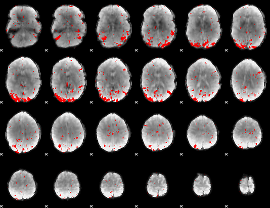 |
HAFNI result component: 388
Overlap rate: 0.5162
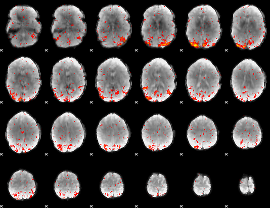 |
HAFNI result component: 144
Overlap rate: 0.44691
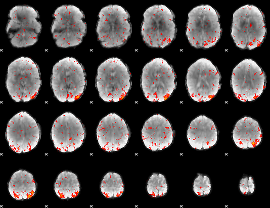 |
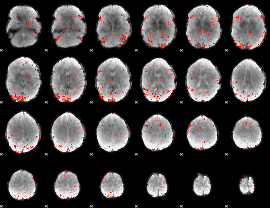
HAFNI result component: 177
Overlap rate: 0.48026
 |
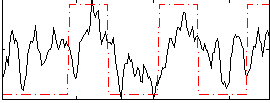
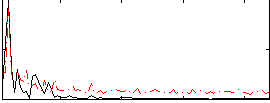
Correlation: 0.4212
  |
Correlation: 0.25502
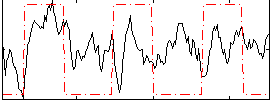  |
Correlation: 0.51655
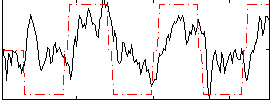 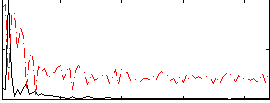 |
Subject: 47 |
GLM result cope: 01, threshold: 2.5
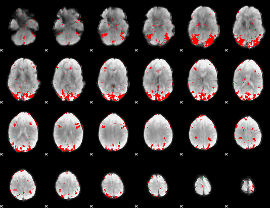 |
GLM result cope: 02, threshold: 2.5
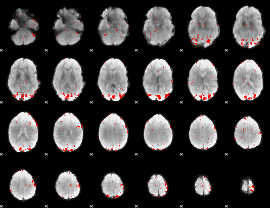 |
GLM result cope: 03, threshold: 2.5
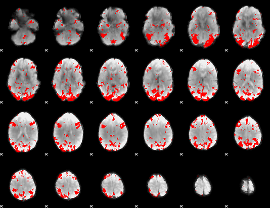 |
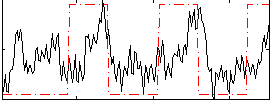
HAFNI result component: 251
Overlap rate: 0.37239
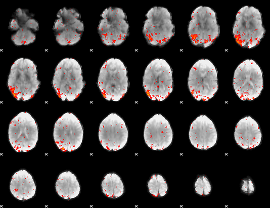 |
HAFNI result component: 325
Overlap rate: 0.37784
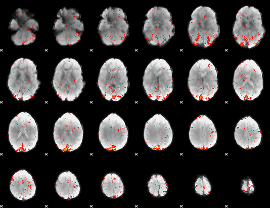 |
HAFNI result component: 275
Overlap rate: 0.33325
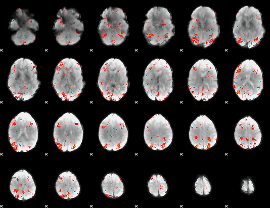 |
Correlation: 0.3204
 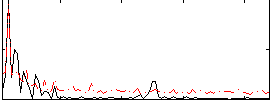 |
Correlation: 0.19629
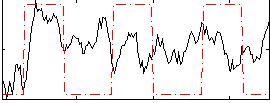  |
Correlation: 0.58727
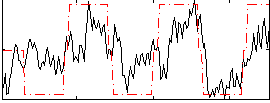 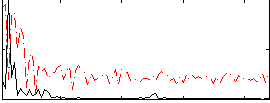 |
Subject: 48 |
GLM result cope: 01, threshold: 2.5
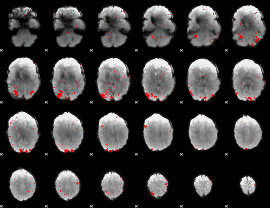 |
GLM result cope: 02, threshold: 2
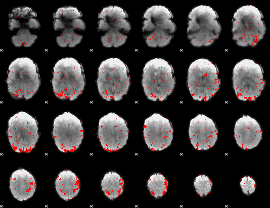 |
GLM result cope: 03, threshold: 2.5
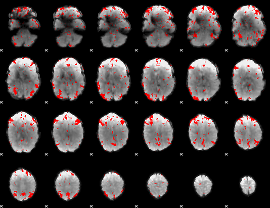 |
HAFNI result component: 361
Overlap rate: 0.51282
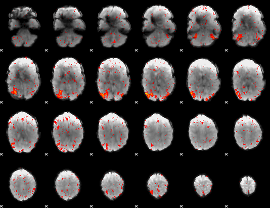 |
HAFNI result component: 013
Overlap rate: 0.40544
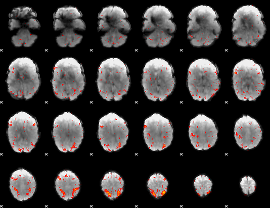 |
HAFNI result component: 116
Overlap rate: 0.30573
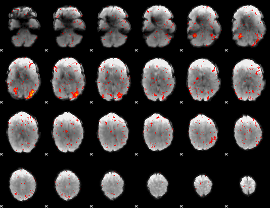 |
Correlation: 0.35178
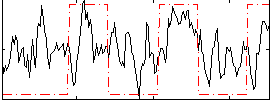 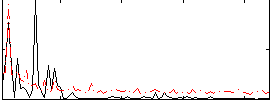 |
Correlation: 0.4153
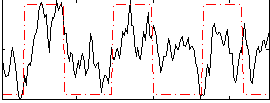  |
Correlation: 0.51434
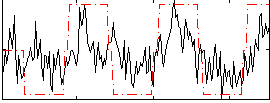  |
Subject: 49 |
GLM result cope: 01, threshold: 3.5
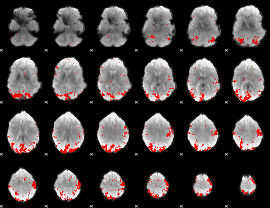 |
GLM result cope: 02, threshold: 3
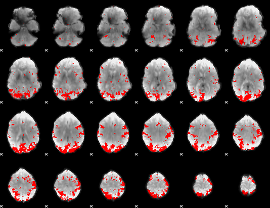 |
GLM result cope: 03, threshold: 3
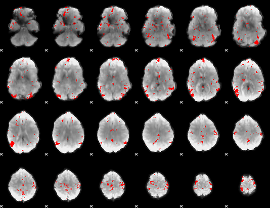 |
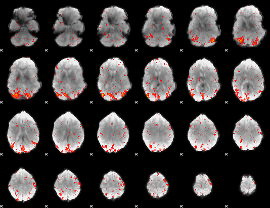
HAFNI result component: 329
Overlap rate: 0.4609
 |
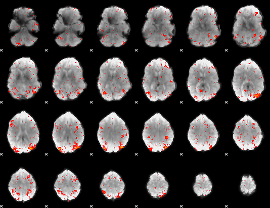
HAFNI result component: 196
Overlap rate: 0.20508
 |
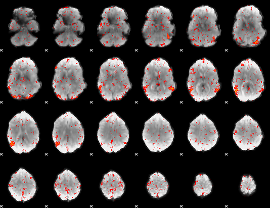
HAFNI result component: 387
Overlap rate: 0.71462
 |
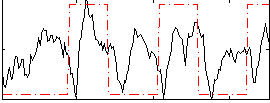
Correlation: 0.22155
 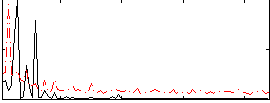 |
Correlation: 0.61732
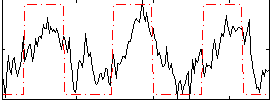 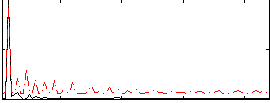 |
Correlation: 0.41508
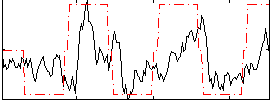  |
Subject: 50 |
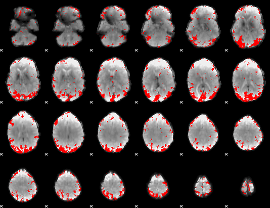
GLM result cope: 01, threshold: 3.5
 |
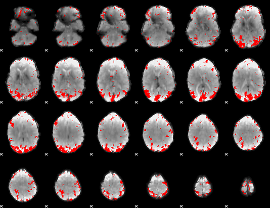
GLM result cope: 02, threshold: 3
 |
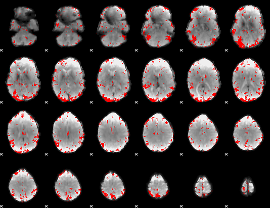
GLM result cope: 03, threshold: 3
 |
HAFNI result component: 155
Overlap rate: 0.39243
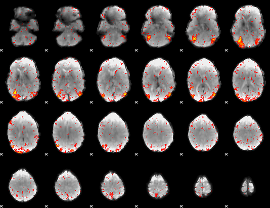 |
HAFNI result component: 232
Overlap rate: 0.48793
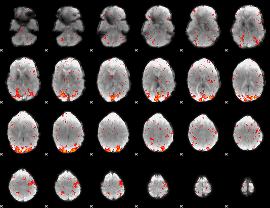 |
HAFNI result component: 155
Overlap rate: 0.54415
 |
Correlation: 0.50784
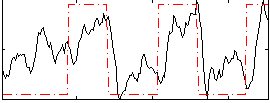 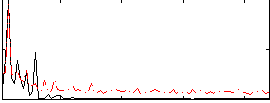 |
Correlation: 0.18665
  |
Correlation: 0.51895
  |
Subject: 51 |
GLM result cope: 01, threshold: 2.5
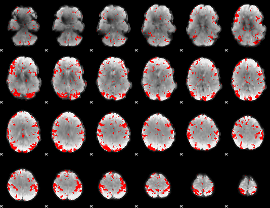 |
GLM result cope: 02, threshold: 2.5
 |
GLM result cope: 03, threshold: 2.5
 |
HAFNI result component: 139
Overlap rate: 0.22557
 |
HAFNI result component: 294
Overlap rate: 0.56458
 |
HAFNI result component: 361
Overlap rate: 0.3372
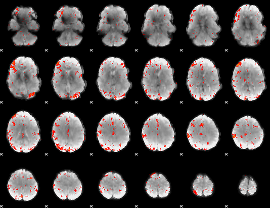 |
Correlation: 0.3279
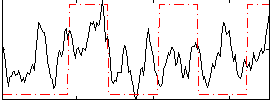 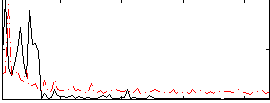 |
Correlation: 0.16369
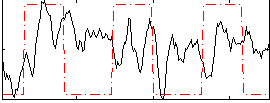  |
Correlation: 0.19693
  |
Subject: 52 |
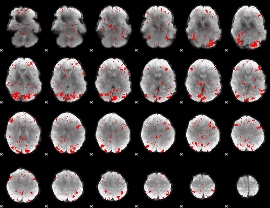
GLM result cope: 01, threshold: 2.5
 |
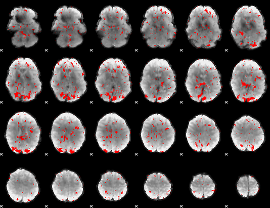
GLM result cope: 02, threshold: 2
 |
GLM result cope: 03, threshold: 3
 |
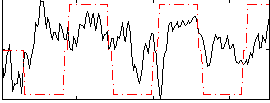
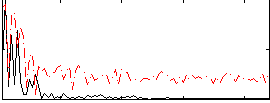
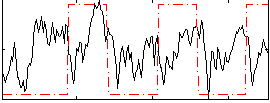
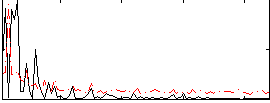
HAFNI result component: 297
Overlap rate: 0.63029
 |
HAFNI result component: 364
Overlap rate: 0.81441
 |
HAFNI result component: 399
Overlap rate: 0.58912
 |
Correlation: 0.12959
  |
Correlation: 0.27609
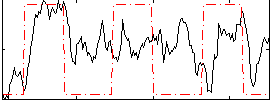 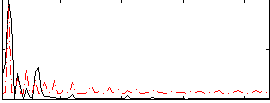 |
Correlation: 0.43211
  |
Subject: 53 |
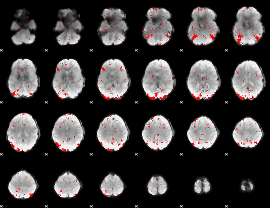
GLM result cope: 01, threshold: 2.5
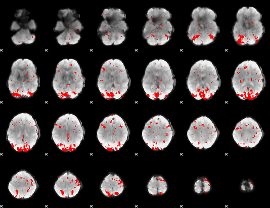 |
GLM result cope: 02, threshold: 2.5
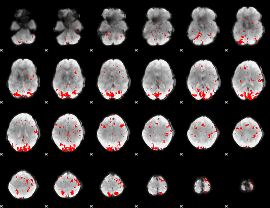 |
GLM result cope: 03, threshold: 2.5
 |
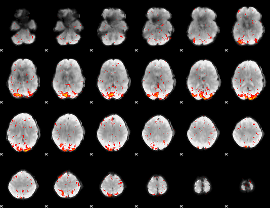
HAFNI result component: 373
Overlap rate: 0.691
 |
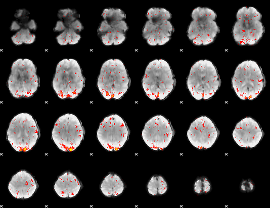
HAFNI result component: 061
Overlap rate: 0.52554
 |
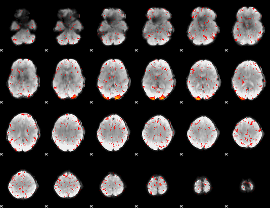
HAFNI result component: 078
Overlap rate: 0.39272
 |
Correlation: 0.14397
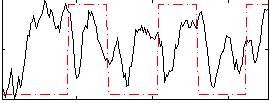 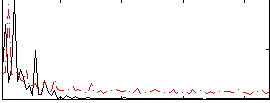 |
Correlation: 0.37468
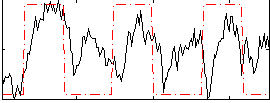 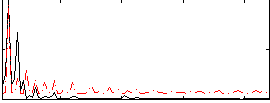 |
Correlation: 0.36525
  |
Subject: 54 |
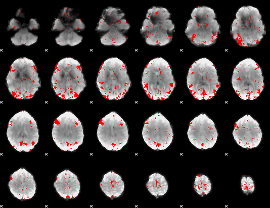
GLM result cope: 01, threshold: 3
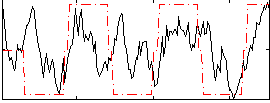
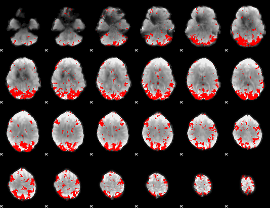 |
GLM result cope: 02, threshold: 3
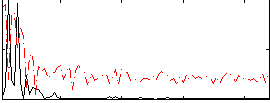
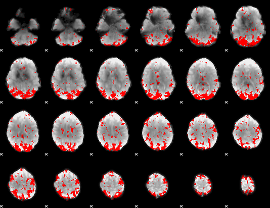 |
GLM result cope: 03, threshold: 2.5
 |
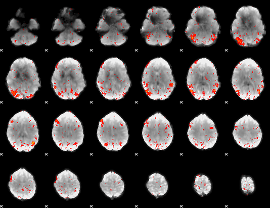
HAFNI result component: 123
Overlap rate: 0.24544
 |
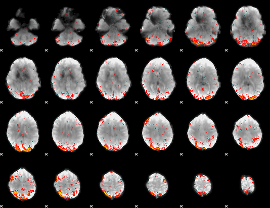
HAFNI result component: 238
Overlap rate: 0.46677
 |
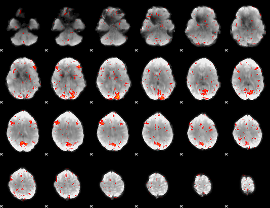
HAFNI result component: 318
Overlap rate: 0.35279
 |
Correlation: 0.28198
  |
Correlation: 0.24771
  |
Correlation: 0.29425
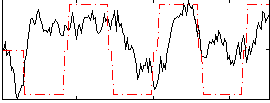 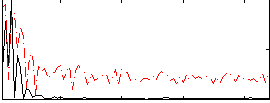 |
Subject: 55 |
GLM result cope: 01, threshold: 2.5
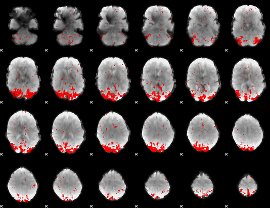 |
GLM result cope: 02, threshold: 3
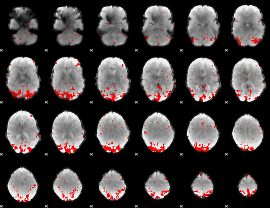 |
GLM result cope: 03, threshold: 2.5
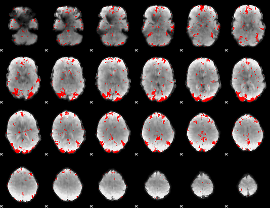 |
HAFNI result component: 006
Overlap rate: 0.55167
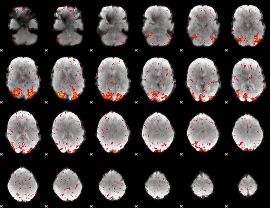 |
HAFNI result component: 399
Overlap rate: 0.56381
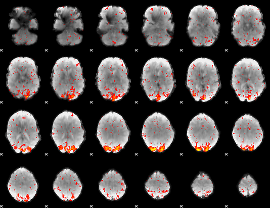 |
HAFNI result component: 390
Overlap rate: 0.52972
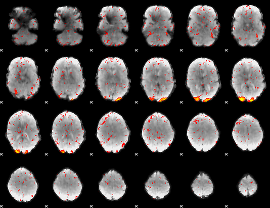 |
Correlation: 0.44769
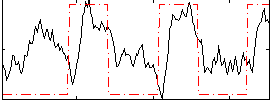 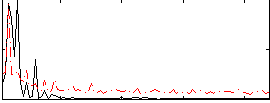 |
Correlation: 0.36902
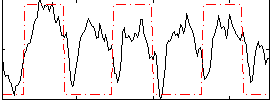 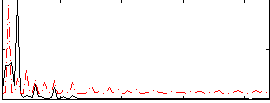 |
Correlation: 0.65456
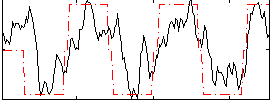 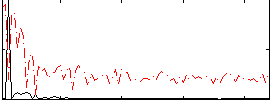 |
Subject: 56 |
GLM result cope: 01, threshold: 2
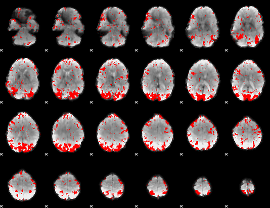 |
GLM result cope: 02, threshold: 2.5
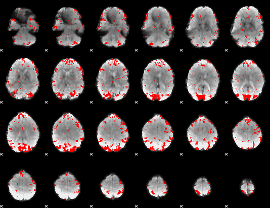 |
GLM result cope: 03, threshold: 2.5
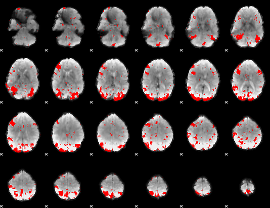 |
HAFNI result component: 203
Overlap rate: 0.61295
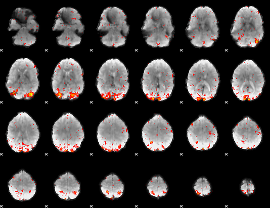 |
HAFNI result component: 168
Overlap rate: 0.5845
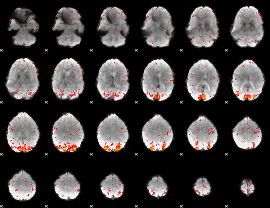 |
HAFNI result component: 010
Overlap rate: 0.55434
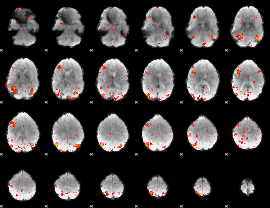 |
Correlation: 0.32773
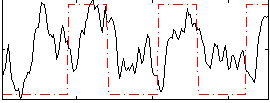 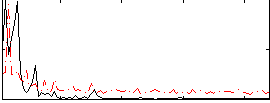 |
Correlation: 0.24835
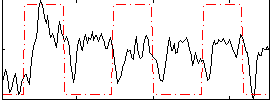 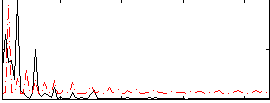 |
Correlation: 0.60094
  |
Subject: 57 |
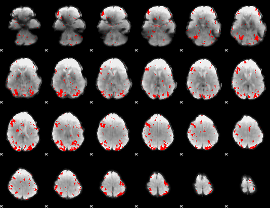
GLM result cope: 01, threshold: 2.5
 |
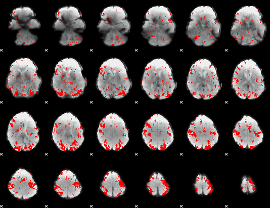
GLM result cope: 02, threshold: 2
 |
GLM result cope: 03, threshold: 2.5
 |
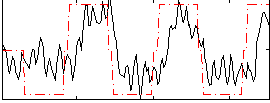
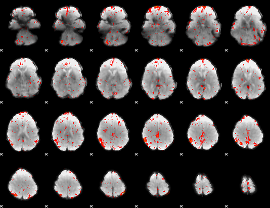
HAFNI result component: 224
Overlap rate: 0.36386
 |
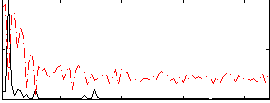
HAFNI result component: 244
Overlap rate: 0.56416
 |
HAFNI result component: 266
Overlap rate: 0.30665
 |
Correlation: 0.46273
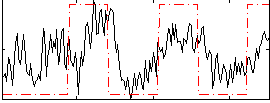 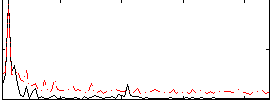 |
Correlation: 0.3924
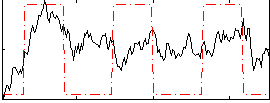 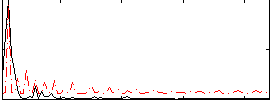 |
Correlation: 0.39491
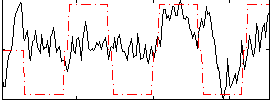 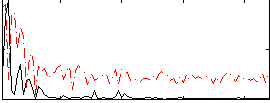 |
Subject: 58 |
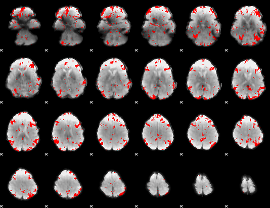
GLM result cope: 01, threshold: 2.5
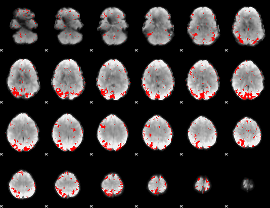 |
GLM result cope: 02, threshold: 2.5
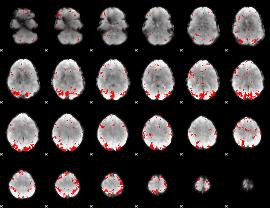 |
GLM result cope: 03, threshold: 2
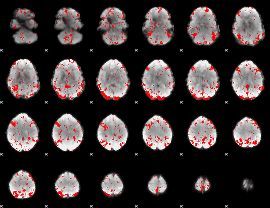 |
HAFNI result component: 353
Overlap rate: 0.45789
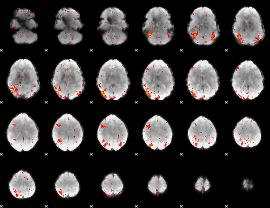 |
HAFNI result component: 051
Overlap rate: 0.57587
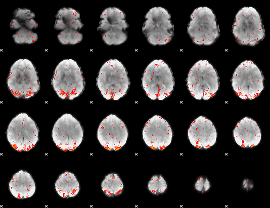 |
HAFNI result component: 309
Overlap rate: 0.33707
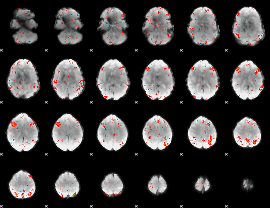 |
Correlation: 0.51475
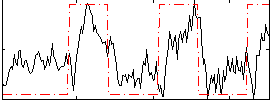 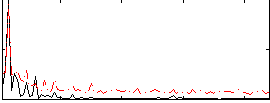 |
Correlation: 0.53271
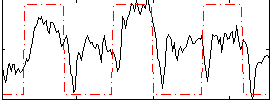  |
Correlation: 0.31156
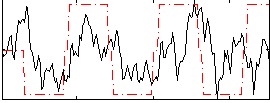 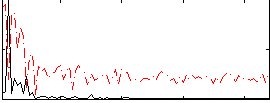 |
Subject: 59 |
GLM result cope: 01, threshold: 2.5
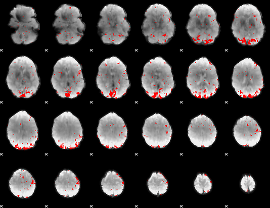 |
GLM result cope: 02, threshold: 2.5
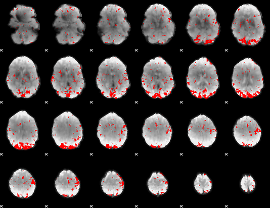 |
GLM result cope: 03, threshold: 2
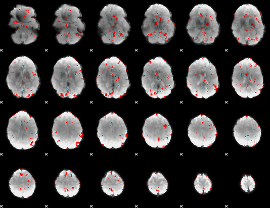 |
HAFNI result component: 147
Overlap rate: 0.48513
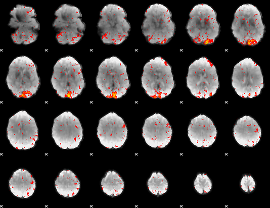 |
HAFNI result component: 327
Overlap rate: 0.7319
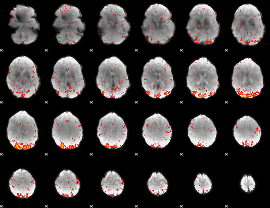 |
HAFNI result component: 370
Overlap rate: 0.46335
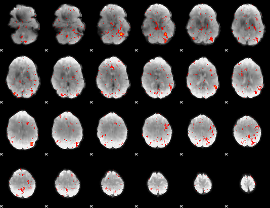 |
Correlation: 0.25031
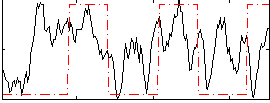 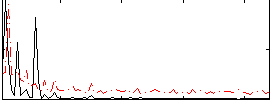 |
Correlation: 0.2848
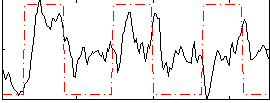 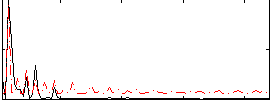 |
Correlation: 0.52182
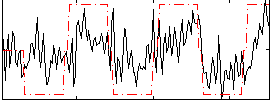 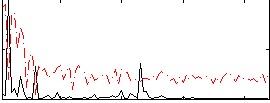 |
Subject: 60 |
GLM result cope: 01, threshold: 3
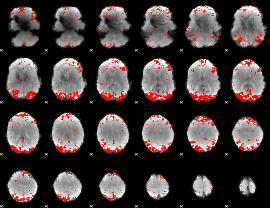 |
GLM result cope: 02, threshold: 3
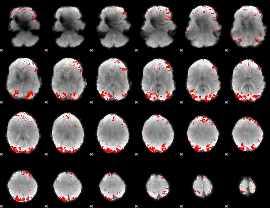 |
GLM result cope: 03, threshold: 3
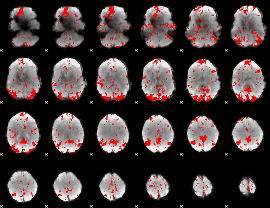 |
HAFNI result component: 350
Overlap rate: 0.44564
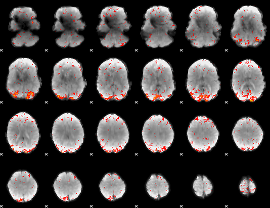 |
HAFNI result component: 158
Overlap rate: 0.25797
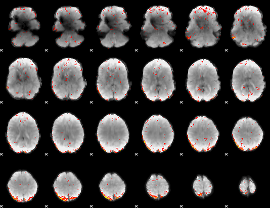 |
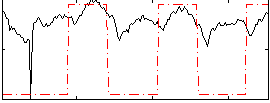
HAFNI result component: 392
Overlap rate: 0.36262
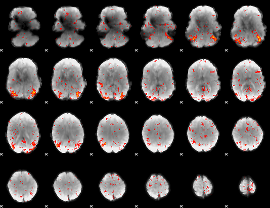 |
Correlation: 0.43957
 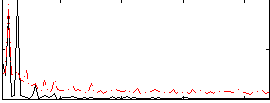 |
Correlation: 0.086956
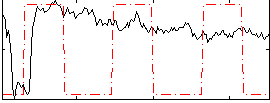  |
Correlation: 0.72177
  |
Subject: 61 |
GLM result cope: 01, threshold: 3
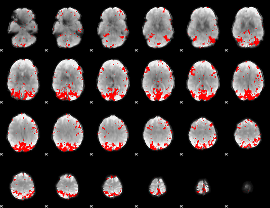 |
GLM result cope: 02, threshold: 3
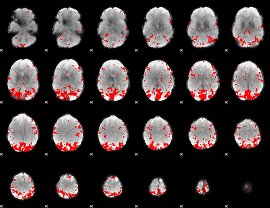 |
GLM result cope: 03, threshold: 2.5
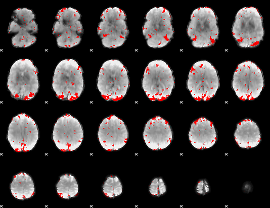 |
HAFNI result component: 149
Overlap rate: 0.29981
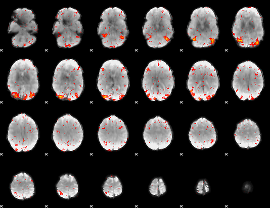 |
HAFNI result component: 072
Overlap rate: 0.37622
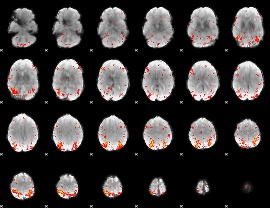 |
HAFNI result component: 277
Overlap rate: 0.49374
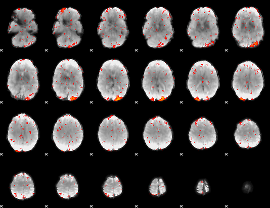 |
Correlation: 0.30748
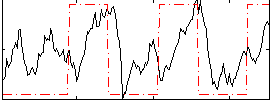  |
Correlation: 0.18963
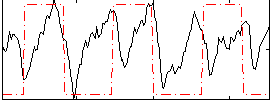 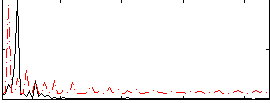 |
Correlation: 0.50283
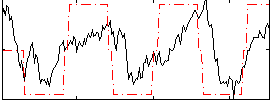 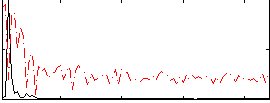 |
Subject: 62 |
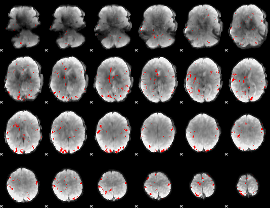
GLM result cope: 01, threshold: 3
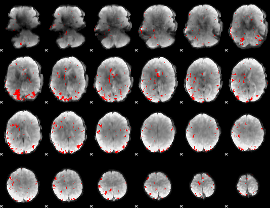 |
GLM result cope: 02, threshold: 3
 |
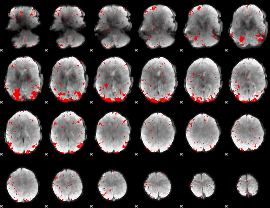
GLM result cope: 03, threshold: 2.5
 |
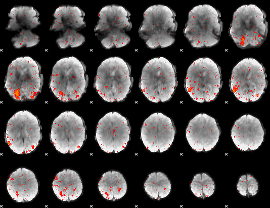
HAFNI result component: 091
Overlap rate: 0.37532
 |
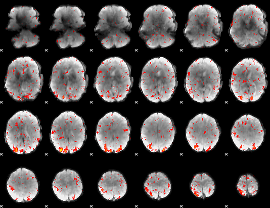
HAFNI result component: 199
Overlap rate: 0.63412
 |
HAFNI result component: 218
Overlap rate: 0.76343
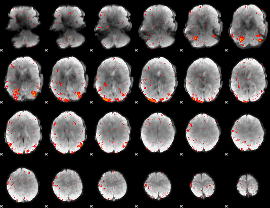 |
Correlation: 0.43191
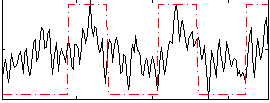 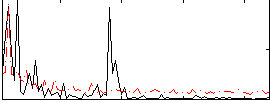 |
Correlation: 0.29032
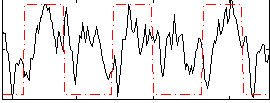  |
Correlation: 0.70167
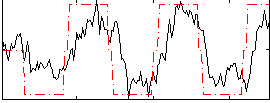 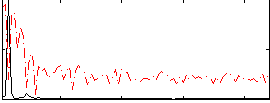 |
Subject: 63 |
GLM result cope: 01, threshold: 2.5
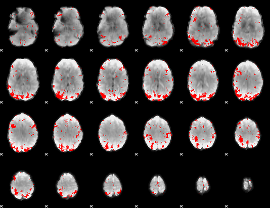 |
GLM result cope: 02, threshold: 2
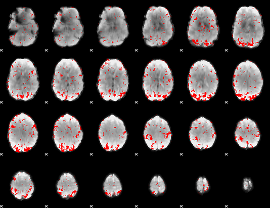 |
GLM result cope: 03, threshold: 2.5
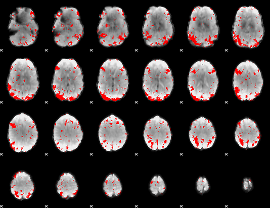 |
HAFNI result component: 162
Overlap rate: 0.66706
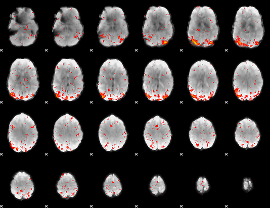 |
HAFNI result component: 392
Overlap rate: 0.62465
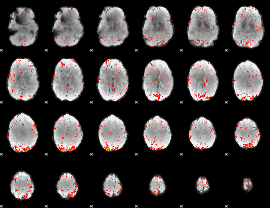 |
HAFNI result component: 199
Overlap rate: 0.40489
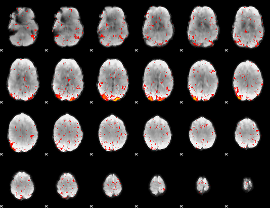 |
Correlation: 0.49742
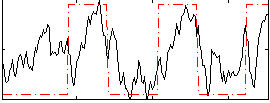 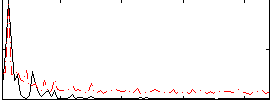 |
Correlation: 0.15381
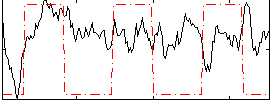 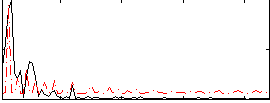 |
Correlation: 0.69253
  |
Subject: 64 |
GLM result cope: 01, threshold: 2.5
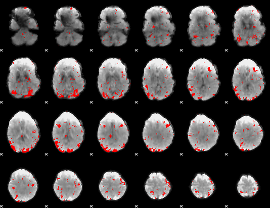 |
GLM result cope: 02, threshold: 2
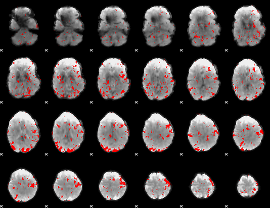 |
GLM result cope: 03, threshold: 2.5
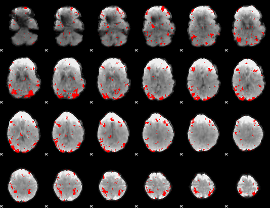 |
HAFNI result component: 020
Overlap rate: 0.70833
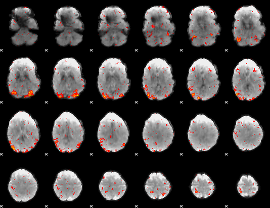 |
HAFNI result component: 372
Overlap rate: 0.66058
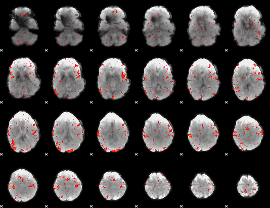 |
HAFNI result component: 209
Overlap rate: 0.34607
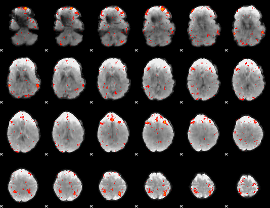 |
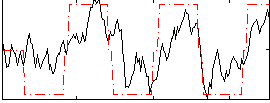
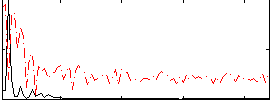
Correlation: 0.58643
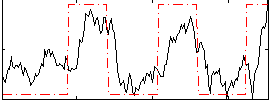 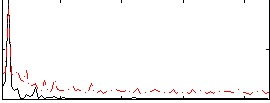 |
Correlation: 0.25002
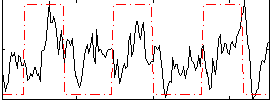 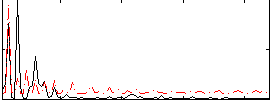 |
Correlation: 0.61362
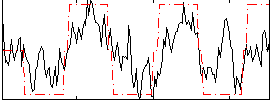 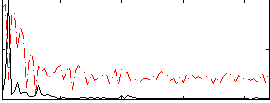 |
Subject: 65 |
GLM result cope: 01, threshold: 2.5
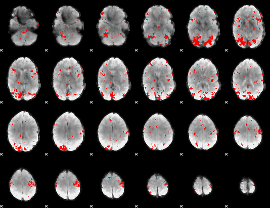 |
GLM result cope: 02, threshold: 2.5
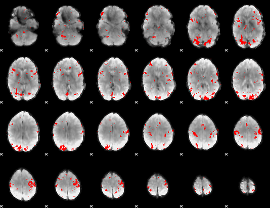 |
GLM result cope: 03, threshold: 2.5
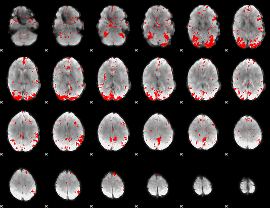 |
HAFNI result component: 153
Overlap rate: 0.65253
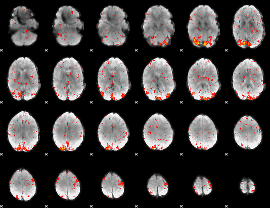 |
HAFNI result component: 273
Overlap rate: 0.57877
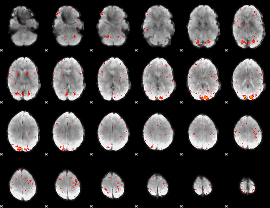 |
HAFNI result component: 143
Overlap rate: 0.42908
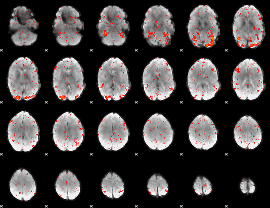 |
Correlation: 0.14388
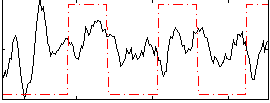 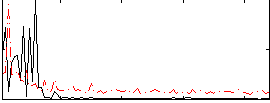 |
Correlation: 0.38831
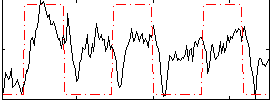 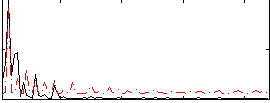 |
Correlation: 0.67188
  |
Subject: 66 |
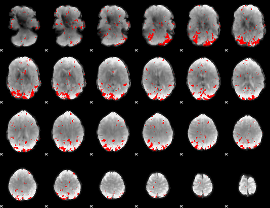
GLM result cope: 01, threshold: 2.5
 |
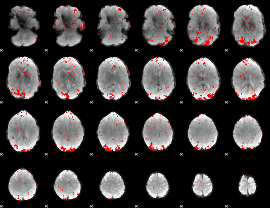
GLM result cope: 02, threshold: 2.5
 |
GLM result cope: 03, threshold: 3
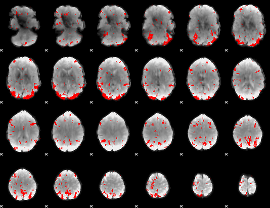 |
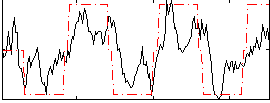
HAFNI result component: 221
Overlap rate: 0.63818
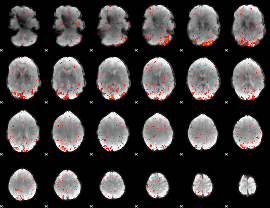 |
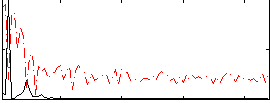
HAFNI result component: 299
Overlap rate: 0.62776
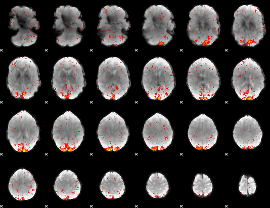 |
HAFNI result component: 314
Overlap rate: 0.64456
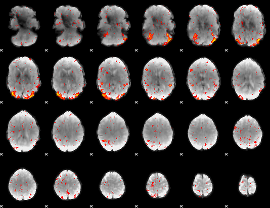 |
Correlation: 0.27468
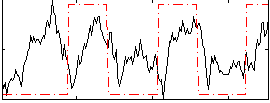 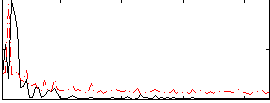 |
Correlation: 0.45519
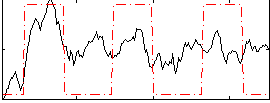  |
Correlation: 0.63449
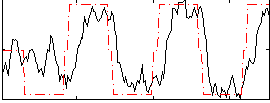  |
Subject: 67 |
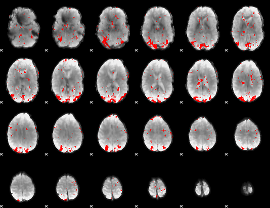
GLM result cope: 01, threshold: 2.5
 |
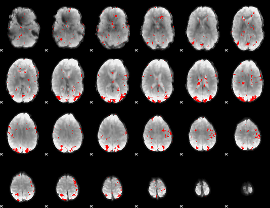
GLM result cope: 02, threshold: 2.5
 |
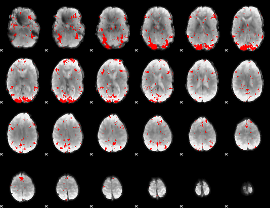
GLM result cope: 03, threshold: 2.5
 |
HAFNI result component: 017
Overlap rate: 0.52737
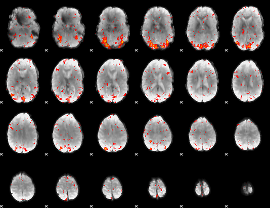 |
HAFNI result component: 221
Overlap rate: 0.5808
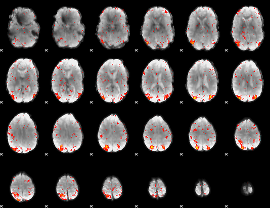 |
HAFNI result component: 201
Overlap rate: 0.64067
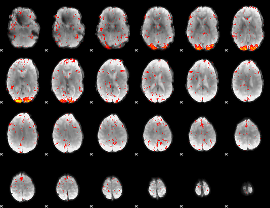 |
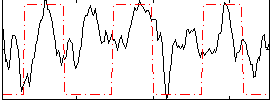
Correlation: 0.5635
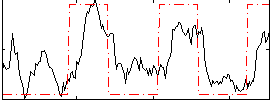 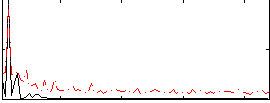 |
Correlation: 0.24043
 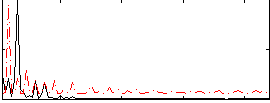 |
Correlation: 0.634
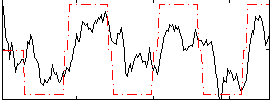  |
Subject: 68 |
GLM result cope: 01, threshold: 2.5
 |
GLM result cope: 02, threshold: 3
 |
GLM result cope: 03, threshold: 2.5
 |
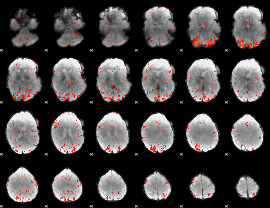
HAFNI result component: 389
Overlap rate: 0.59813
 |
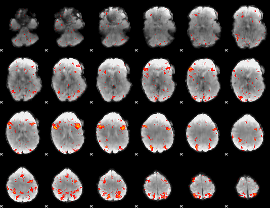
HAFNI result component: 344
Overlap rate: 0.46869
 |
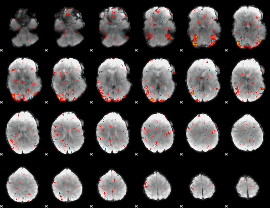
HAFNI result component: 042
Overlap rate: 0.8824
 |
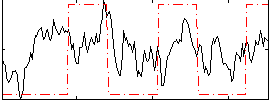
Correlation: 0.27747
  |
Correlation: 0.3354
  |
Correlation: 0.52931
  |















































































































































































































































































































































































































































































































































































































































































































































































































































